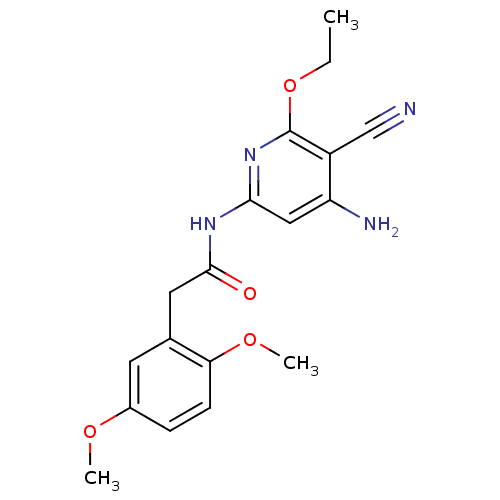
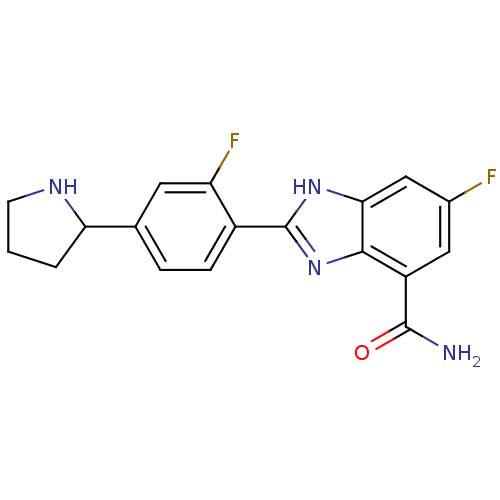
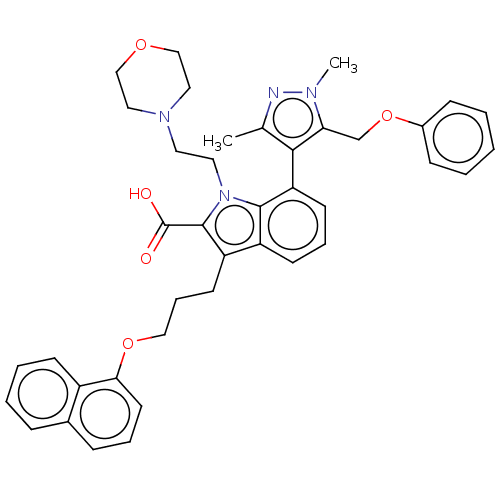
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Abbvie
Curated by ChEMBL
Abbvie
Curated by ChEMBL
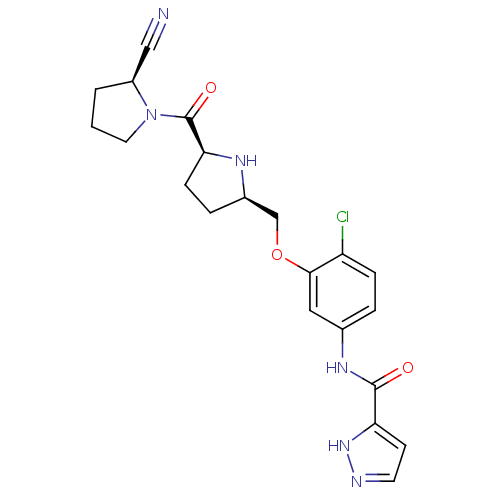
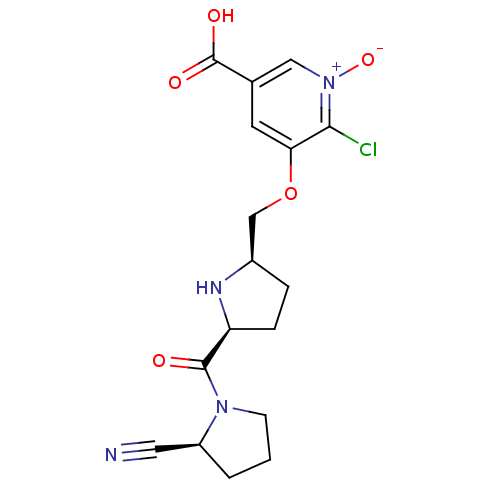
Affinity DataKi: 0.200nMAssay Description:Displacement of F-Bak (GQVGRQLAIIGDK(6-FAM)INR-amide) from MCL1 (unknown origin) after 1 hr by TR-FRET assayMore data for this Ligand-Target Pair
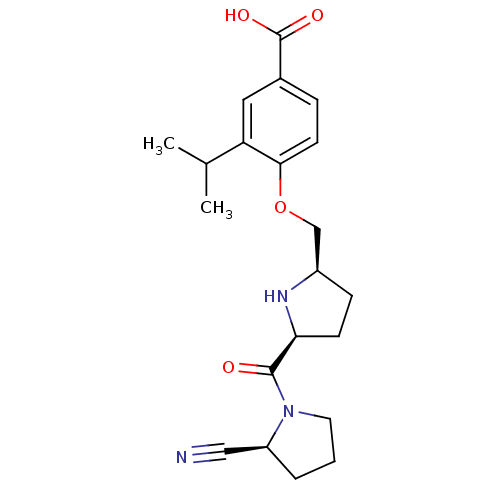
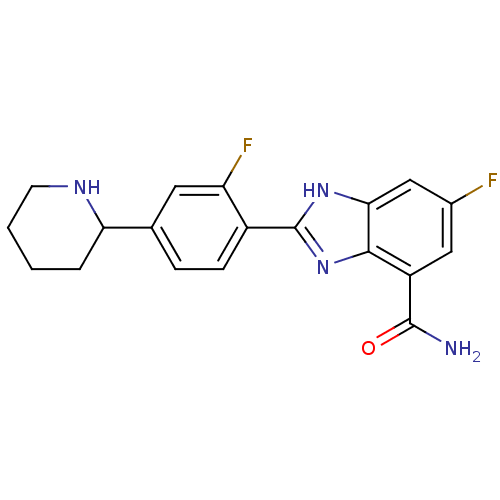
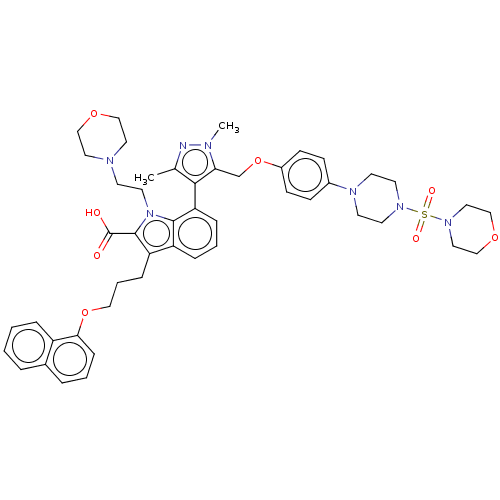
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Abbvie
Curated by ChEMBL
Abbvie
Curated by ChEMBL
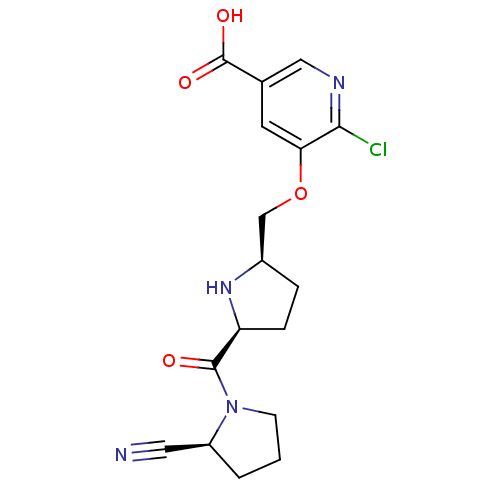
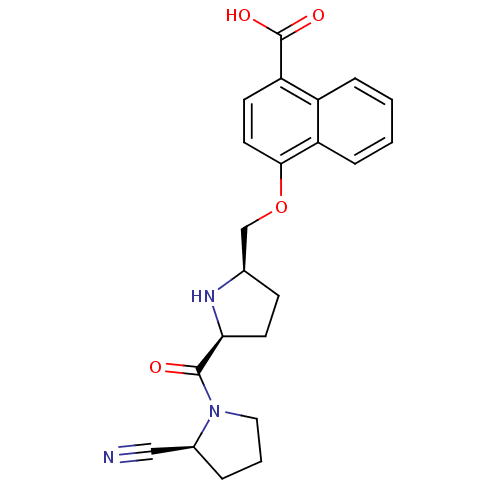
Affinity DataKi: 0.240nMAssay Description:Displacement of F-Bak (GQVGRQLAIIGDK(6-FAM)INR-amide) from MCL1 (unknown origin) after 1 hr by TR-FRET assayMore data for this Ligand-Target Pair
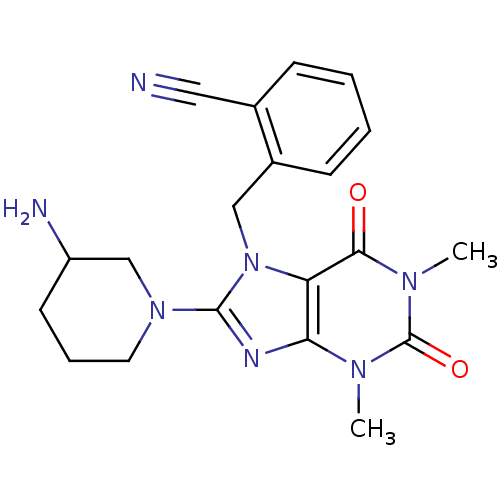
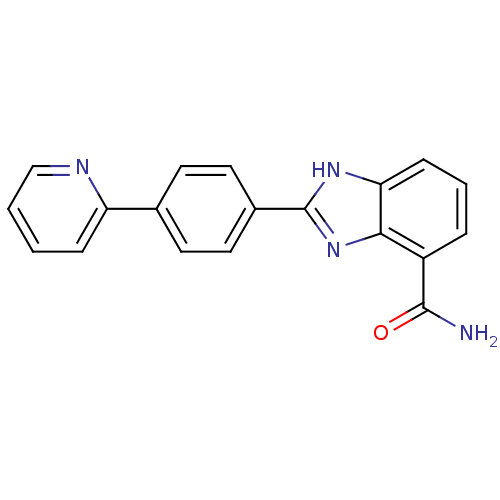
