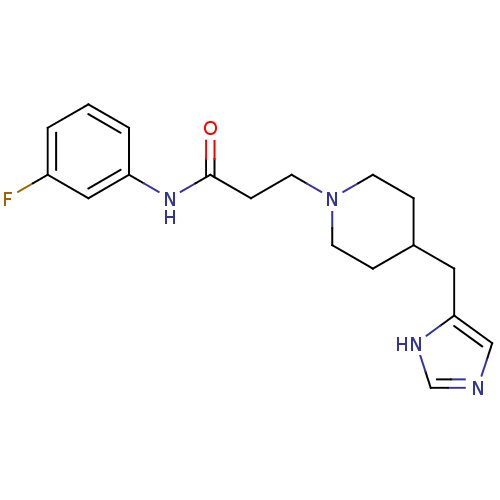
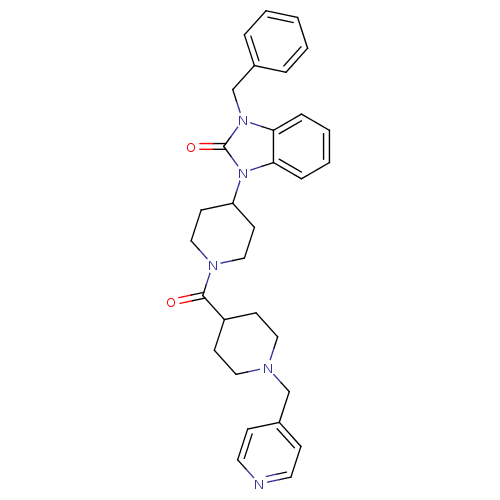
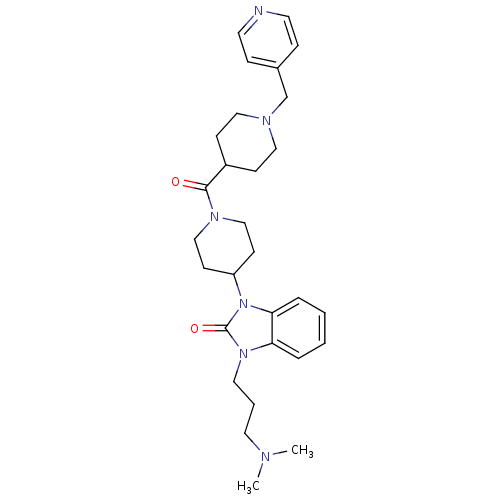
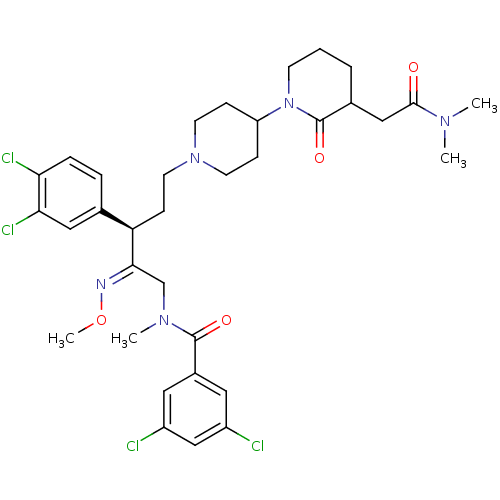
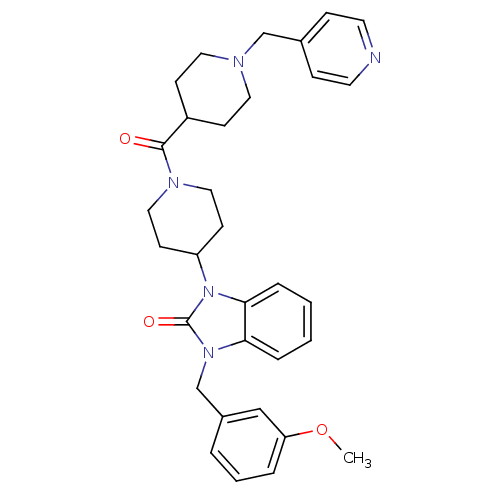
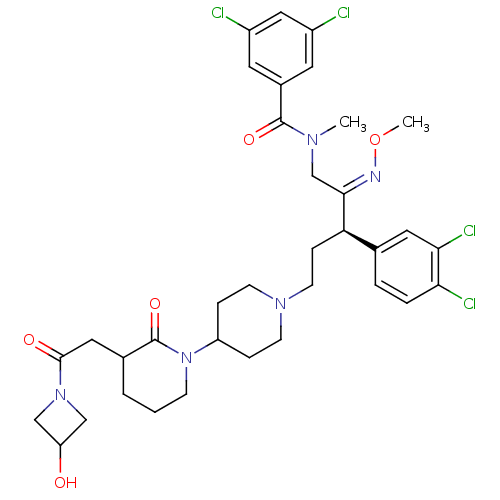
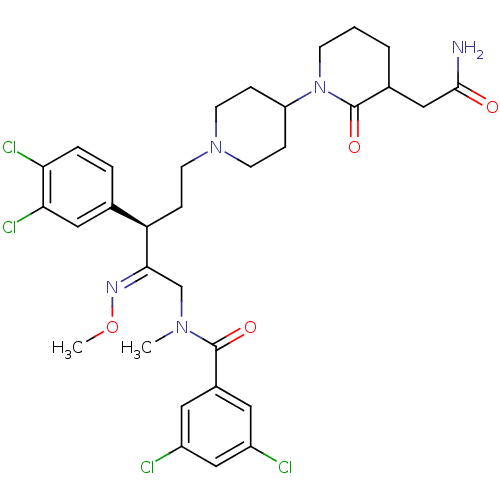
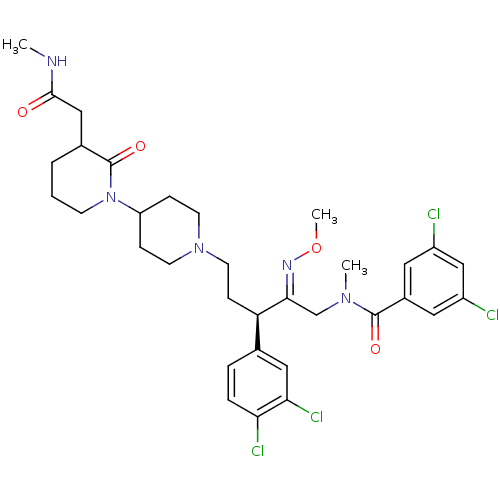
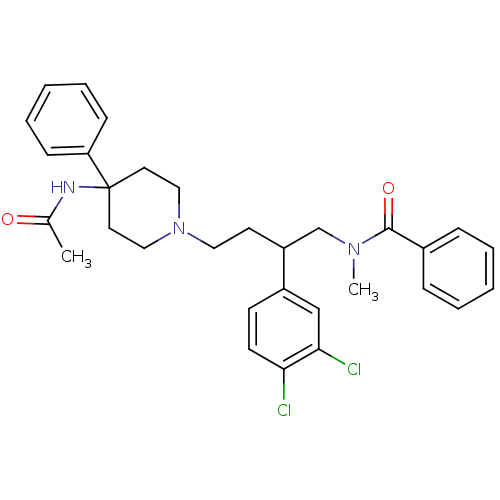
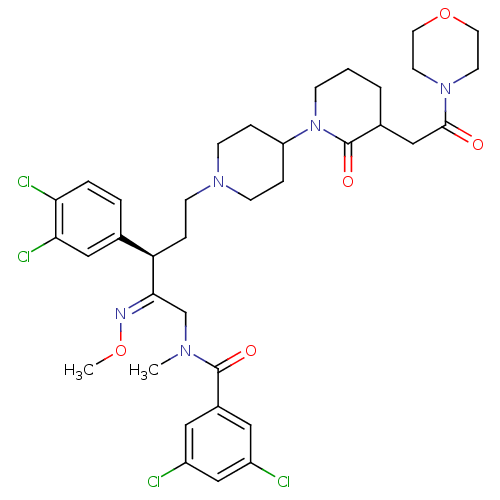
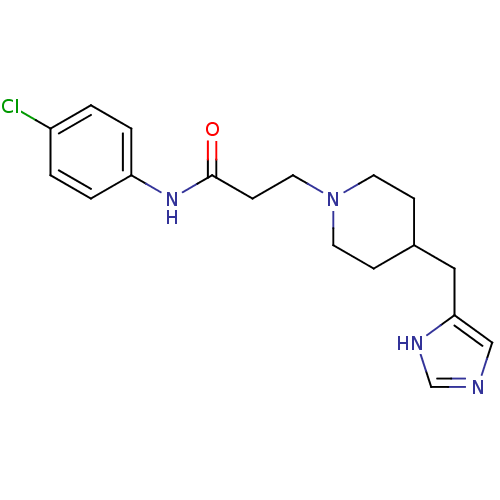
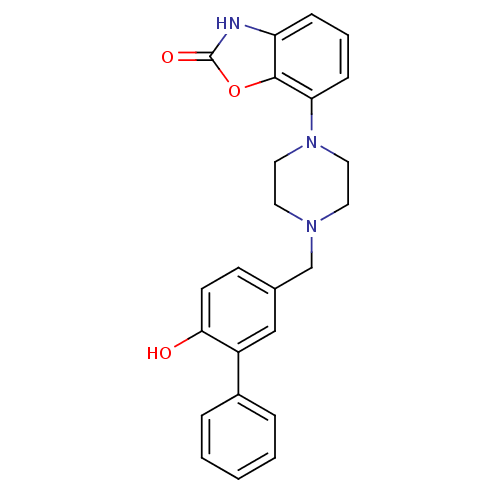
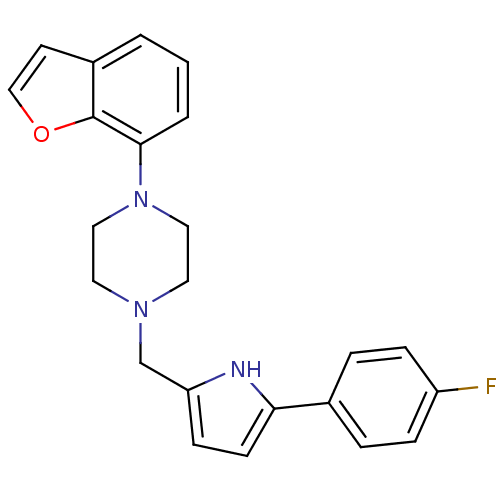
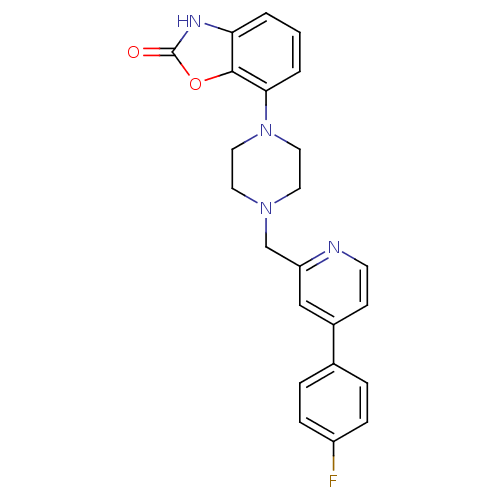
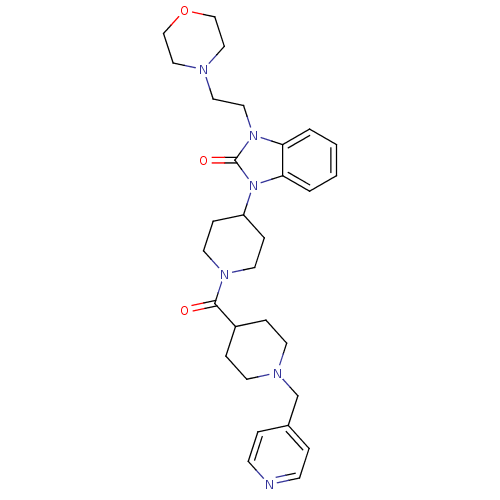
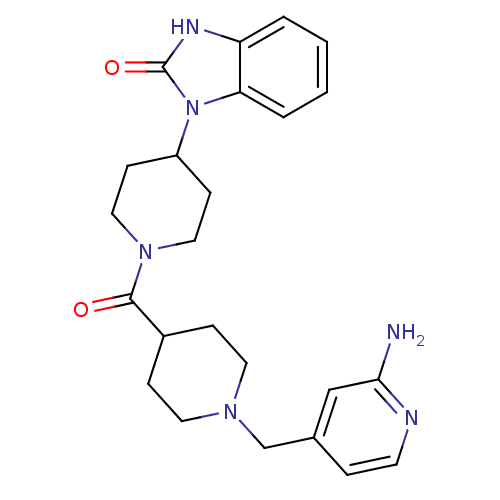
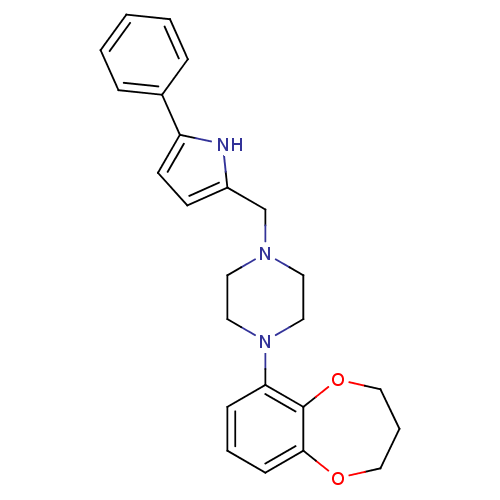
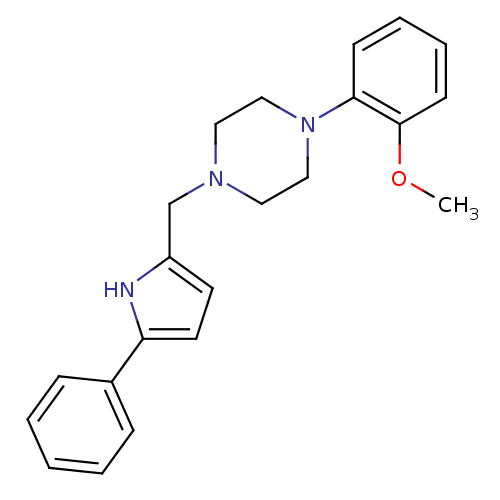
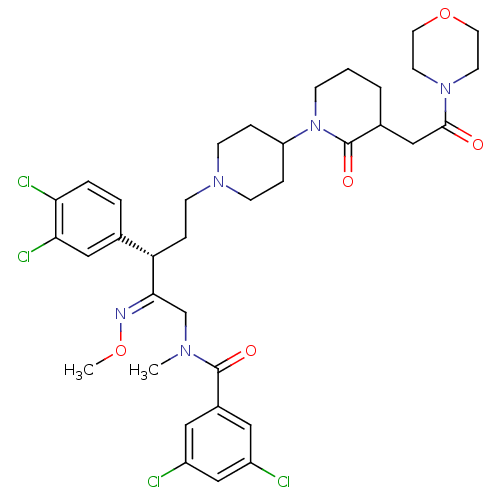
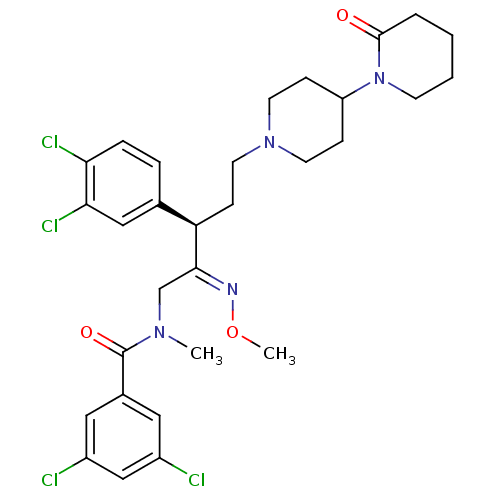
Affinity DataKi: 0.150nMAssay Description:Displacement of [3H]Nalpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]Nalpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]Nalpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]Nalpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:In vitro binding to Tachykinin receptor 2More data for this Ligand-Target Pair
