Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Beta-secretase 1
Ligand
BDBM50383846
Substrate
n/a
Meas. Tech.
ChEMBL_819409 (CHEMBL2033076)
IC50
0.700000±n/a nM
Citation
 Monenschein, H; Horne, DB; Bartberger, MD; Hitchcock, SA; Nguyen, TT; Patel, VF; Pennington, LD; Zhong, W Structure guided P1' modifications of HEA derivedß-secretase inhibitors for the treatment of Alzheimer's disease. Bioorg Med Chem Lett 22:3607-11 (2012) [PubMed] Article
Monenschein, H; Horne, DB; Bartberger, MD; Hitchcock, SA; Nguyen, TT; Patel, VF; Pennington, LD; Zhong, W Structure guided P1' modifications of HEA derivedß-secretase inhibitors for the treatment of Alzheimer's disease. Bioorg Med Chem Lett 22:3607-11 (2012) [PubMed] Article More Info.:
Target
Name:
Beta-secretase 1
Synonyms:
ASP2 | Asp 2 | Aspartyl protease 2 | BACE | BACE1 | BACE1_HUMAN | Beta-secretase (BACE) | Beta-secretase 1 | Beta-secretase 1 (BACE 1) | Beta-secretase 1 (BACE-1) | Beta-secretase 1 (BACE1) | Beta-site APP cleaving enzyme 1 | Beta-site amyloid precursor protein cleaving enzyme 1 | KIAA1149 | Memapsin-2 | Membrane-associated aspartic protease 2 | beta-Secretase (BACE-1) | beta-Secretase (BACE1)
Type:
Protein
Mol. Mass.:
55755.10
Organism:
Homo sapiens (Human)
Description:
P56817
Residue:
501
Sequence:
MAQALPWLLLWMGAGVLPAHGTQHGIRLPLRSGLGGAPLGLRLPRETDEEPEEPGRRGSFVEMVDNLRGKSGQGYYVEMTVGSPPQTLNILVDTGSSNFAVGAAPHPFLHRYYQRQLSSTYRDLRKGVYVPYTQGKWEGELGTDLVSIPHGPNVTVRANIAAITESDKFFINGSNWEGILGLAYAEIARPDDSLEPFFDSLVKQTHVPNLFSLQLCGAGFPLNQSEVLASVGGSMIIGGIDHSLYTGSLWYTPIRREWYYEVIIVRVEINGQDLKMDCKEYNYDKSIVDSGTTNLRLPKKVFEAAVKSIKAASSTEKFPDGFWLGEQLVCWQAGTTPWNIFPVISLYLMGEVTNQSFRITILPQQYLRPVEDVATSQDDCYKFAISQSSTGTVMGAVIMEGFYVVFDRARKRIGFAVSACHVHDEFRTAAVEGPFVTLDMEDCGYNIPQTDESTLMTIAYVMAAICALFMLPLCLMVCQWRCLRCLRQQHDDFADDISLLK
Inhibitor
Name:
BDBM50383846
Synonyms:
CHEMBL2031142
Type:
Small organic molecule
Emp. Form.:
C29H39F2N3O3
Mol. Mass.:
515.6351
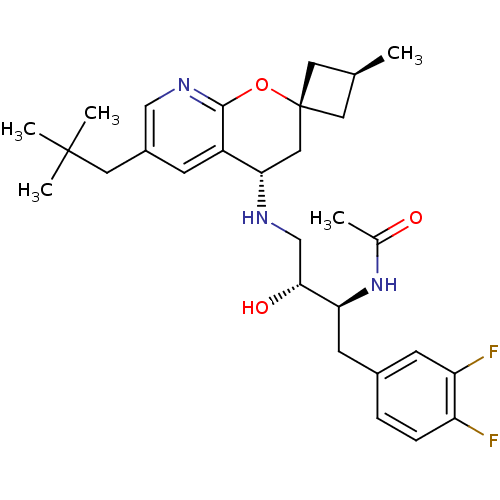
SMILES:
C[C@H]1C[C@]2(C1)C[C@H](NC[C@@H](O)[C@H](Cc1ccc(F)c(F)c1)NC(C)=O)c1cc(CC(C)(C)C)cnc1O2 |r,wU:6.7,1.0,wD:9.10,11.22,3.3,(31.56,-19.68,;30.47,-20.77,;30.47,-22.31,;28.93,-22.31,;28.93,-20.77,;27.59,-21.54,;26.25,-22.32,;24.91,-21.56,;23.58,-22.33,;22.25,-21.57,;22.24,-20.03,;20.91,-22.34,;20.92,-23.88,;19.59,-24.66,;19.61,-26.19,;18.28,-26.97,;16.93,-26.21,;15.61,-26.99,;16.93,-24.67,;15.59,-23.9,;18.26,-23.89,;19.58,-21.58,;19.57,-20.04,;18.23,-19.28,;20.9,-19.26,;26.26,-23.86,;24.93,-24.63,;24.93,-26.18,;23.6,-26.95,;23.6,-28.49,;22.26,-29.25,;24.93,-29.26,;23.58,-30.02,;26.27,-26.95,;27.6,-26.18,;27.6,-24.63,;28.93,-23.86,)|