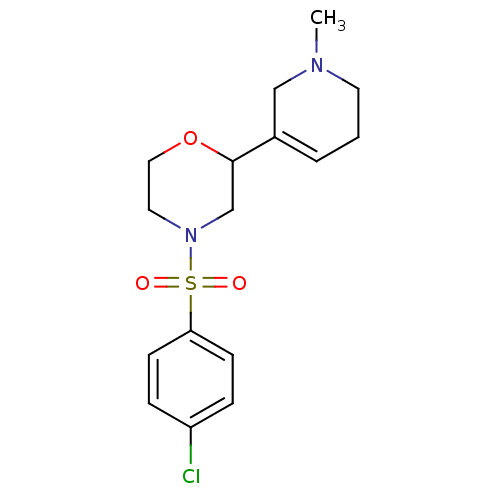
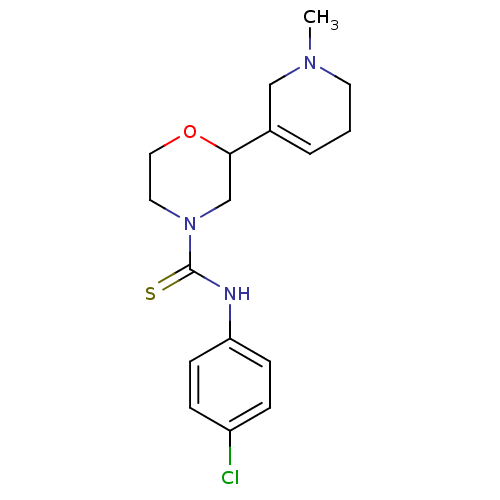
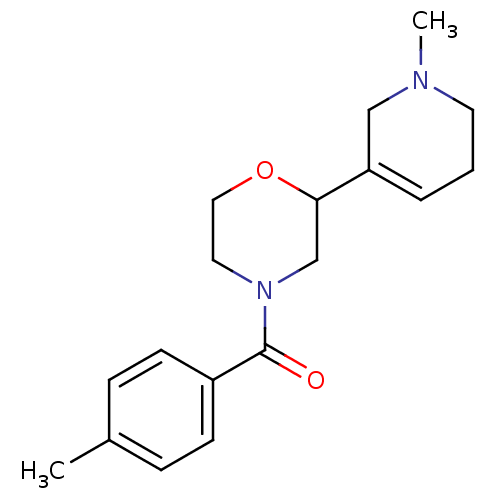
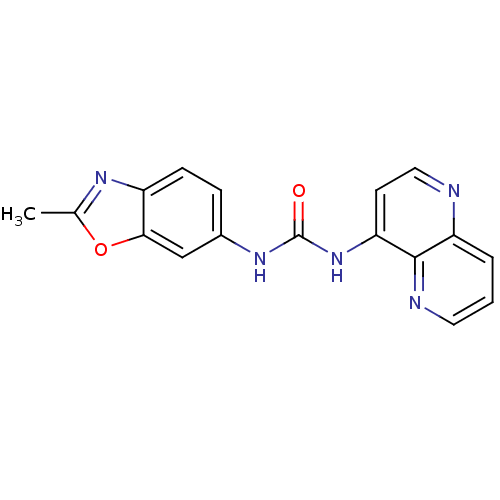
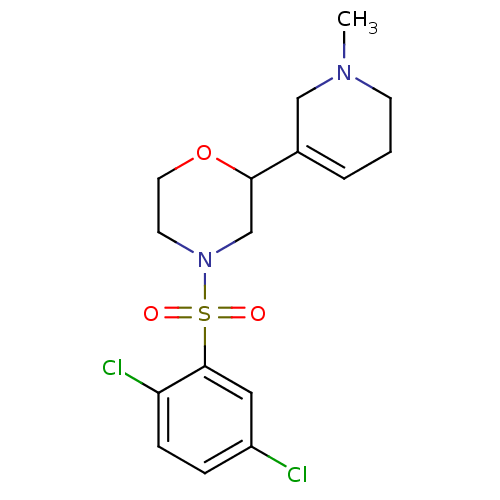
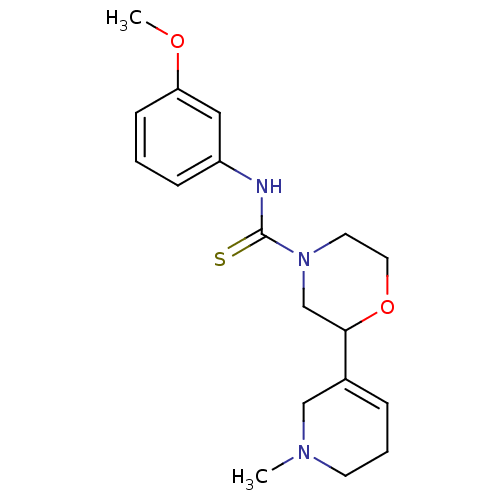
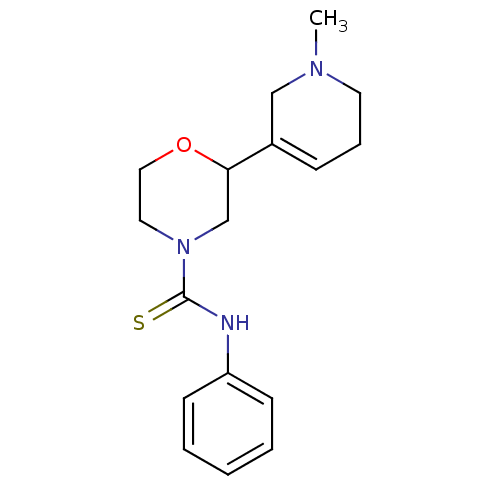
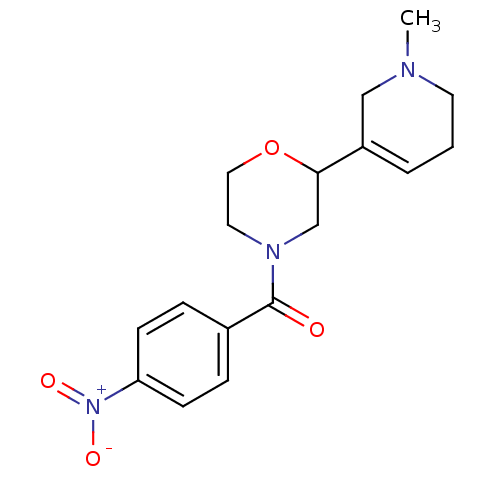
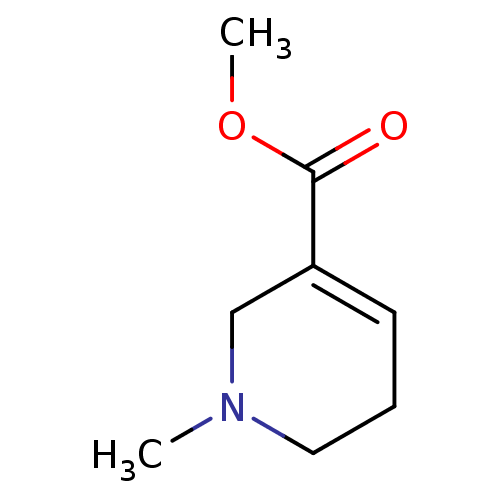
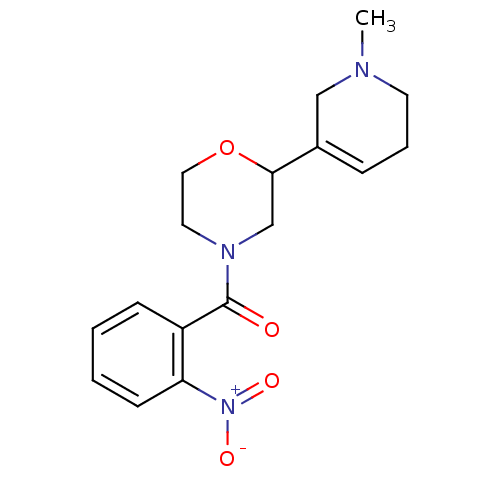
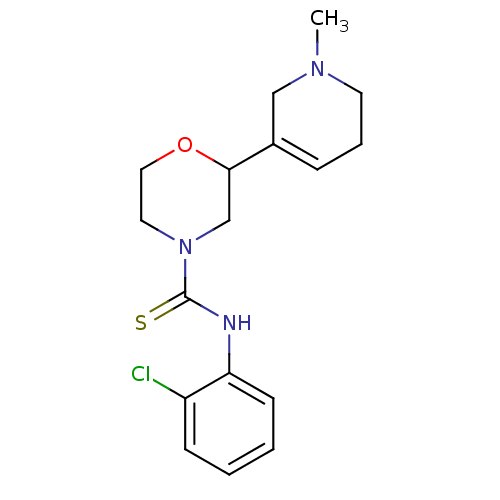
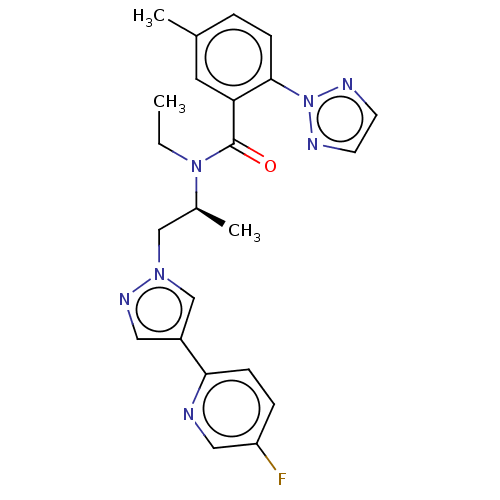
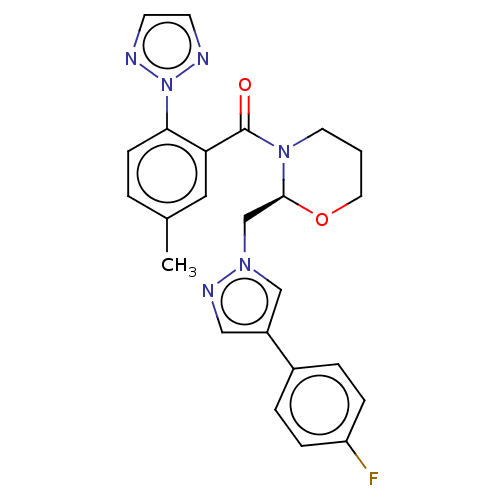
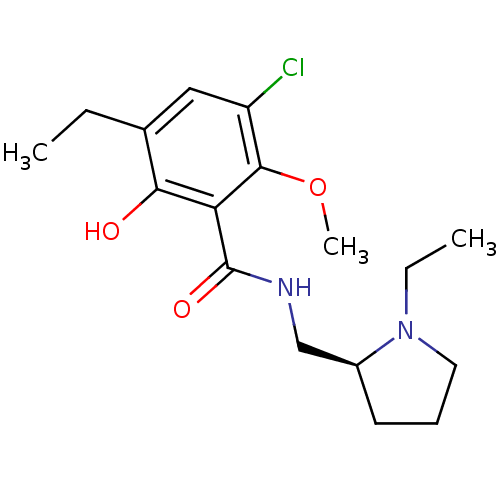
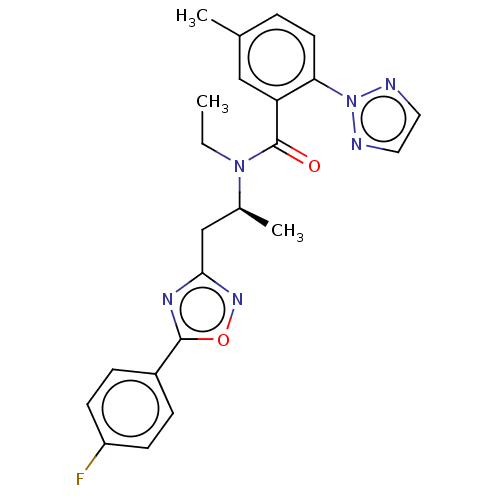
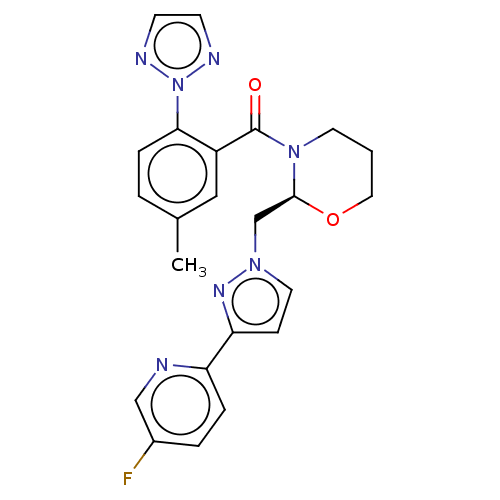
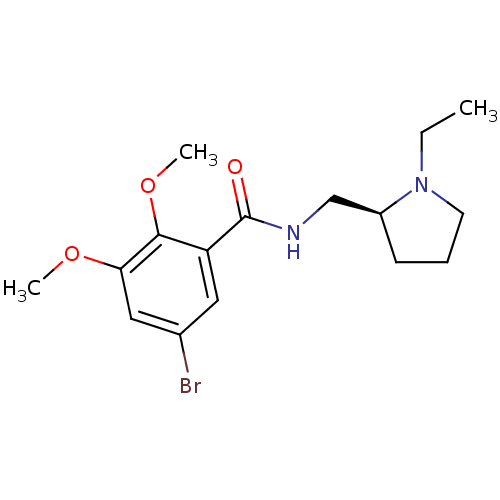
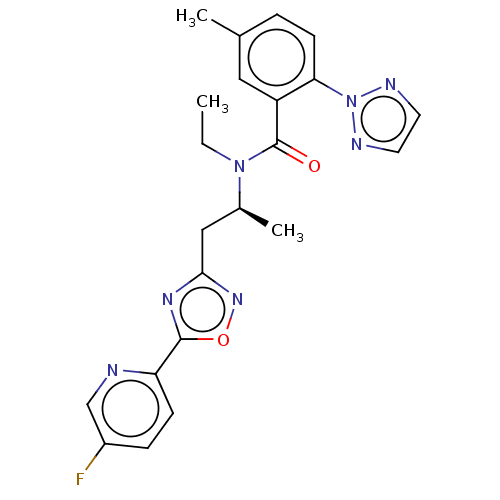
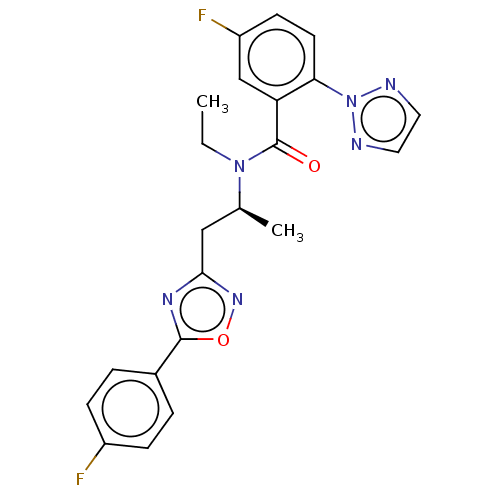
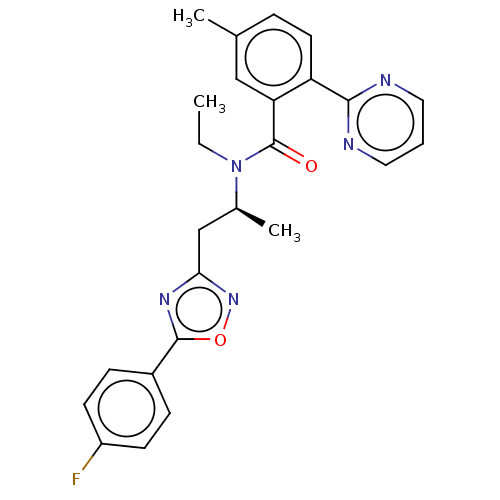
Affinity DataKi: 260nMAssay Description:Displacement of [3H]Quinuclidinyl benzillate from muscarinic M1 receptor in Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 310nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 420nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor in Wistar rat brain cortex after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 670nMAssay Description:Binding affinity to adenosine 2A receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 841nMAssay Description:Inhibition of Plasmodium falciparum Plm4 using synthetic substrate pretreated for 5 mins followed by substrate addition measured immediately by spect...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nMAssay Description:Binding affinity to 5HT2c receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+3nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor in Wistar rat brain cortex after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.92E+3nMAssay Description:Inhibition of Plasmodium falciparum Plm2 using synthetic substrate pretreated for 5 mins followed by substrate addition measured immediately by spect...More data for this Ligand-Target Pair