Affinity DataKi: 0.25nMAssay Description:Displacement of [3H]Nalpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:In vitro binding to Tachykinin receptor 2More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:In vitro binding to Tachykinin receptor 1More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Displacement of [3H]Nalpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:In vitro binding to Tachykinin receptor 2More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:In vitro binding to Tachykinin receptor 2More data for this Ligand-Target Pair
TargetNeuromedin-K receptor(Homo sapiens (Human))
Schering-Plough Research Institute
Curated by PDSP Ki Database
Schering-Plough Research Institute
Curated by PDSP Ki Database
TargetSubstance-K receptor(Homo sapiens (Human))
Schering-Plough Research Institute
Curated by PDSP Ki Database
Schering-Plough Research Institute
Curated by PDSP Ki Database
TargetSubstance-K receptor(Homo sapiens (Human))
Schering-Plough Research Institute
Curated by PDSP Ki Database
Schering-Plough Research Institute
Curated by PDSP Ki Database
Affinity DataKi: 0.400nMAssay Description:In vitro binding to Tachykinin receptor 2More data for this Ligand-Target Pair
Affinity DataKi: 0.400nMAssay Description:In vitro binding to Tachykinin receptor 2More data for this Ligand-Target Pair
Affinity DataKi: 0.400nMAssay Description:In vitro binding to Tachykinin receptor 1More data for this Ligand-Target Pair
