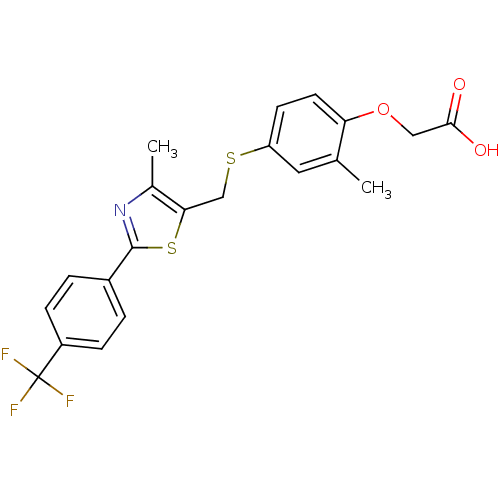
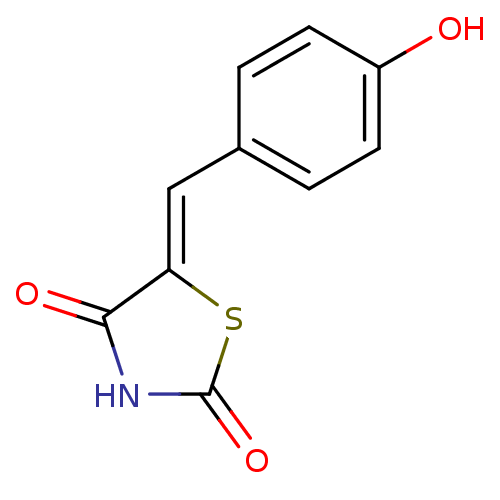
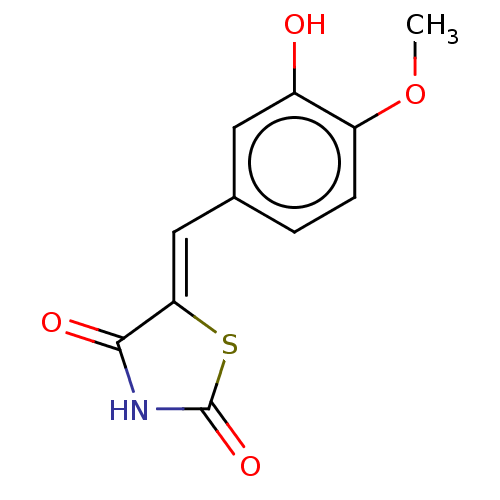
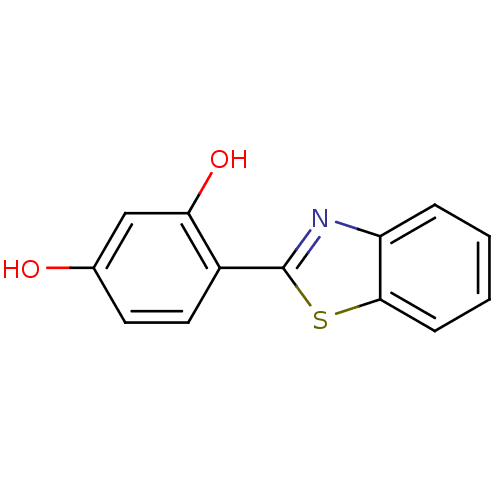
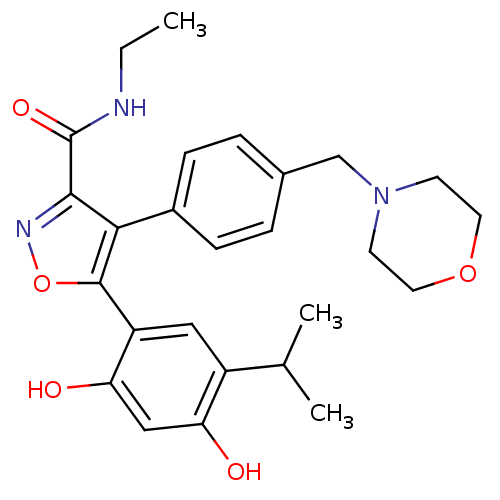
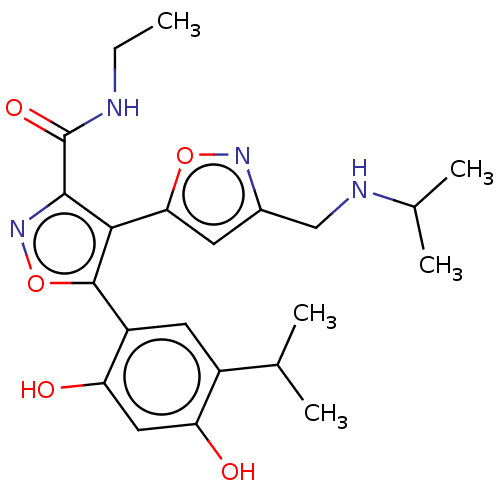
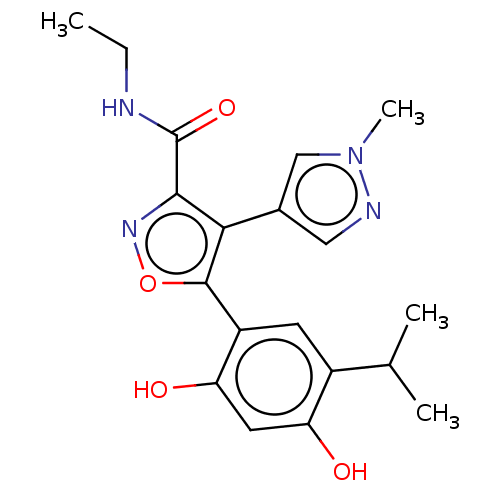
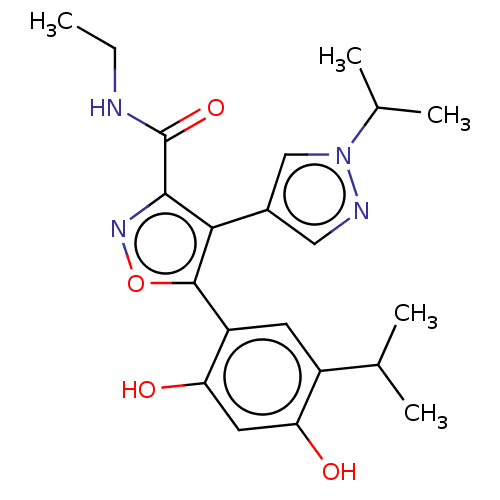
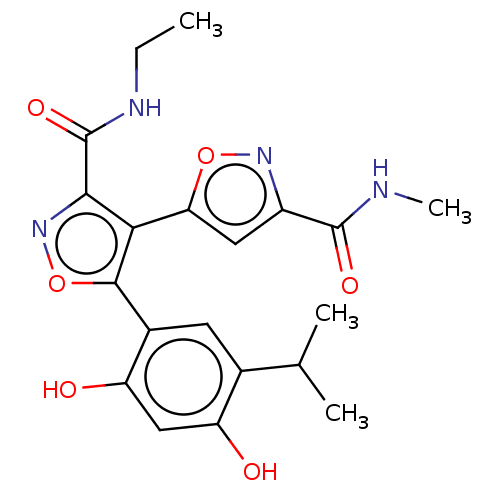
TargetPeroxisome proliferator-activated receptor alpha(Homo sapiens (Human))
Chonnam National University
Curated by ChEMBL
Chonnam National University
Curated by ChEMBL
Affinity DataKi: 10nMAssay Description:Binding affinity to PPARalpha (unknown origin) by TR-FRET based LanthaScreen competitive binding assayMore data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Homo sapiens (Human))
Chonnam National University
Curated by ChEMBL
Chonnam National University
Curated by ChEMBL
Affinity DataKi: 70nMAssay Description:Binding affinity to PPARgamma (unknown origin) by TR-FRET based LanthaScreen competitive binding assayMore data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor delta(Homo sapiens (Human))
Chonnam National University
Curated by ChEMBL
Chonnam National University
Curated by ChEMBL
Affinity DataKi: 80nMAssay Description:Binding affinity to PPARdelta (unknown origin) by TR-FRET based LanthaScreen competitive binding assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
Pusan National University
Curated by ChEMBL
Pusan National University
Curated by ChEMBL
Affinity DataKi: 374nMAssay Description:Competitive inhibition of mushroom tyrosinase using L-tyrosine as substrate at 1.25 uM by Line-Weaver-Burk plot analysisMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
Pusan National University
Curated by ChEMBL
Pusan National University
Curated by ChEMBL
Affinity DataKi: 413nMAssay Description:Competitive inhibition of mushroom tyrosinase using L-tyrosine as substrate at 1.25 uM by Line-Weaver-Burk plot analysisMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
Pusan National University
Curated by ChEMBL
Pusan National University
Curated by ChEMBL
Affinity DataKi: 567nMAssay Description:Inhibition of mushroom tyrosinase at 1.25 uM by Lineweaver-burk plot analysisMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
Pusan National University
Curated by ChEMBL
Pusan National University
Curated by ChEMBL
Affinity DataKi: 673nMAssay Description:Inhibition of mushroom tyrosinase at 1.25 uM by Lineweaver-burk plot analysisMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
Pusan National University
Curated by ChEMBL
Pusan National University
Curated by ChEMBL