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Abbvie
Curated by ChEMBL
Abbvie
Curated by ChEMBL
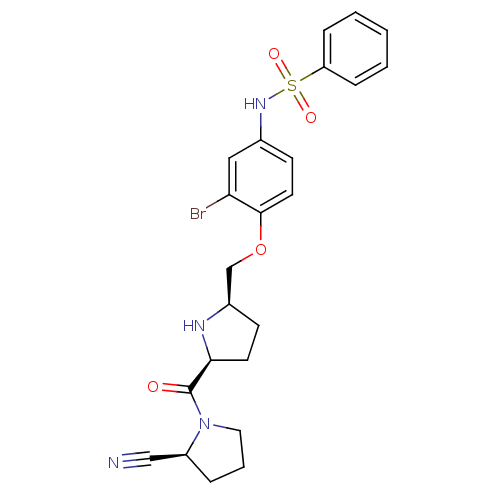
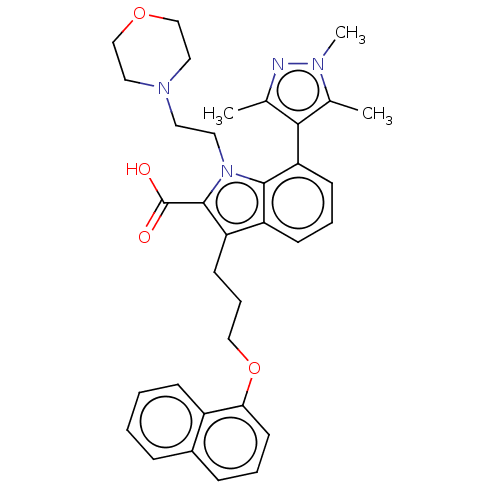
Affinity DataKi: 0.310nMAssay Description:Displacement of F-Bak (GQVGRQLAIIGDK(6-FAM)INR-amide) from MCL1 (unknown origin) after 1 hr by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.410nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
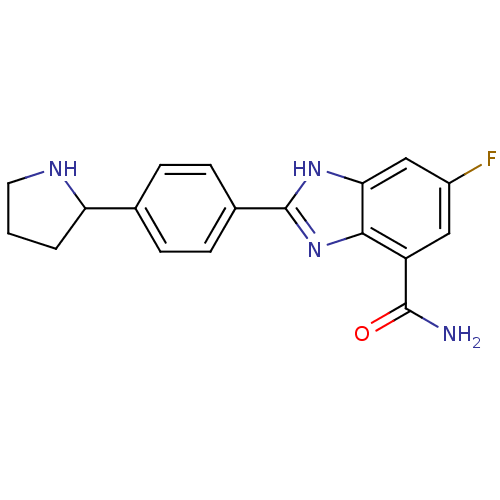
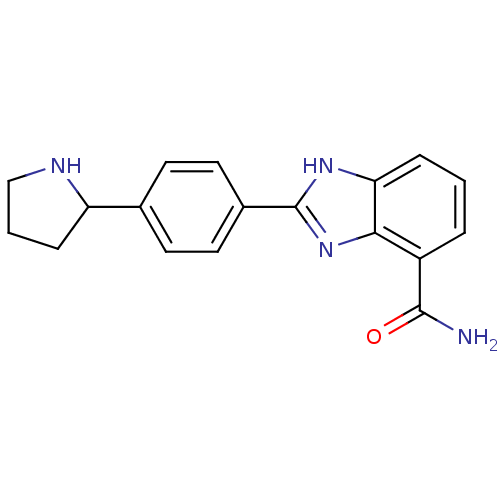
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Abbvie
Curated by ChEMBL
Abbvie
Curated by ChEMBL
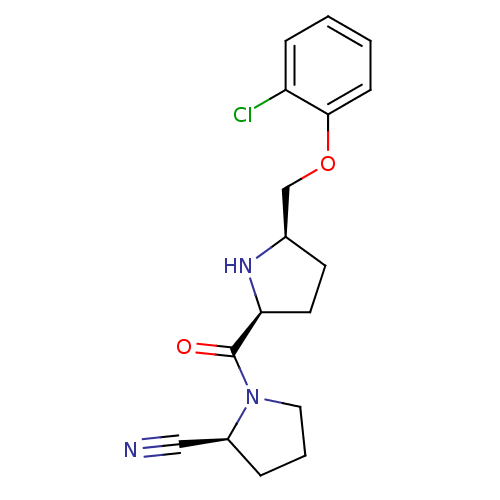
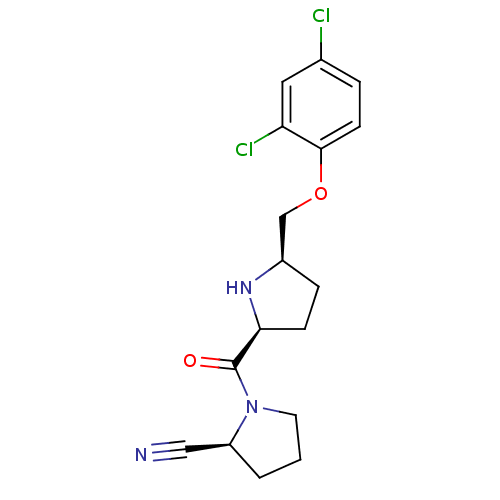
Affinity DataKi: 0.430nMAssay Description:Displacement of F-Bak (GQVGRQLAIIGDK(6-FAM)INR-amide) from MCL1 (unknown origin) after 1 hr by TR-FRET assayMore data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Abbvie
Curated by ChEMBL
Abbvie
Curated by ChEMBL
Affinity DataKi: 0.450nMAssay Description:Displacement of F-Bak (GQVGRQLAIIGDK(6-FAM)INR-amide) from MCL1 (unknown origin) after 1 hr by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.450nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Abbvie
