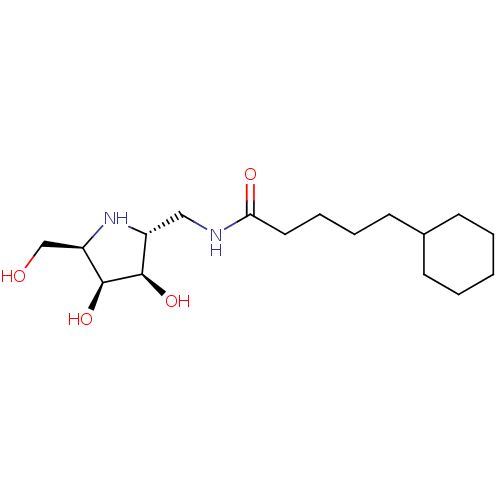
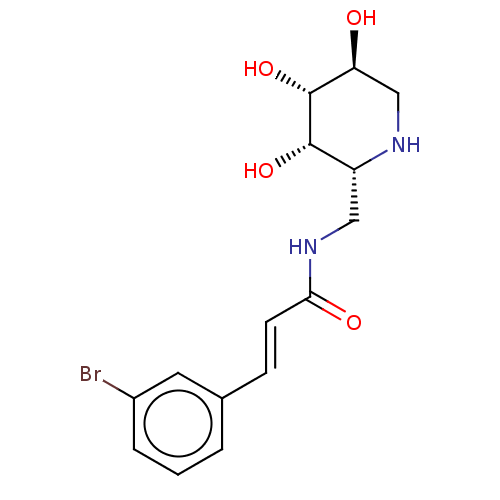
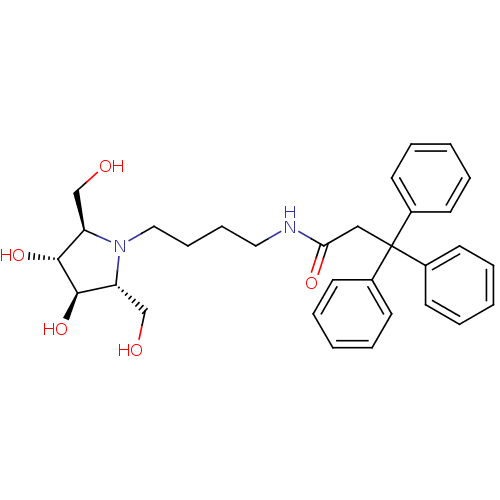
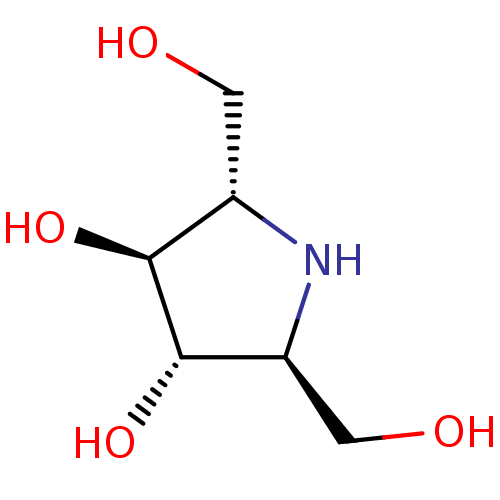
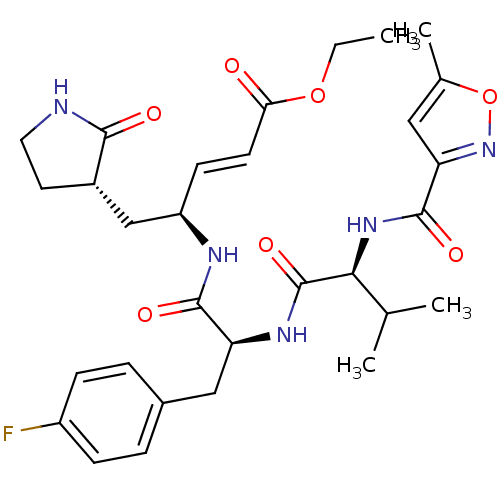
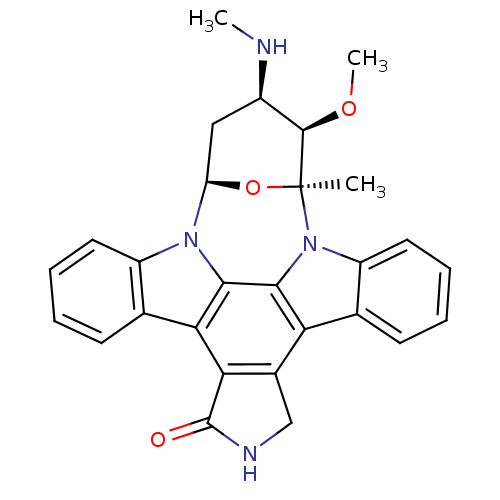
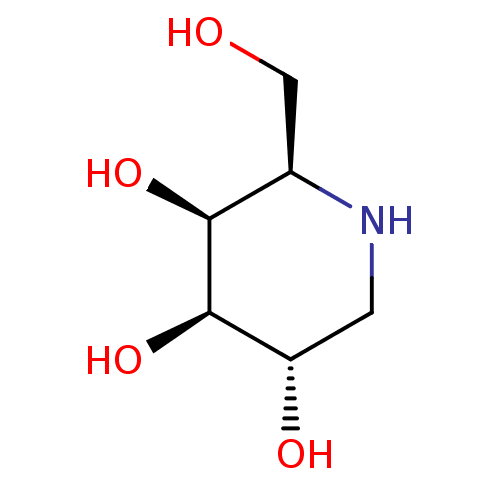
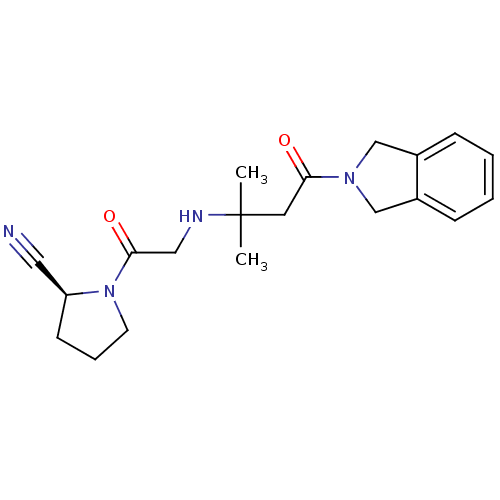
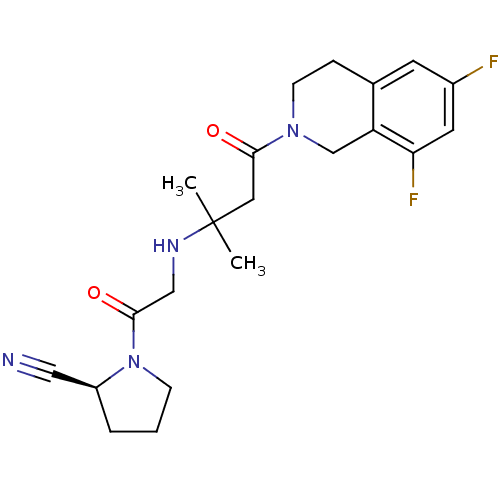
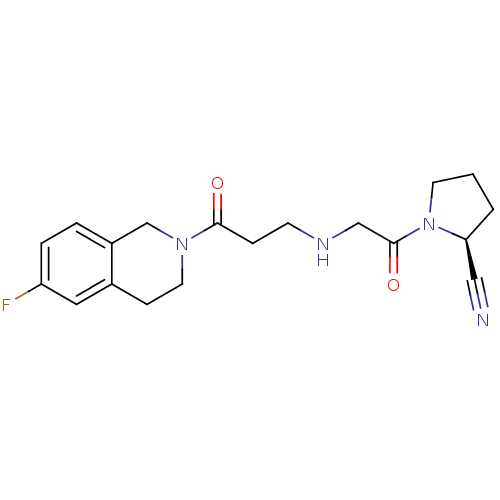
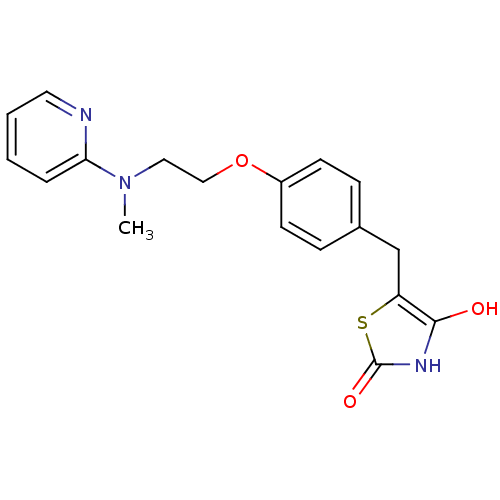
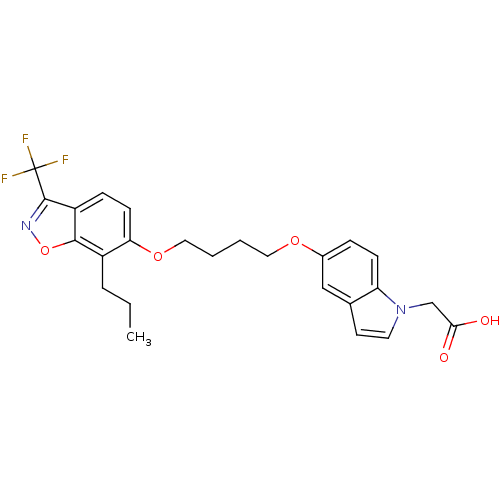
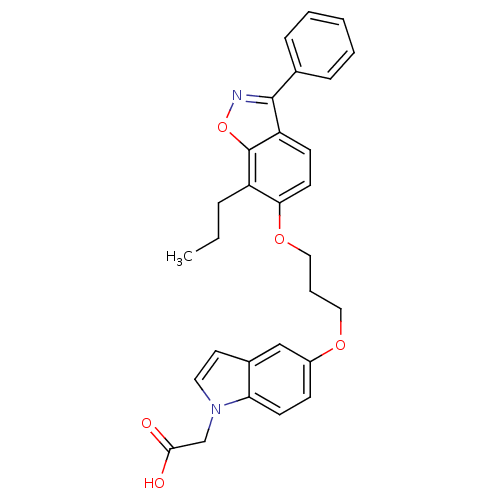
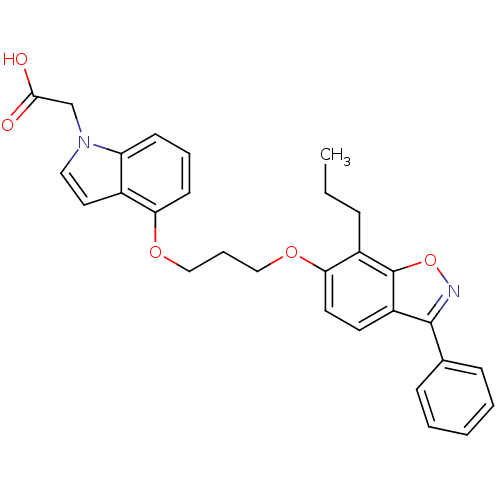
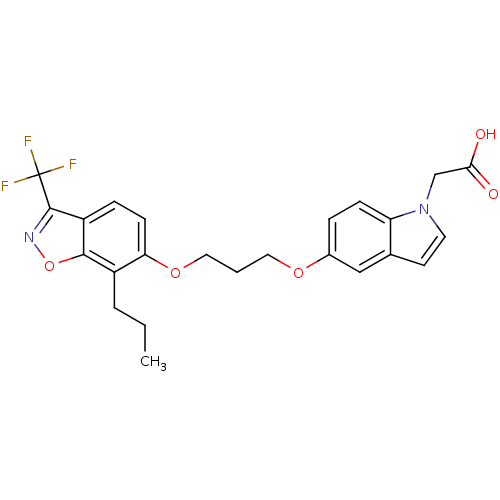