Affinity DataKi: 840nMAssay Description:Competitive inhibition of mushroom tyrosinase using L-tyrosine as substrate at 20 uM by Line-Weaver-Burk plot analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 1(Homo sapiens (Human))
Kyungpook National University
Curated by ChEMBL
Kyungpook National University
Curated by ChEMBL
Affinity DataKi: 920nMAssay Description:Inhibition of PTP1B (unknown origin) assessed as decrease in p-nitrophenolate formation using using pNPP as substrate by Dixon plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 990nMAssay Description:Inhibition of BChE (unknown origin) by Dixon plot analysisMore data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor delta(Homo sapiens (Human))
Chonnam National University
Curated by ChEMBL
Chonnam National University
Curated by ChEMBL
Affinity DataKi: 1.00E+3nMAssay Description:Binding affinity to PPARdelta (unknown origin) by TR-FRET based LanthaScreen competitive binding assayMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 1(Homo sapiens (Human))
Kyungpook National University
Curated by ChEMBL
Kyungpook National University
Curated by ChEMBL
Affinity DataKi: 1.02E+3nMAssay Description:Inhibition of PTP1B (unknown origin) assessed as decrease in p-nitrophenolate formation using using pNPP as substrate by Dixon plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 1.17E+3nMAssay Description:Inhibition of AChE (unknown origin) by Dixon plot analysisMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
Pusan National University
Curated by ChEMBL
Pusan National University
Curated by ChEMBL
Affinity DataKi: 1.38E+3nMAssay Description:Competitive inhibition of mushroom tyrosinase using L-tyrosine as substrate at 20 uM by Line-Weaver-Burk plot analysisMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
Pusan National University
Curated by ChEMBL
Pusan National University
Curated by ChEMBL
Affinity DataKi: 1.70E+3nMAssay Description:Inhibition of mushroom tyrosinase at 20 uM by Lineweaver-burk plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 2.49E+3nMAssay Description:Inhibition of AChE (unknown origin) by Dixon plot analysisMore data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Homo sapiens (Human))
Chonnam National University
Curated by ChEMBL
Chonnam National University
Curated by ChEMBL
Affinity DataKi: 2.70E+3nMAssay Description:Binding affinity to PPARgamma (unknown origin) by TR-FRET based LanthaScreen competitive binding assayMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 1(Homo sapiens (Human))
Kyungpook National University
Curated by ChEMBL
Kyungpook National University
Curated by ChEMBL
