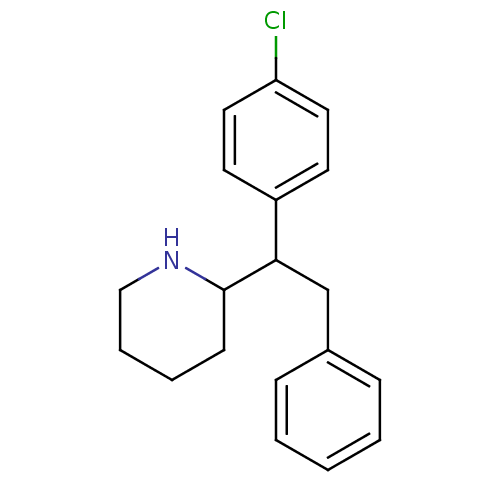
BDBM50202399 (RR/SS)-2-[1-(4-chlorophenyl)-2-phenylethyl]piperidine::(RS/SR)-2-[1-(4-chlorophenyl)-2-phenylethyl]piperidine::CHEMBL373773
SMILES Clc1ccc(cc1)C(Cc1ccccc1)C1CCCCN1
InChI Key InChIKey=SOCRIYWOGGANHT-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 12 hits for monomerid = 50202399
Found 12 hits for monomerid = 50202399
TargetSodium-dependent dopamine transporter(Human)
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Affinity DataIC50: 370nMAssay Description:Inhibition of [3H]DA uptake at human DAT expressing HEK293 cellsMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Affinity DataIC50: 390nMAssay Description:Inhibition of [3H]DA uptake at human DAT expressing HEK293 cellsMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Affinity DataKi: 440nMAssay Description:Displacement of [125I]RTI-55 from human DAT expressing HEK293 cellsMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Human)
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Affinity DataKi: 440nMAssay Description:Displacement of [125I]RTI-55 from human DAT expressing HEK293 cellsMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Human)
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of [3H]5-HT uptake into human SERT expressing HEK293 cellsMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Human)
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Affinity DataKi: 1.10E+3nMAssay Description:Displacement of [125I]RTI-55 from human SERT expressing HEK293 cellsMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Human)
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of [3H]5-HT uptake into human SERT expressing HEK293 cellsMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Human)
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Affinity DataKi: 1.10E+3nMAssay Description:Displacement of [125I]RTI-55 from human SERT expressing HEK293 cellsMore data for this Ligand-Target Pair
TargetSodium-dependent noradrenaline transporter(Human)
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Affinity DataKi: 2.90E+3nMAssay Description:Displacement of [125I]RTI-55 from human NET expressing HEK293 cellsMore data for this Ligand-Target Pair
TargetSodium-dependent noradrenaline transporter(Human)
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Affinity DataIC50: 2.90E+3nMAssay Description:Inhibition of [3H]NE uptake into human NET expressing HEK293 cellsMore data for this Ligand-Target Pair
TargetSodium-dependent noradrenaline transporter(Human)
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Affinity DataKi: 2.90E+3nMAssay Description:Displacement of [125I]RTI-55 from human NET expressing HEK293 cellsMore data for this Ligand-Target Pair
TargetSodium-dependent noradrenaline transporter(Human)
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Massachusetts College of Pharmacy and Health Sciences
Curated by ChEMBL
Affinity DataIC50: 8.80E+3nMAssay Description:Inhibition of [3H]NE uptake into human NET expressing HEK293 cellsMore data for this Ligand-Target Pair