Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
A disintegrin and metalloproteinase with thrombospondin motifs 13
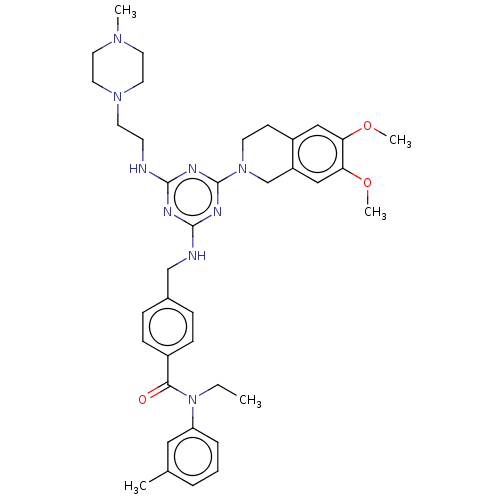
Ligand
BDBM50122449
Substrate
n/a
Meas. Tech.
ChEMBL_1519717 (CHEMBL3624062)
IC50
>10000±n/a nM
Citation
 Ding, Y; O'Keefe, H; DeLorey, JL; Israel, DI; Messer, JA; Chiu, CH; Skinner, SR; Matico, RE; Murray-Thompson, MF; Li, F; Clark, MA; Cuozzo, JW; Arico-Muendel, C; Morgan, BA Discovery of Potent and Selective Inhibitors for ADAMTS-4 through DNA-Encoded Library Technology (ELT). ACS Med Chem Lett 6:888-93 (2015) [PubMed] Article
Ding, Y; O'Keefe, H; DeLorey, JL; Israel, DI; Messer, JA; Chiu, CH; Skinner, SR; Matico, RE; Murray-Thompson, MF; Li, F; Clark, MA; Cuozzo, JW; Arico-Muendel, C; Morgan, BA Discovery of Potent and Selective Inhibitors for ADAMTS-4 through DNA-Encoded Library Technology (ELT). ACS Med Chem Lett 6:888-93 (2015) [PubMed] Article More Info.:
Target
Name:
A disintegrin and metalloproteinase with thrombospondin motifs 13
Synonyms:
ADAM-TS 13 | ADAM-TS13 | ADAMTS-13 | ADAMTS13 | ATS13_HUMAN | C9orf8 | vWF-CP | vWF-cleaving protease | von Willebrand factor-cleaving protease
Type:
PROTEIN
Mol. Mass.:
153616.59
Organism:
Homo sapiens (Human)
Description:
ChEMBL_953678
Residue:
1427
Sequence:
MHQRHPRARCPPLCVAGILACGFLLGCWGPSHFQQSCLQALEPQAVSSYLSPGAPLKGRPPSPGFQRQRQRQRRAAGGILHLELLVAVGPDVFQAHQEDTERYVLTNLNIGAELLRDPSLGAQFRVHLVKMVILTEPEGAPNITANLTSSLLSVCGWSQTINPEDDTDPGHADLVLYITRFDLELPDGNRQVRGVTQLGGACSPTWSCLITEDTGFDLGVTIAHEIGHSFGLEHDGAPGSGCGPSGHVMASDGAAPRAGLAWSPCSRRQLLSLLSAGRARCVWDPPRPQPGSAGHPPDAQPGLYYSANEQCRVAFGPKAVACTFAREHLDMCQALSCHTDPLDQSSCSRLLVPLLDGTECGVEKWCSKGRCRSLVELTPIAAVHGRWSSWGPRSPCSRSCGGGVVTRRRQCNNPRPAFGGRACVGADLQAEMCNTQACEKTQLEFMSQQCARTDGQPLRSSPGGASFYHWGAAVPHSQGDALCRHMCRAIGESFIMKRGDSFLDGTRCMPSGPREDGTLSLCVSGSCRTFGCDGRMDSQQVWDRCQVCGGDNSTCSPRKGSFTAGRAREYVTFLTVTPNLTSVYIANHRPLFTHLAVRIGGRYVVAGKMSISPNTTYPSLLEDGRVEYRVALTEDRLPRLEEIRIWGPLQEDADIQVYRRYGEEYGNLTRPDITFTYFQPKPRQAWVWAAVRGPCSVSCGAGLRWVNYSCLDQARKELVETVQCQGSQQPPAWPEACVLEPCPPYWAVGDFGPCSASCGGGLRERPVRCVEAQGSLLKTLPPARCRAGAQQPAVALETCNPQPCPARWEVSEPSSCTSAGGAGLALENETCVPGADGLEAPVTEGPGSVDEKLPAPEPCVGMSCPPGWGHLDATSAGEKAPSPWGSIRTGAQAAHVWTPAAGSCSVSCGRGLMELRFLCMDSALRVPVQEELCGLASKPGSRREVCQAVPCPARWQYKLAACSVSCGRGVVRRILYCARAHGEDDGEEILLDTQCQGLPRPEPQEACSLEPCPPRWKVMSLGPCSASCGLGTARRSVACVQLDQGQDVEVDEAACAALVRPEASVPCLIADCTYRWHVGTWMECSVSCGDGIQRRRDTCLGPQAQAPVPADFCQHLPKPVTVRGCWAGPCVGQGTPSLVPHEEAAAPGRTTATPAGASLEWSQARGLLFSPAPQPRRLLPGPQENSVQSSACGRQHLEPTGTIDMRGPGQADCAVAIGRPLGEVVTLRVLESSLNCSAGDMLLLWGRLTWRKMCRKLLDMTFSSKTNTLVVRQRCGRPGGGVLLRYGSQLAPETFYRECDMQLFGPWGEIVSPSLSPATSNAGGCRLFINVAPHARIAIHALATNMGAGTEGANASYILIRDTHSLRTTAFHGQQVLYWESESSQAEMEFSEGFLKAQASLRGQYWTLQSWVPEMQDPQSWKGKEGT