Affinity DataKi: 3.00E+3nM ΔG°: -32.8kJ/mole IC50: 4.80E+3nMpH: 6.0 T: 2°CAssay Description:In each 96-well plates (total 200 μL of volume), there were 2 mM p-NPP and PTP1B (0.05-0.1 μg) in a buffer containing 50 mM citrate (pH 6.0...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor delta(Homo sapiens (Human))
Chonnam National University
Curated by ChEMBL
Chonnam National University
Curated by ChEMBL
Affinity DataKi: 3.40E+3nMAssay Description:Binding affinity to PPARdelta (unknown origin) by TR-FRET based LanthaScreen competitive binding assayMore data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor delta(Homo sapiens (Human))
Chonnam National University
Curated by ChEMBL
Chonnam National University
Curated by ChEMBL
Affinity DataKi: 4.30E+3nMAssay Description:Binding affinity to PPARdelta (unknown origin) by TR-FRET based LanthaScreen competitive binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.78E+3nMAssay Description:Inhibition of BChE (unknown origin) by Dixon plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 4.88E+3nMAssay Description:Inhibition of BChE (unknown origin) by Dixon plot analysisMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
Pusan National University
Curated by ChEMBL
Pusan National University
Curated by ChEMBL
Affinity DataKi: 7.70E+3nMAssay Description:Inhibition of mushroom tyrosinase at 20 uM by Lineweaver-burk plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 7.97E+3nMAssay Description:Inhibition of AChE (unknown origin) by Dixon plot analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 1(Homo sapiens (Human))
Kyungpook National University
Curated by ChEMBL
Kyungpook National University
Curated by ChEMBL
Affinity DataKi: 9.70E+3nM ΔG°: -29.8kJ/mole IC50: 1.38E+4nMpH: 6.0 T: 2°CAssay Description:In each 96-well plates (total 200 μL of volume), there were 2 mM p-NPP and PTP1B (0.05-0.1 μg) in a buffer containing 50 mM citrate (pH 6.0...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 1(Homo sapiens (Human))
Kyungpook National University
Curated by ChEMBL
Kyungpook National University
Curated by ChEMBL
Affinity DataKi: 1.11E+4nM ΔG°: -29.4kJ/mole IC50: 1.46E+4nMpH: 6.0 T: 2°CAssay Description:In each 96-well plates (total 200 μL of volume), there were 2 mM p-NPP and PTP1B (0.05-0.1 μg) in a buffer containing 50 mM citrate (pH 6.0...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 1(Homo sapiens (Human))
Kyungpook National University
Curated by ChEMBL
Kyungpook National University
Curated by ChEMBL
Affinity DataKi: 1.13E+4nM ΔG°: -29.4kJ/mole IC50: 1.45E+4nMpH: 6.0 T: 2°CAssay Description:In each 96-well plates (total 200 μL of volume), there were 2 mM p-NPP and PTP1B (0.05-0.1 μg) in a buffer containing 50 mM citrate (pH 6.0...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Homo sapiens (Human))