Affinity DataKi: 1.98E+3nMAssay Description:Inhibition of Plasmodium falciparum Plm4 using synthetic substrate pretreated for 5 mins followed by substrate addition measured immediately by spect...More data for this Ligand-Target Pair
Affinity DataKi: 3.35E+3nMAssay Description:Inhibition of Plasmodium falciparum Plm4 using synthetic substrate pretreated for 5 mins followed by substrate addition measured immediately by spect...More data for this Ligand-Target Pair
Affinity DataKi: 4.00E+3nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 4.00E+3nMAssay Description:Displacement of [3H]Quinuclidinyl benzillate from muscarinic M1 receptor in Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 7.00E+3nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 1.40E+4nMAssay Description:Displacement of [3H]Quinuclidinyl benzillate from muscarinic M1 receptor in Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 1.50E+4nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 1.90E+4nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor in Wistar rat brain cortex after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.70E+4nMAssay Description:Displacement of [3H]Quinuclidinyl benzillate from muscarinic M1 receptor in Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 3.10E+4nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 3.50E+4nMAssay Description:Displacement of [3H]Quinuclidinyl benzillate from muscarinic M1 receptor in Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 4.10E+4nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 4.80E+4nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor in Wistar rat brain cortex after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4.90E+4nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 6.20E+4nMAssay Description:Displacement of [3H]Quinuclidinyl benzillate from muscarinic M1 receptor in Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 6.40E+4nMAssay Description:Displacement of [3H]Quinuclidinyl benzillate from muscarinic M1 receptor in Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 6.50E+4nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor in Wistar rat brain cortex after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 8.50E+4nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 8.60E+4nMAssay Description:Displacement of [3H]Quinuclidinyl benzillate from muscarinic M1 receptor in Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 8.80E+4nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor in Wistar rat brain cortex after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 9.60E+4nMAssay Description:Displacement of [3H]Quinuclidinyl benzillate from muscarinic M1 receptor in Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 9.80E+4nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 1.26E+5nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor in Wistar rat brain cortex after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.14E+5nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 0.380nMAssay Description:Inhibition of human plasma renin assessed as reduction in endogenous angiotensin 1 production in plasmaMore data for this Ligand-Target Pair