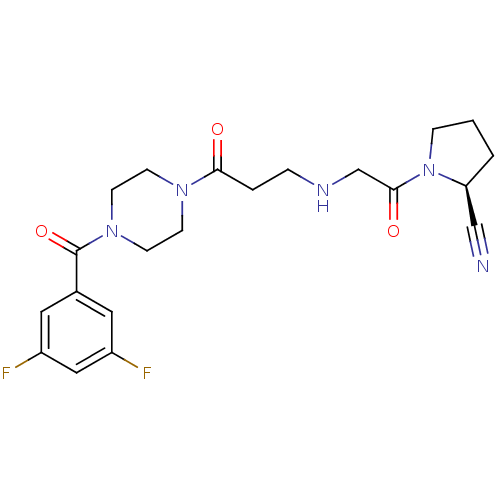
TargetSphingosine 1-phosphate receptor 1(Homo sapiens (Human))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
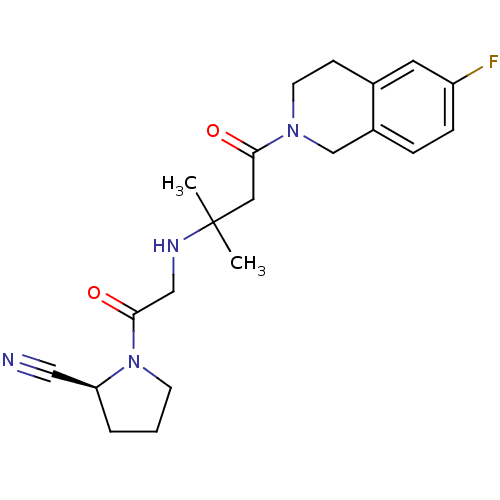
Affinity DataKi: 18nMAssay Description:Antagonist activity at human S1P1 receptor expressed in CHO cells assessed as inhibition of SEW2871-induced [35S]GTPgamma bindingMore data for this Ligand-Target Pair
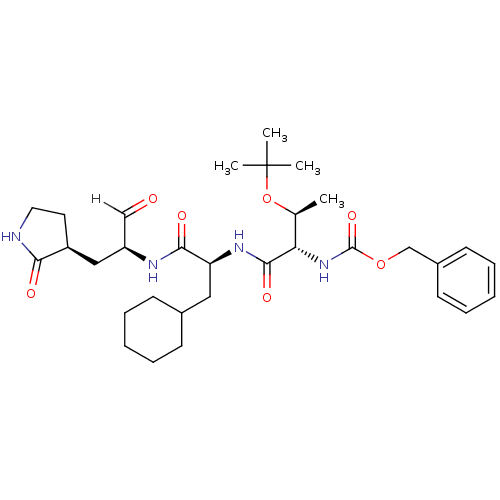
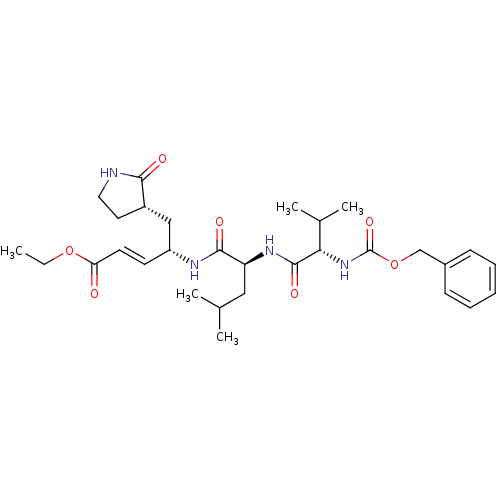
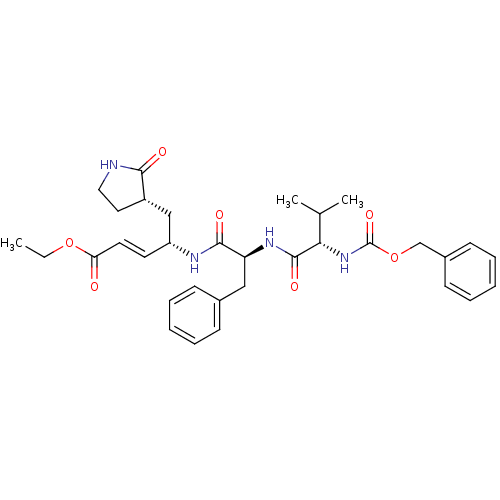
TargetReplicase polyprotein 1ab(Human SARS coronavirus (SARS-CoV) (Severe acute re...)