Curated by ChEMBL
Abbvie
Curated by ChEMBL
Affinity DataKi: 0.480nMAssay Description:Displacement of F-Bak (GQVGRQLAIIGDK(6-FAM)INR-amide) from MCL1 (unknown origin) after 1 hr by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.600nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 0.630nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 0.660nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 0.660nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 0.680nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 0.680nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 0.720nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 0.730nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 0.820nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 0.850nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 0.930nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inhibition of PARP2 by top count scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inhibition of PARP2 by top count scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inhibition of PARP1 using [3H]NAD+ by top count scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inhibition of PARP1 using [3H]NAD+ by top count scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Ser/Thr-kinase selectivity assays were performed using a radioactive FlashPlate-based assay platform. Substrate incorporated radioactivity was counte...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Inhibition of PARP2 by top count scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Abbvie
Curated by ChEMBL
Abbvie
Curated by ChEMBL
Affinity DataKi: 1.5nMAssay Description:Displacement of F-Bak (GQVGRQLAIIGDK(6-FAM)INR-amide) from MCL1 (unknown origin) after 1 hr by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Abbvie
Curated by ChEMBL
Abbvie
Curated by ChEMBL
Affinity DataKi: 1.80nMAssay Description:Displacement of F-Bak (GQVGRQLAIIGDK(6-FAM)INR-amide) from MCL1 (unknown origin) after 1 hr by TR-FRET assayMore data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Abbvie