Chonnam National University
Curated by ChEMBL
Chonnam National University
Curated by ChEMBL
Affinity DataKi: 1.19E+4nMAssay Description:Binding affinity to PPARgamma (unknown origin) by TR-FRET based LanthaScreen competitive binding assayMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 1(Homo sapiens (Human))
Kyungpook National University
Curated by ChEMBL
Kyungpook National University
Curated by ChEMBL
Affinity DataKi: 1.39E+4nM ΔG°: -28.8kJ/mole IC50: 1.59E+4nMpH: 6.0 T: 2°CAssay Description:In each 96-well plates (total 200 μL of volume), there were 2 mM p-NPP and PTP1B (0.05-0.1 μg) in a buffer containing 50 mM citrate (pH 6.0...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 1(Homo sapiens (Human))
Kyungpook National University
Curated by ChEMBL
Kyungpook National University
Curated by ChEMBL
Affinity DataKi: 1.45E+4nM ΔG°: -28.7kJ/mole IC50: 1.32E+4nMpH: 6.0 T: 2°CAssay Description:In each 96-well plates (total 200 μL of volume), there were 2 mM p-NPP and PTP1B (0.05-0.1 μg) in a buffer containing 50 mM citrate (pH 6.0...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
Pusan National University
Curated by ChEMBL
Pusan National University
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of mushroom tyrosinase using L-tyrosine as a substrateMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 12nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 12nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 13nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 13nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 14nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 16nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 18nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 19nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 19nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 23nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 24nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 24nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 24nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 27nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 28nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 33nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 38nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 38nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha/90-beta(Homo sapiens (Human))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 42nMAssay Description:Inhibition of HSP90 (unknown origin) assessed as HER2 degradation by cell-based assayMore data for this Ligand-Target Pair

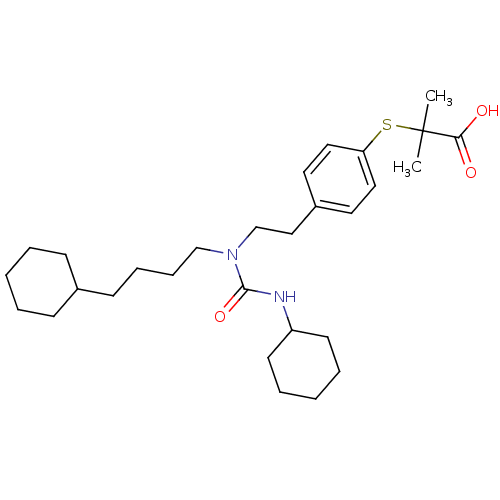
 3D Structure (crystal)
3D Structure (crystal)