Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Androgen receptor
Ligand
BDBM50125052
Substrate
n/a
Meas. Tech.
ChEMBL_1519272 (CHEMBL3624769)
IC50
30000±n/a nM
Citation
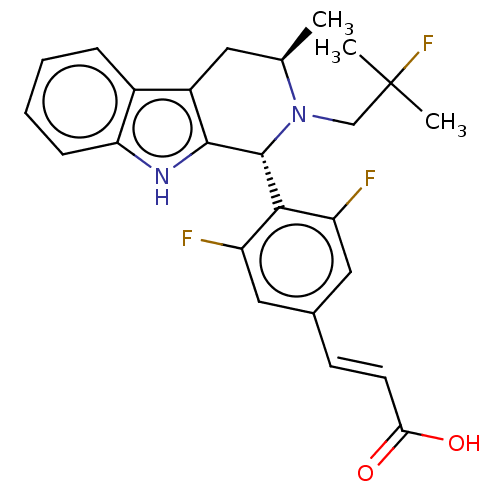
 De Savi, C; Bradbury, RH; Rabow, AA; Norman, RA; de Almeida, C; Andrews, DM; Ballard, P; Buttar, D; Callis, RJ; Currie, GS; Curwen, JO; Davies, CD; Donald, CS; Feron, LJ; Gingell, H; Glossop, SC; Hayter, BR; Hussain, S; Karoutchi, G; Lamont, SG; MacFaul, P; Moss, TA; Pearson, SE; Tonge, M; Walker, GE; Weir, HM; Wilson, Z Optimization of a Novel Binding Motif to (E)-3-(3,5-Difluoro-4-((1R,3R)-2-(2-fluoro-2-methylpropyl)-3-methyl-2,3,4,9-tetrahydro-1H-pyrido[3,4-b]indol-1-yl)phenyl)acrylic Acid (AZD9496), a Potent and Orally Bioavailable Selective Estrogen Receptor Downregulator and Antagonist. J Med Chem 58:8128-40 (2015) [PubMed] Article
De Savi, C; Bradbury, RH; Rabow, AA; Norman, RA; de Almeida, C; Andrews, DM; Ballard, P; Buttar, D; Callis, RJ; Currie, GS; Curwen, JO; Davies, CD; Donald, CS; Feron, LJ; Gingell, H; Glossop, SC; Hayter, BR; Hussain, S; Karoutchi, G; Lamont, SG; MacFaul, P; Moss, TA; Pearson, SE; Tonge, M; Walker, GE; Weir, HM; Wilson, Z Optimization of a Novel Binding Motif to (E)-3-(3,5-Difluoro-4-((1R,3R)-2-(2-fluoro-2-methylpropyl)-3-methyl-2,3,4,9-tetrahydro-1H-pyrido[3,4-b]indol-1-yl)phenyl)acrylic Acid (AZD9496), a Potent and Orally Bioavailable Selective Estrogen Receptor Downregulator and Antagonist. J Med Chem 58:8128-40 (2015) [PubMed] Article More Info.:
Target
Name:
Androgen receptor
Synonyms:
ANDR_HUMAN | AR | Androgen Receptor | Androgen receptor (AR) | Androgen receptor/Baculoviral IAP repeat-containing protein 2 | DHTR | Dihydrotestosterone receptor | NR3C4 | Nuclear receptor subfamily 3 group C member 4
Type:
Receptor
Mol. Mass.:
99185.27
Organism:
Homo sapiens (Human)
Description:
CHO cells were stably transfected with human AR gene.
Residue:
920
Sequence:
MEVQLGLGRVYPRPPSKTYRGAFQNLFQSVREVIQNPGPRHPEAASAAPPGASLLLLQQQQQQQQQQQQQQQQQQQQQQQETSPRQQQQQQGEDGSPQAHRRGPTGYLVLDEEQQPSQPQSALECHPERGCVPEPGAAVAASKGLPQQLPAPPDEDDSAAPSTLSLLGPTFPGLSSCSADLKDILSEASTMQLLQQQQQEAVSEGSSSGRAREASGAPTSSKDNYLGGTSTISDNAKELCKAVSVSMGLGVEALEHLSPGEQLRGDCMYAPLLGVPPAVRPTPCAPLAECKGSLLDDSAGKSTEDTAEYSPFKGGYTKGLEGESLGCSGSAAAGSSGTLELPSTLSLYKSGALDEAAAYQSRDYYNFPLALAGPPPPPPPPHPHARIKLENPLDYGSAWAAAAAQCRYGDLASLHGAGAAGPGSGSPSAAASSSWHTLFTAEEGQLYGPCGGGGGGGGGGGGGGGGGGGGGGGEAGAVAPYGYTRPPQGLAGQESDFTAPDVWYPGGMVSRVPYPSPTCVKSEMGPWMDSYSGPYGDMRLETARDHVLPIDYYFPPQKTCLICGDEASGCHYGALTCGSCKVFFKRAAEGKQKYLCASRNDCTIDKFRRKNCPSCRLRKCYEAGMTLGARKLKKLGNLKLQEEGEASSTTSPTEETTQKLTVSHIEGYECQPIFLNVLEAIEPGVVCAGHDNNQPDSFAALLSSLNELGERQLVHVVKWAKALPGFRNLHVDDQMAVIQYSWMGLMVFAMGWRSFTNVNSRMLYFAPDLVFNEYRMHKSRMYSQCVRMRHLSQEFGWLQITPQEFLCMKALLLFSIIPVDGLKNQKFFDELRMNYIKELDRIIACKRKNPTSCSRRFYQLTKLLDSVQPIARELHQFTFDLLIKSHMVSVDFPEMMAEIISVQVPKILSGKVKPIYFHTQ