Affinity DataIC50: 0.767nMAssay Description:Antagonist activity at human OX1R expressed in CHO cells assessed as reduction in [Ala6,12]orexin-A-induced intracellular Ca2+ mobilization incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 0.784nMAssay Description:Antagonist activity at human recombinant OX2R expressed in CHOK1 cells assessed as inhibition of orexin A-induced intracellular Ca2+ release preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 0.820nMAssay Description:Antagonist activity at human OX1R expressed in CHO cells assessed as reduction in [Ala6,12]orexin-A-induced intracellular Ca2+ mobilization incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 0.920nMAssay Description:Antidopamine activity in vitro by ability to displace [3H]spiperone from rat brain striatal preparations.More data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:Antagonist activity at human OX1R expressed in CHO cells assessed as reduction in [Ala6,12]orexin-A-induced intracellular Ca2+ mobilization incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:Antagonist activity at human OX1R expressed in CHO cells assessed as reduction in [Ala6,12]orexin-A-induced intracellular Ca2+ mobilization incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:Antagonist activity at human recombinant OX1R expressed in CHOK1 cells assessed as inhibition of orexin A-induced intracellular Ca2+ release preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:The affinity for the Dopamine receptor D2 was assessed by the inhibition of [3H]-spiperone binding in rat striatal membranes in vitro.More data for this Ligand-Target Pair