Taigen Biotechnology
Taigen Biotechnology
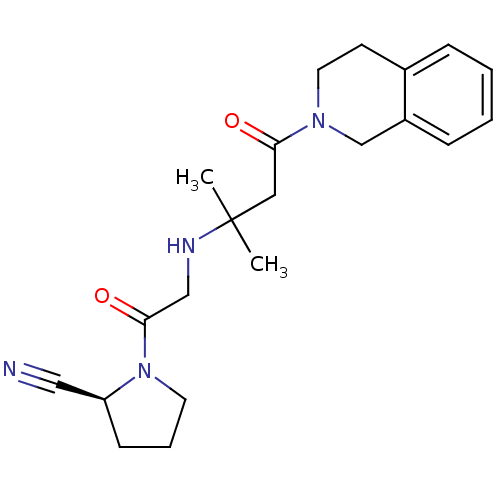
Affinity DataKi: 53nM ΔG°: -41.5kJ/molepH: 7.5 T: 2°CAssay Description:The effects of compound on enzyme activity were measured by using peptide cleavage assay. Cleavage products were resolved and analyzed with a reverse...More data for this Ligand-Target Pair
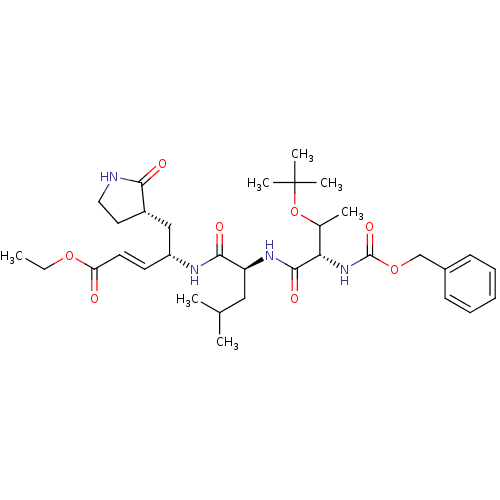
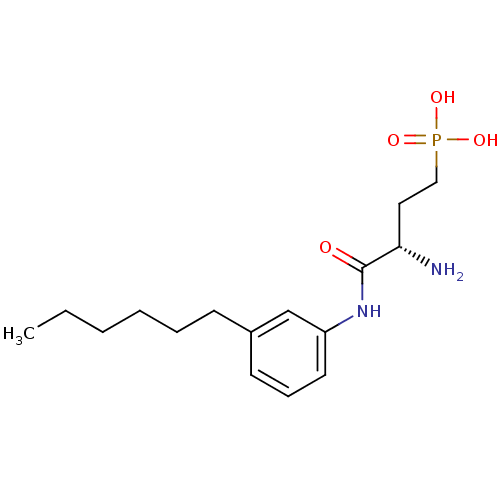
TargetReplicase polyprotein 1ab(Human SARS coronavirus (SARS-CoV) (Severe acute re...)
Taigen Biotechnology
Taigen Biotechnology
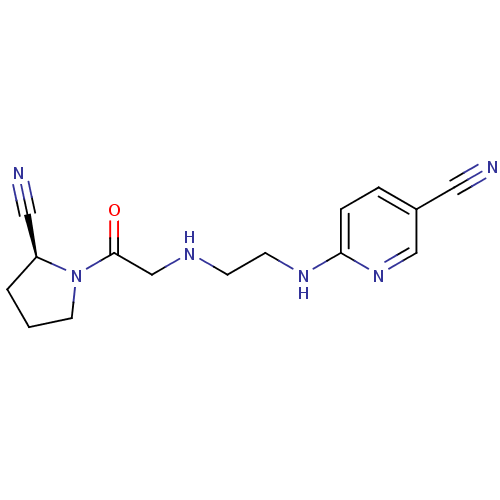
