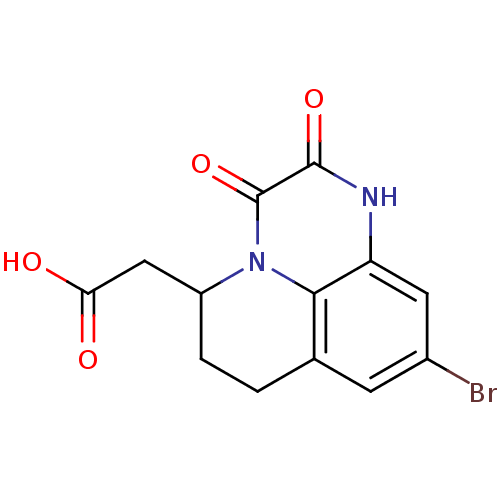
BDBM50038043 (9-Bromo-2,3-dioxo-2,3,6,7-tetrahydro-1H,5H-pyrido[1,2,3-de]quinoxalin-5-yl)-acetic acid::CHEMBL290555
SMILES OC(=O)CC1CCc2cc(Br)cc3[nH]c(=O)c(=O)n1c23
InChI Key InChIKey=RDSPPHOSNHTINE-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 5 hits for monomerid = 50038043
Found 5 hits for monomerid = 50038043
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 9.90nMAssay Description:Displacement of [3H]DCKA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 9.90nMAssay Description:Affinity for the glycine binding site of the NMDA receptor using [3H]- 5,7- dichloro -kynurenic acid as radio-ligandMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: >3.00E+4nMAssay Description:Displacement of [3H]kainic acid from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 4.60E+3nMAssay Description:Displacement of [3H]AMPA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: >3.00E+4nMAssay Description:Displacement of [3H]CGP-39653 from NMDA receptor (unknown origin)More data for this Ligand-Target Pair