Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Sodium-dependent dopamine transporter
Ligand
BDBM50022723
Substrate
n/a
Meas. Tech.
ChEMBL_792746 (CHEMBL1930661)
EC50
24.8±n/a nM
Citation
 Lewin, AH; Miller, GM; Gilmour, B Trace amine-associated receptor 1 is a stereoselective binding site for compounds in the amphetamine class. Bioorg Med Chem 19:7044-8 (2011) [PubMed] Article
Lewin, AH; Miller, GM; Gilmour, B Trace amine-associated receptor 1 is a stereoselective binding site for compounds in the amphetamine class. Bioorg Med Chem 19:7044-8 (2011) [PubMed] Article More Info.:
Target
Name:
Sodium-dependent dopamine transporter
Synonyms:
DA transporter | DAT | DAT1 | Dopamine transporter (DAT) | Dopamine transporter protein (DAT) | Monoamine transporter | SC6A3_HUMAN | SLC6A3 | Sodium-dependent dopamine transporter | Sodium-dependent dopamine transporter (DAT) | Solute carrier family 6 member 3
Type:
Multi-pass membrane protein
Mol. Mass.:
68497.11
Organism:
Homo sapiens (Human)
Description:
Q01959
Residue:
620
Sequence:
MSKSKCSVGLMSSVVAPAKEPNAVGPKEVELILVKEQNGVQLTSSTLTNPRQSPVEAQDRETWGKKIDFLLSVIGFAVDLANVWRFPYLCYKNGGGAFLVPYLLFMVIAGMPLFYMELALGQFNREGAAGVWKICPILKGVGFTVILISLYVGFFYNVIIAWALHYLFSSFTTELPWIHCNNSWNSPNCSDAHPGDSSGDSSGLNDTFGTTPAAEYFERGVLHLHQSHGIDDLGPPRWQLTACLVLVIVLLYFSLWKGVKTSGKVVWITATMPYVVLTALLLRGVTLPGAIDGIRAYLSVDFYRLCEASVWIDAATQVCFSLGVGFGVLIAFSSYNKFTNNCYRDAIVTTSINSLTSFSSGFVVFSFLGYMAQKHSVPIGDVAKDGPGLIFIIYPEAIATLPLSSAWAVVFFIMLLTLGIDSAMGGMESVITGLIDEFQLLHRHRELFTLFIVLATFLLSLFCVTNGGIYVFTLLDHFAAGTSILFGVLIEAIGVAWFYGVGQFSDDIQQMTGQRPSLYWRLCWKLVSPCFLLFVVVVSIVTFRPPHYGAYIFPDWANALGWVIATSSMAMVPIYAAYKFCSLPGSFREKLAYAIAPEKDRELVDRGEVRQFTLRHWLKV
Inhibitor
Name:
BDBM50022723
Synonyms:
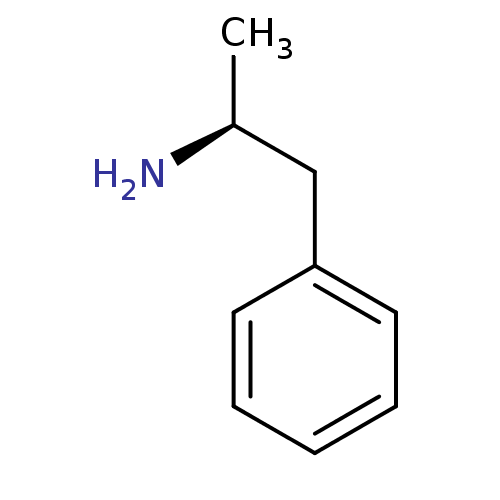
(+)-(S)-amphetamine | (+)-alpha-methylphenethylamine | (+)-alpha-methylphenylethylamine | (2S)-1-phenylpropan-2-amine | (S)-(+)-amphetamine | (S)-(+)-beta-phenylisopropylamine | (S)-1-phenyl-2-aminopropane | (S)-1-phenyl-2-propylamine | (S)-alpha-methylbenzeneethanamine | (S)-amphetamine | (alphaS)-alpha-methylbenzeneethanamine | CHEMBL612 | Dextroamphetamine | d-amphetamine | dexamphetamine
Type:
Small organic molecule
Emp. Form.:
C9H13N
Mol. Mass.:
135.2062
SMILES:
C[C@H](N)Cc1ccccc1 |r|