Affinity DataKi: 58nM ΔG°: -41.3kJ/molepH: 7.5 T: 2°CAssay Description:The effects of compound on enzyme activity were measured by using peptide cleavage assay. Cleavage products were resolved and analyzed with a reverse...More data for this Ligand-Target Pair
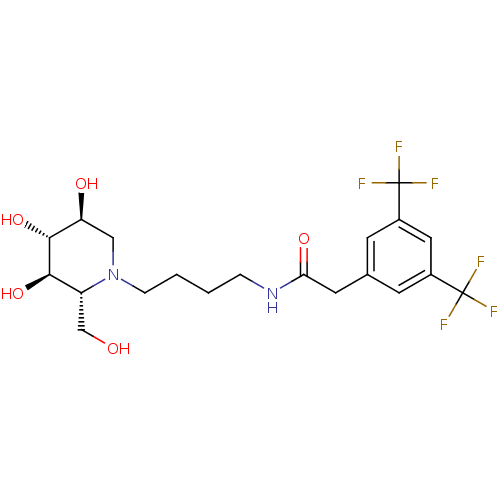
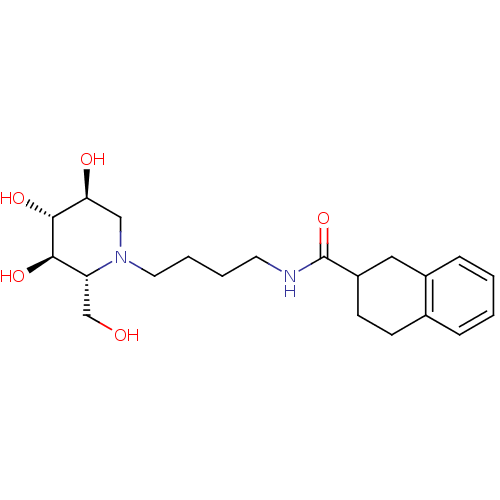
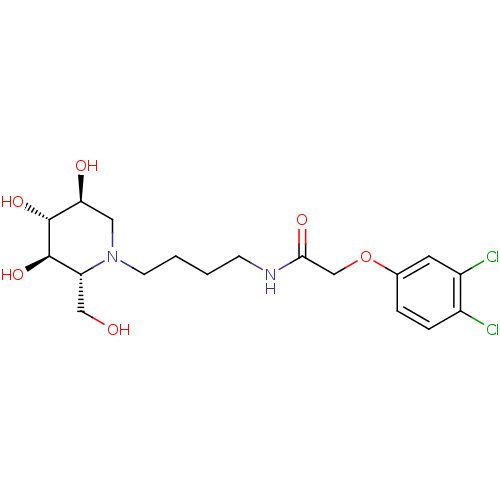
TargetLysosomal acid glucosylceramidase(Homo sapiens (Human))
Genomics Research Center
Curated by ChEMBL
Genomics Research Center
Curated by ChEMBL
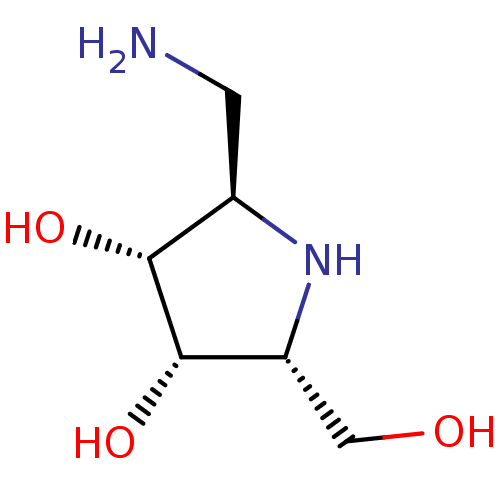