Affinity DataIC50: 1.30nMAssay Description:Antagonist activity at human recombinant OX2R expressed in CHOK1 cells assessed as inhibition of orexin A-induced intracellular Ca2+ release preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30nMAssay Description:Antagonist activity at human OX1R expressed in CHO cells assessed as reduction in [Ala6,12]orexin-A-induced intracellular Ca2+ mobilization incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30nMAssay Description:Antagonist activity at human OX1R expressed in CHO cells assessed as reduction in [Ala6,12]orexin-A-induced intracellular Ca2+ mobilization incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30nMAssay Description:Antagonist activity at human OX1R expressed in CHO cells assessed as reduction in [Ala6,12]orexin-A-induced intracellular Ca2+ mobilization incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Antagonist activity at human OX1R expressed in CHO cells assessed as reduction in [Ala6,12]orexin-A-induced intracellular Ca2+ mobilization incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:The affinity for the Dopamine receptor D2 was assessed by the inhibition of [3H]-spiperone binding in rat striatal membranes in vitro.More data for this Ligand-Target Pair
Affinity DataIC50: 1.5nMAssay Description:Antidopamine activity in vitro by ability to displace [3H]spiperone from rat brain striatal preparations.More data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Antagonist activity at human recombinant OX2R expressed in CHOK1 cells assessed as inhibition of orexin A-induced intracellular Ca2+ release preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Antagonist activity at human OX1R expressed in CHO cells assessed as reduction in [Ala6,12]orexin-A-induced intracellular Ca2+ mobilization incubated...More data for this Ligand-Target Pair