Curated by ChEMBL
Abbvie
Curated by ChEMBL
Affinity DataKi: 2nMAssay Description:Displacement of F-Bak (GQVGRQLAIIGDK(6-FAM)INR-amide) from MCL1 (unknown origin) after 1 hr by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Ser/Thr-kinase selectivity assays were performed using a radioactive FlashPlate-based assay platform. Substrate incorporated radioactivity was counte...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -49.2kJ/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of PARP1 using [3H]NAD+ by top count scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of PARP1 using [3H]NAD+ by top count scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of PARP1 using [3H]NAD+ by top count scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of PARP1 using [3H]NAD+ by top count scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of PARP1 using [3H]NAD+ by top count scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of PARP2 by top count scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of PARP2 by top count scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of PARP1 using [3H]NAD+ by top count scintillation countingMore data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Abbvie
Curated by ChEMBL
Abbvie
Curated by ChEMBL
Affinity DataKi: 2.30nMAssay Description:Displacement of F-Bak (GQVGRQLAIIGDK(6-FAM)INR-amide) from MCL1 (unknown origin) after 1 hr by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:The DPP4 activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and mea...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Abbvie
Curated by ChEMBL
Abbvie
Curated by ChEMBL
Affinity DataKi: 2.70nMAssay Description:Displacement of F-Bak (GQVGRQLAIIGDK(6-FAM)INR-amide) from MCL1 (unknown origin) after 1 hr by TR-FRET assay in presence of 10% human serumMore data for this Ligand-Target Pair
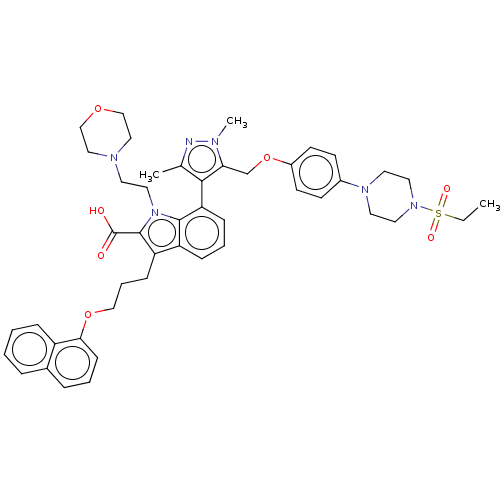













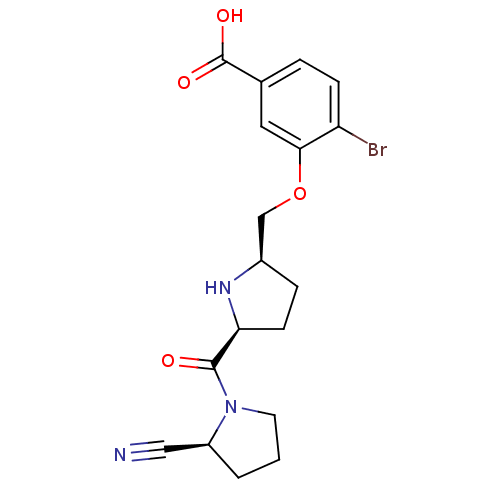

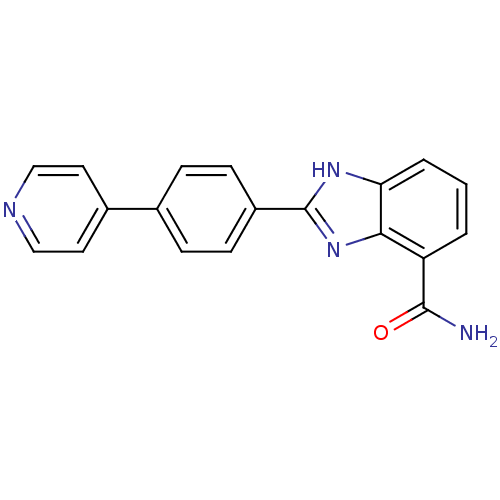

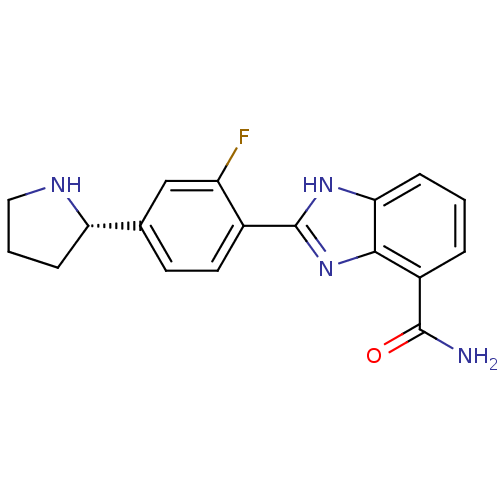

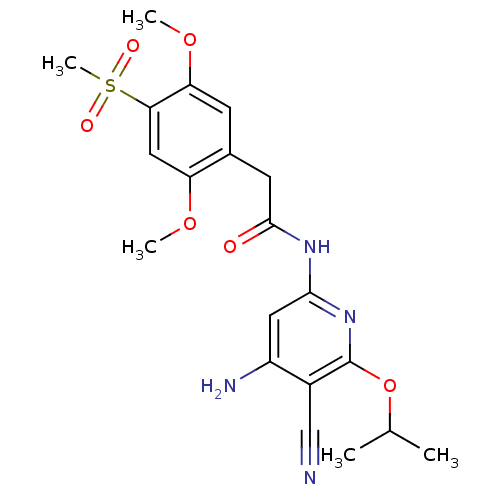


 3D Structure (crystal)
3D Structure (crystal)