Affinity DataKi: 71nMAssay Description:Competitive inhibition of human beta-glucocerebrosidase by Lineweaver-Burk double reciprocal plot methodMore data for this Ligand-Target Pair
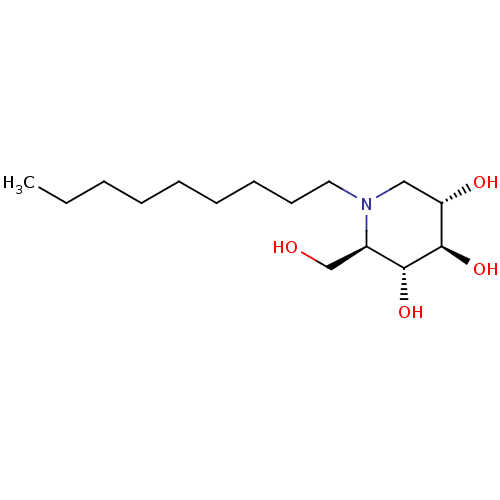
TargetSphingosine 1-phosphate receptor 1(Homo sapiens (Human))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 77nMAssay Description:Antagonist activity at human S1P1 receptor expressed in CHO cells assessed as inhibition of S1P-induced [35S]GTPgamma bindingMore data for this Ligand-Target Pair
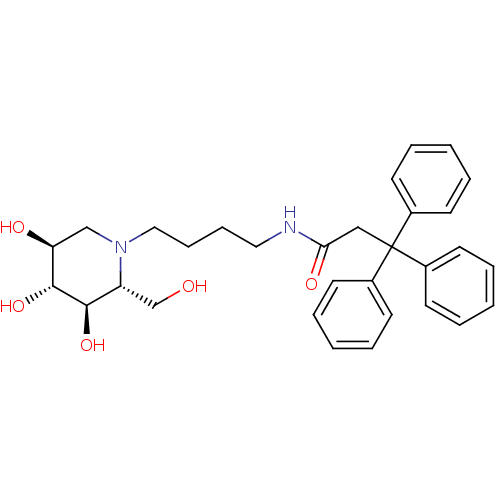
TargetLysosomal acid glucosylceramidase(Homo sapiens (Human))
Genomics Research Center
Curated by ChEMBL
Genomics Research Center
Curated by ChEMBL
Affinity DataKi: 400nMAssay Description:Competitive inhibition of human beta-glucocerebrosidase by Lineweaver-Burk double reciprocal plot methodMore data for this Ligand-Target Pair
TargetLysosomal acid glucosylceramidase(Homo sapiens (Human))
Genomics Research Center
Curated by ChEMBL
Genomics Research Center
Curated by ChEMBL
Affinity DataKi: 400nMAssay Description:Competitive inhibition of human beta-glucocerebrosidase by Lineweaver-Burk double reciprocal plot methodMore data for this Ligand-Target Pair
TargetReplicase polyprotein 1ab(Human SARS coronavirus (SARS-CoV) (Severe acute re...)
