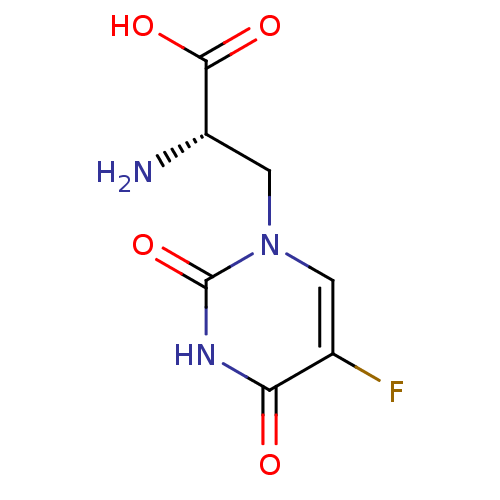
BDBM50060635 (S)-2-Amino-3-(5-fluoro-2,4-dioxo-3,4-dihydro-2H-pyrimidin-1-yl)-propionic acid::(S)-2-amino-3-(5-fluoro-2,4-dioxo-3,4-dihydropyrimidin-1(2H)-yl)propanoic acid::(S)-5-Fluorowillardiine::2-AMINO-3-(5-FLUORO-2,4-DIOXO-3,4-DIHYDRO-2H-PYRIMIDIN-1-YL)-PROPIONIC ACID::CHEMBL123132::Fluorowillardiine
SMILES C1=C(C(=O)NC(=O)N1C[C@@H](C(=O)O)N)F
InChI Key InChIKey=DBWPFHJYSTVBCZ-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 23 hits for monomerid = 50060635
Found 23 hits for monomerid = 50060635
TargetGlutamate receptor 1(Rat)
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
Affinity DataKi: 2.90nMAssay Description:Displacement of (R,S)-[5-methyl-3H]AMPA from rat recombinant flop iGluR1 expressed in Sf9 cellsMore data for this Ligand-Target Pair
TargetGlutamate receptor 2(Rat)
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
Affinity DataKi: 4nMAssay Description:Displacement of (R,S)-[5-methyl-3H]AMPA from rat recombinant flop iGluR2(R) expressed in Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]AMPA from human Ionotropic glutamate receptor AMPA 1 expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Displacement of [3H]AMPA from human Ionotropic glutamate receptor AMPA 2 expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 70nMAssay Description:Displacement of (R,S)-[5-methyl-3H]AMPA from rat recombinant flop iGluR3 expressed in Sf9 cellsMore data for this Ligand-Target Pair
TargetGlutamate receptor 4(Rat)
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
Affinity DataKi: 123nMAssay Description:Displacement of (R,S)-[5-methyl-3H]AMPA from rat recombinant flop iGluR4 expressed in Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 305nMAssay Description:Displacement of [3H]AMPA from human Ionotropic glutamate receptor AMPA 4 expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataEC50: 380nMAssay Description:Agonist activity at recombinant GluA1 receptor expressed in Xenopus oocytesMore data for this Ligand-Target Pair
TargetGlutamate receptor 1(Rat)
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
Affinity DataEC50: 380nMAssay Description:Agonist activity at cyclothiazide-desensitized rat recombinant flip iGluR1 expressed in Xenopus laevis oocytes by two electrode voltage-clamp electro...More data for this Ligand-Target Pair
Affinity DataEC50: 382nMAssay Description:Agonist activity at recombinant GluA1 receptor flip isoform expressed in Xenopus oocytesMore data for this Ligand-Target Pair
TargetGlutamate receptor 2(Rat)
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
Affinity DataEC50: 460nMAssay Description:Agonist activity at cyclothiazide-desensitized rat recombinant flip iGluR2(Q) expressed in Xenopus laevis oocytes by two electrode voltage-clamp elec...More data for this Ligand-Target Pair
Affinity DataEC50: 463nMAssay Description:Agonist activity at recombinant GluA2 receptor flip isoform expressed in Xenopus oocytesMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, kainate 1(Rat)
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
Affinity DataKi: 1.38E+3nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5(Q)1b expressed in Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 1.82E+3nMAssay Description:Displacement of [3H]kainate from human Ionotropic glutamate receptor ionotropic kainate 1 expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGlutamate receptor 4(Rat)
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
Affinity DataEC50: 1.19E+4nMAssay Description:Agonist activity at cyclothiazide-desensitized rat recombinant flip iGluR4 expressed in Xenopus laevis oocytes by two electrode voltage-clamp electro...More data for this Ligand-Target Pair
Affinity DataEC50: 2.09E+4nMAssay Description:Agonist activity at recombinant GluA3 receptor expressed in Xenopus oocytesMore data for this Ligand-Target Pair
Affinity DataEC50: 2.09E+4nMAssay Description:Agonist activity at cyclothiazide-desensitized rat recombinant flip iGluR3 expressed in Xenopus laevis oocytes by two electrode voltage-clamp electro...More data for this Ligand-Target Pair
Affinity DataEC50: 2.09E+4nMAssay Description:Agonist activity at recombinant GluA3 receptor flip isoform expressed in Xenopus oocytesMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, kainate 2(Rat)
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
European Research Centre For Drug Discovery and Development (Natsyndrugs)
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]kainic acid from rat recombinant iGluR6(V,C,R) receptor expressed in Sf9 cellsMore data for this Ligand-Target Pair
 3D Structure (crystal)
3D Structure (crystal)