Affinity DataKi: 0.400nMAssay Description:Displacement of [3H]Nalpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.400nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
TargetHistamine H3 receptor(Homo sapiens (Human))
The Schering Plough Research Institute
Curated by ChEMBL
The Schering Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 0.400nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from histamine H3 receptor in human brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.400nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.400nMAssay Description:In vitro binding affinity for serotonin reuptake sites in rat frontal cortex membranes by [3H]-paroxetine displacement.More data for this Ligand-Target Pair
Affinity DataKi: 0.410nMAssay Description:Binding affinity for dopamine receptor D2 determined using [3H]-spiperoneMore data for this Ligand-Target Pair
Affinity DataKi: 0.420nMAssay Description:Binding affinity for dopamine receptor D2 determined using [3H]-spiperoneMore data for this Ligand-Target Pair
Affinity DataKi: 0.450nMAssay Description:Displacement of [3H]-Spiperone from Dopamine receptor D2 in rat striatumMore data for this Ligand-Target Pair
Affinity DataKi: 0.470nMAssay Description:Binding affinity for dopamine receptor D2 determined using [3H]-spiperoneMore data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Displacement of [3H]-N-alpha-methyl histamine from histamine H3 receptor in guinea pig brain homogenatesMore data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Displacement of [3H]Nalpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.600nMAssay Description:In vitro binding to Tachykinin receptor 1More data for this Ligand-Target Pair
Affinity DataKi: 0.600nMAssay Description:In vitro binding to Tachykinin receptor 2More data for this Ligand-Target Pair
Affinity DataKi: 0.600nMAssay Description:In vitro binding to Tachykinin receptor 1More data for this Ligand-Target Pair
Affinity DataKi: 0.600nMAssay Description:In vitro binding affinity for serotonin reuptake sites in rat frontal cortex membranes by [3H]-paroxetine displacement.More data for this Ligand-Target Pair
Affinity DataKi: 0.600nMAssay Description:Displacement of [3H]-N-alpha-methyl histamine from histamine H3 receptor in guinea pig brain homogenatesMore data for this Ligand-Target Pair
Affinity DataKi: 0.650nMAssay Description:Displacement of [3H]Nalpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.680nMAssay Description:Displacement of [3H]-Spiperone from Dopamine receptor D2 in rat striatumMore data for this Ligand-Target Pair
Affinity DataKi: 0.700nMAssay Description:In vitro binding to Tachykinin receptor 1More data for this Ligand-Target Pair
Affinity DataKi: 0.700nMAssay Description:Displacement of [3H]-N-alpha-methyl histamine from histamine H3 receptor in guinea pig brain homogenatesMore data for this Ligand-Target Pair
Affinity DataKi: 0.700nMAssay Description:In vitro binding to Tachykinin receptor 2More data for this Ligand-Target Pair
Affinity DataKi: 0.760nMAssay Description:Displacement of [3H]-Spiperone from Dopamine receptor D2 in rat striatumMore data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:In vitro binding to Tachykinin receptor 1More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:In vitro binding to Tachykinin receptor 1More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:In vitro binding to Tachykinin receptor 2More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:In vitro binding to Tachykinin receptor 1More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:In vitro binding to Tachykinin receptor 2More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Displacement of [3H]Nalpha-methylhistamine from histamine H3 receptor in guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:In vitro binding affinity for serotonin reuptake sites in rat frontal cortex membranes by [3H]-paroxetine displacement.More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:In vitro binding to Tachykinin receptor 2More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:In vitro binding to Tachykinin receptor 1More data for this Ligand-Target Pair