Taigen Biotechnology
Taigen Biotechnology
Affinity DataKi: 660nM ΔG°: -35.3kJ/molepH: 7.5 T: 2°CAssay Description:The effects of compound on enzyme activity were measured by using peptide cleavage assay. Cleavage products were resolved and analyzed with a reverse...More data for this Ligand-Target Pair
TargetLysosomal acid glucosylceramidase(Homo sapiens (Human))
Genomics Research Center
Curated by ChEMBL
Genomics Research Center
Curated by ChEMBL
Affinity DataKi: 1.00E+3nMAssay Description:Competitive inhibition of human beta-glucocerebrosidase by Lineweaver-Burk double reciprocal plot methodMore data for this Ligand-Target Pair
TargetLysosomal acid glucosylceramidase(Homo sapiens (Human))
Genomics Research Center
Curated by ChEMBL
Genomics Research Center
Curated by ChEMBL
Affinity DataKi: 1.10E+3nMAssay Description:Non-competitive inhibition of human beta-glucocerebrosidase by Lineweaver-Burk double reciprocal plot methodMore data for this Ligand-Target Pair
TargetLysosomal acid glucosylceramidase(Homo sapiens (Human))
Genomics Research Center
Curated by ChEMBL
Genomics Research Center
Curated by ChEMBL
Affinity DataKi: 1.30E+3nMAssay Description:Competitive inhibition of human beta-glucocerebrosidase by Lineweaver-Burk double reciprocal plot methodMore data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nMAssay Description:Competitive inhibition of recombinant human lysosomal alpha-glucosidase assessed as inhibition constant by Lineweaver-Burk plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nMAssay Description:Non-competitive inhibition of recombinant human alpha GAL-A using varying levels of 4-methylumbelliferyl alpha-D-galactopyranoside substrate at pH 7 ...More data for this Ligand-Target Pair
TargetReplicase polyprotein 1ab(Human SARS coronavirus (SARS-CoV) (Severe acute re...)
Taigen Biotechnology
Taigen Biotechnology
Affinity DataKi: 2.26E+3nM ΔG°: -32.2kJ/molepH: 7.5 T: 2°CAssay Description:The effects of compound on enzyme activity were measured by using peptide cleavage assay. Cleavage products were resolved and analyzed with a reverse...More data for this Ligand-Target Pair
TargetSphingosine 1-phosphate receptor 1(Homo sapiens (Human))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 2.84E+3nMAssay Description:Antagonist activity at human S1P1 receptor expressed in CHO cells assessed as inhibition of SEW2871-induced [35S]GTPgamma bindingMore data for this Ligand-Target Pair
TargetLysosomal acid glucosylceramidase(Homo sapiens (Human))
Genomics Research Center
Curated by ChEMBL
Genomics Research Center
Curated by ChEMBL
Affinity DataKi: 3.20E+3nMAssay Description:Competitive inhibition of human beta-glucocerebrosidase by Lineweaver-Burk double reciprocal plot methodMore data for this Ligand-Target Pair
Affinity DataKi: 3.50E+3nMAssay Description:Competitive inhibition of human recombinant alpha-galactosidase A using 4-MU-alpha-d-galactopyranoside as substrate by Lineweaver-Burk plot methodMore data for this Ligand-Target Pair
TargetLysosomal acid glucosylceramidase(Homo sapiens (Human))
Genomics Research Center
Curated by ChEMBL
Genomics Research Center
Curated by ChEMBL
Affinity DataKi: 4.10E+3nMAssay Description:Competitive inhibition of human beta-glucocerebrosidase by Lineweaver-Burk double reciprocal plot methodMore data for this Ligand-Target Pair
TargetSphingosine 1-phosphate receptor 1(Homo sapiens (Human))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 4.63E+3nMAssay Description:Antagonist activity at human S1P1 receptor expressed in CHO cells assessed as inhibition of S1P-induced [35S]GTPgamma bindingMore data for this Ligand-Target Pair
Affinity DataKi: 6.40E+3nMAssay Description:Competitive inhibition of recombinant human lysosomal alpha-glucosidase assessed as inhibition constant by Lineweaver-Burk plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 7.70E+3nMAssay Description:Non-competitive inhibition of recombinant human alpha GAL-A using varying levels of 4-methylumbelliferyl alpha-D-galactopyranoside substrate at pH 4....More data for this Ligand-Target Pair
TargetReplicase polyprotein 1ab(Human SARS coronavirus (SARS-CoV) (Severe acute re...)
