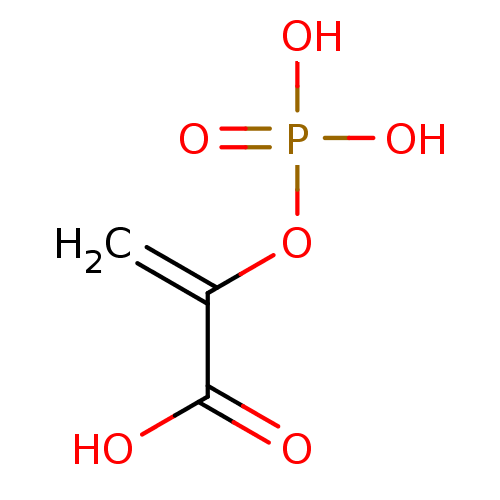
BDBM50366413 PHOSPHOENOLPYRUVATE
SMILES C=C(C(=O)O)OP(=O)(O)O
InChI Key InChIKey=DTBNBXWJWCWCIK-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 4 hits for monomerid = 50366413
Found 4 hits for monomerid = 50366413
Affinity DataKi: 8.5nMpH: 6.0Assay Description:Inhibition of prolidase from swine kidney at a pH 6More data for this Ligand-Target Pair
Affinity DataKi: 300nMAssay Description:Inhibitory constant for inhibition of prolidase in the first phase of biphasic inhibitionMore data for this Ligand-Target Pair
Affinity DataKi: 2.89E+5nMAssay Description:The inhibition of H477R PEPCK by β-sulfopyruvate (βSP), oxalate, and GMPPCP was conducted in the direction of PEP synthesis (OAA → PE...More data for this Ligand-Target Pair
TargetSolute carrier organic anion transporter family member 2A1(Human)
Albert Einstein College of Medicine
Curated by ChEMBL
Albert Einstein College of Medicine
Curated by ChEMBL
Affinity DataKi: 1.30E+7nMAssay Description:TP_TRANSPORTER: inhibition of PGE2 uptake in PGT-expressing HeLa cellsMore data for this Ligand-Target Pair
 3D Structure (crystal)
3D Structure (crystal)