Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Focal adhesion kinase 1
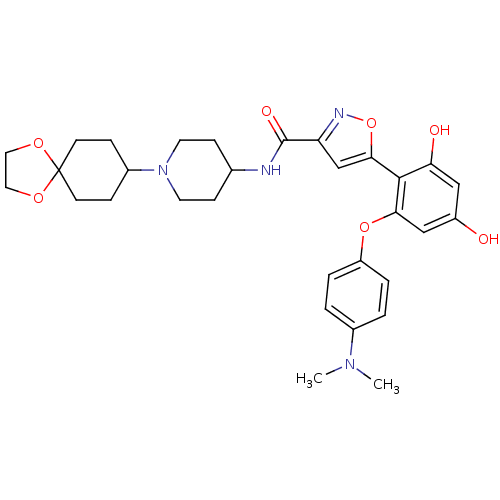
Ligand
BDBM50442757
Substrate
n/a
Meas. Tech.
ChEMBL_989480 (CHEMBL2444007)
IC50
>10.0±n/a nM
Citation
 Brasca, MG; Mantegani, S; Amboldi, N; Bindi, S; Caronni, D; Casale, E; Ceccarelli, W; Colombo, N; De Ponti, A; Donati, D; Ermoli, A; Fachin, G; Felder, ER; Ferguson, RD; Fiorelli, C; Guanci, M; Isacchi, A; Pesenti, E; Polucci, P; Riceputi, L; Sola, F; Visco, C; Zuccotto, F; Fogliatto, G Discovery of NMS-E973 as novel, selective and potent inhibitor of heat shock protein 90 (Hsp90). Bioorg Med Chem 21:7047-63 (2013) [PubMed] Article
Brasca, MG; Mantegani, S; Amboldi, N; Bindi, S; Caronni, D; Casale, E; Ceccarelli, W; Colombo, N; De Ponti, A; Donati, D; Ermoli, A; Fachin, G; Felder, ER; Ferguson, RD; Fiorelli, C; Guanci, M; Isacchi, A; Pesenti, E; Polucci, P; Riceputi, L; Sola, F; Visco, C; Zuccotto, F; Fogliatto, G Discovery of NMS-E973 as novel, selective and potent inhibitor of heat shock protein 90 (Hsp90). Bioorg Med Chem 21:7047-63 (2013) [PubMed] Article More Info.:
Target
Name:
Focal adhesion kinase 1
Synonyms:
FADK 1 | FAK | FAK1 | FAK1_HUMAN | FLT4 | FRNK | Focal adhesion kinase (FAK) | Focal adhesion kinase (PTK2) | Focal adhesion kinase 1 (FAK) | Focal adhesion kinase 1/vascular endothelial growth factor receptor 3 | Focal adhesion kinase-related nonkinase | PPP1R71 | PTK2 | Protein phosphatase 1 regulatory subunit 71 | Protein-tyrosine kinase 2 | VHL/Focal adhesion kinase 1 | p125FAK | pp125FAK
Type:
Tyrosine-protein kinase
Mol. Mass.:
119233.17
Organism:
Homo sapiens (Human)
Description:
Q05397
Residue:
1052
Sequence:
MAAAYLDPNLNHTPNSSTKTHLGTGMERSPGAMERVLKVFHYFESNSEPTTWASIIRHGDATDVRGIIQKIVDSHKVKHVACYGFRLSHLRSEEVHWLHVDMGVSSVREKYELAHPPEEWKYELRIRYLPKGFLNQFTEDKPTLNFFYQQVKSDYMLEIADQVDQEIALKLGCLEIRRSYWEMRGNALEKKSNYEVLEKDVGLKRFFPKSLLDSVKAKTLRKLIQQTFRQFANLNREESILKFFEILSPVYRFDKECFKCALGSSWIISVELAIGPEEGISYLTDKGCNPTHLADFTQVQTIQYSNSEDKDRKGMLQLKIAGAPEPLTVTAPSLTIAENMADLIDGYCRLVNGTSQSFIIRPQKEGERALPSIPKLANSEKQGMRTHAVSVSETDDYAEIIDEEDTYTMPSTRDYEIQRERIELGRCIGEGQFGDVHQGIYMSPENPALAVAIKTCKNCTSDSVREKFLQEALTMRQFDHPHIVKLIGVITENPVWIIMELCTLGELRSFLQVRKYSLDLASLILYAYQLSTALAYLESKRFVHRDIAARNVLVSSNDCVKLGDFGLSRYMEDSTYYKASKGKLPIKWMAPESINFRRFTSASDVWMFGVCMWEILMHGVKPFQGVKNNDVIGRIENGERLPMPPNCPPTLYSLMTKCWAYDPSRRPRFTELKAQLSTILEEEKAQQEERMRMESRRQATVSWDSGGSDEAPPKPSRPGYPSPRSSEGFYPSPQHMVQTNHYQVSGYPGSHGITAMAGSIYPGQASLLDQTDSWNHRPQEIAMWQPNVEDSTVLDLRGIGQVLPTHLMEERLIRQQQEMEEDQRWLEKEERFLKPDVRLSRGSIDREDGSLQGPIGNQHIYQPVGKPDPAAPPKKPPRPGAPGHLGSLASLSSPADSYNEGVKLQPQEISPPPTANLDRSNDKVYENVTGLVKAVIEMSSKIQPAPPEEYVPMVKEVGLALRTLLATVDETIPLLPASTHREIEMAQKLLNSDLGELINKMKLAQQYVMTSLQQEYKKQMLTAAHALAVDAKNLLDVIDQARLKMLGQTRPH