Taigen Biotechnology
Taigen Biotechnology
Affinity DataKi: >1.00E+4nM ΔG°: >-28.5kJ/molepH: 7.5 T: 2°CAssay Description:The effects of compound on enzyme activity were measured by using peptide cleavage assay. Cleavage products were resolved and analyzed with a reverse...More data for this Ligand-Target Pair
Affinity DataKi: 1.15E+5nMAssay Description:Competitive inhibition of recombinant human lysosomal alpha-glucosidase assessed as inhibition constant by Lineweaver-Burk plot analysisMore data for this Ligand-Target Pair
TargetSerine/threonine-protein kinase pim-3(Homo sapiens (Human))
West Virginia University
Curated by ChEMBL
West Virginia University
Curated by ChEMBL
Affinity DataIC50: 0.121nMAssay Description:Inhibition of human PIM3 using RSRHSSYPAGT as substrate in presence of [gamma-33P]-ATPMore data for this Ligand-Target Pair
TargetDeath-associated protein kinase 1(Homo sapiens (Human))
West Virginia University
Curated by ChEMBL
West Virginia University
Curated by ChEMBL
Affinity DataIC50: 9.40nMAssay Description:Inhibition of human DAPK1 using KKLNRTLSFAEPG as substrate in presence of [gamma-33P]-ATPMore data for this Ligand-Target Pair
Affinity DataIC50: 12nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of recombinant human alpha GAL-A using 4-methylumbelliferyl alpha-D-galactopyranoside as substrate at pH 7 after 15 mins by fluorescence a...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 27nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 42nMAssay Description:Inhibition of recombinant human alpha GAL-A using 4-methylumbelliferyl alpha-D-galactopyranoside as substrate at pH 4.6 after 15 mins by fluorescence...More data for this Ligand-Target Pair
Affinity DataIC50: 49nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 51nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 53nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 53nMpH: 7.0Assay Description:Inhibition of human recombinant lysosomal alpha GAL-A using 4-methylberiilyl alpha-D-galactopyranoside as substrate at pH 7 after 15 mins by fluoresc...More data for this Ligand-Target Pair
Affinity DataIC50: 54nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 83nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Homo sapiens (Human))
National Health Research Institutes
Curated by ChEMBL
National Health Research Institutes
Curated by ChEMBL
Affinity DataIC50: 92nMAssay Description:Displacement of [3H]-rosiglitazone from human PPAR gamma by SPA assayMore data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Homo sapiens (Human))
National Health Research Institutes
Curated by ChEMBL
National Health Research Institutes
Curated by ChEMBL
Affinity DataIC50: 105nMAssay Description:Displacement of [3H]-rosiglitazone from human PPAR gamma by SPA assayMore data for this Ligand-Target Pair
Affinity DataIC50: 116nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 119nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Homo sapiens (Human))
National Health Research Institutes
Curated by ChEMBL
National Health Research Institutes
Curated by ChEMBL
Affinity DataIC50: 120nMAssay Description:Displacement of [3H]-rosiglitazone from human PPAR gamma by SPA assayMore data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Homo sapiens (Human))
National Health Research Institutes
Curated by ChEMBL
National Health Research Institutes
Curated by ChEMBL
Affinity DataIC50: 127nMAssay Description:Displacement of [3H]-rosiglitazone from human PPAR gamma by SPA assayMore data for this Ligand-Target Pair
Affinity DataIC50: 132nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Homo sapiens (Human))
National Health Research Institutes
Curated by ChEMBL
National Health Research Institutes
Curated by ChEMBL
Affinity DataIC50: 152nMAssay Description:Displacement of [3H]-rosiglitazone from human PPAR gamma by SPA assayMore data for this Ligand-Target Pair
Affinity DataIC50: 202nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 298nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 317nMpH: 8.0 T: 2°CAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair
Affinity DataIC50: 369nMAssay Description:The enzyme activity resulted in the liberation of free pNA at 405 nm. Reaction progress was monitored using a Molecular Devices SpectraMax Plus micro...More data for this Ligand-Target Pair

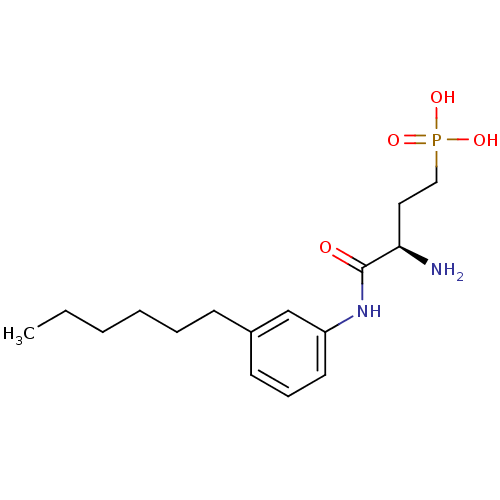
 3D Structure (crystal)
3D Structure (crystal)