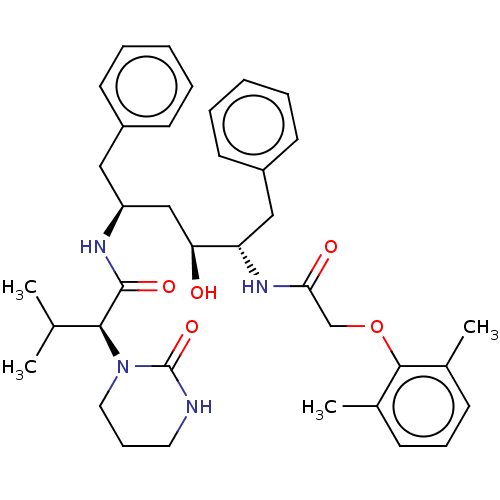
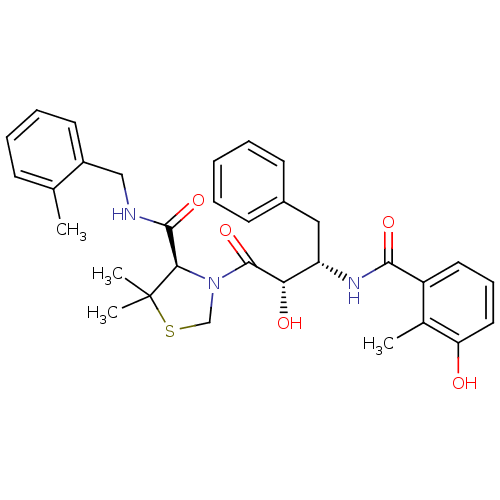
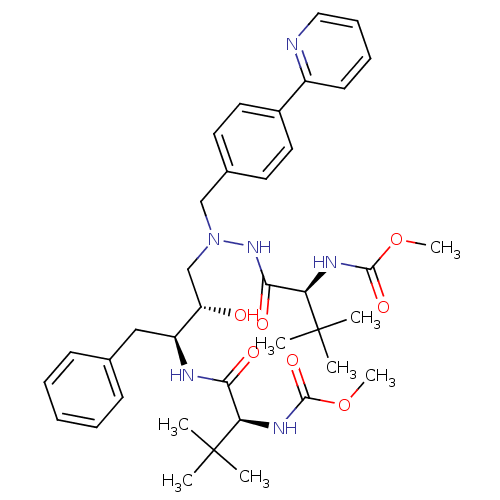
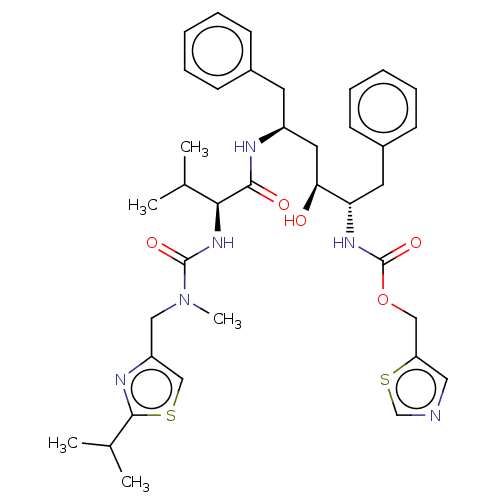
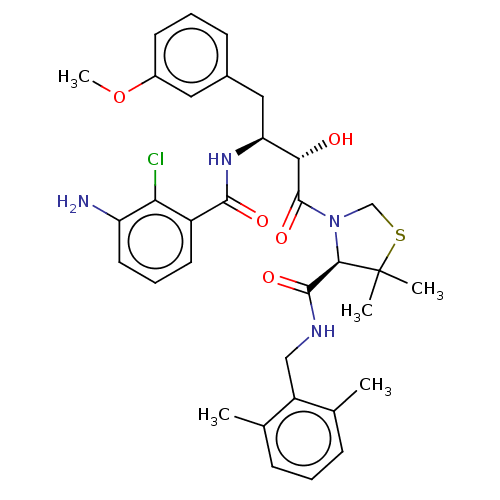
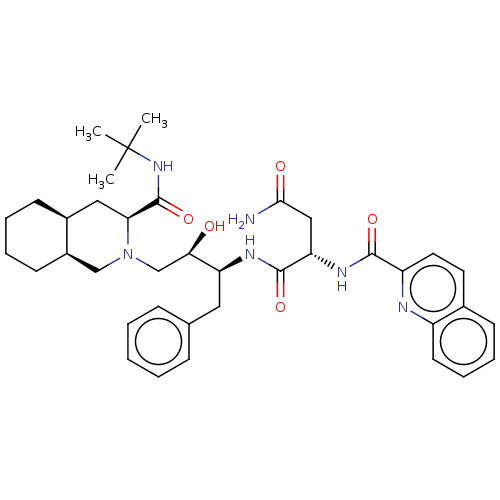
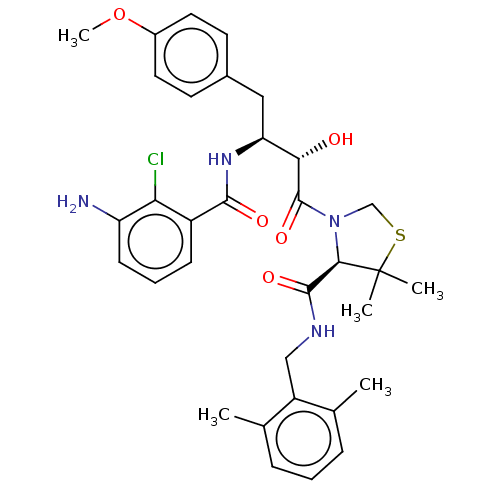
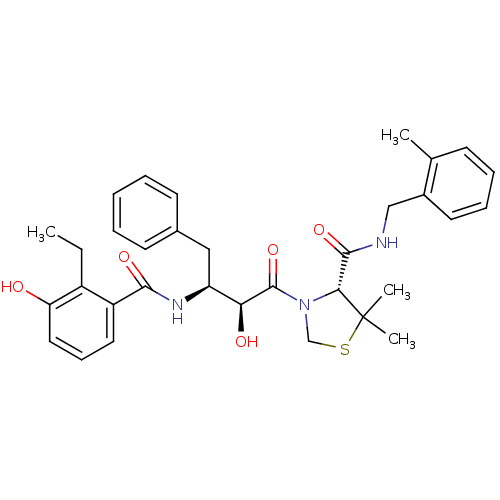
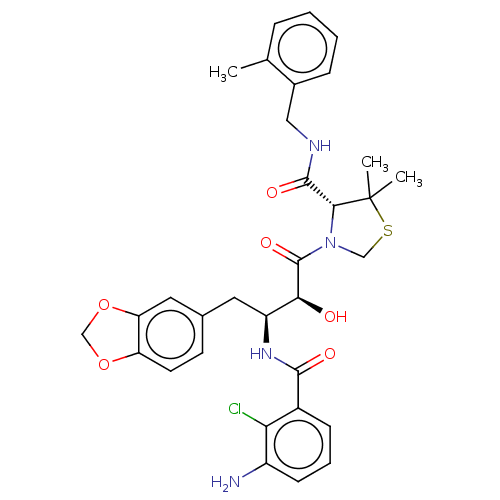
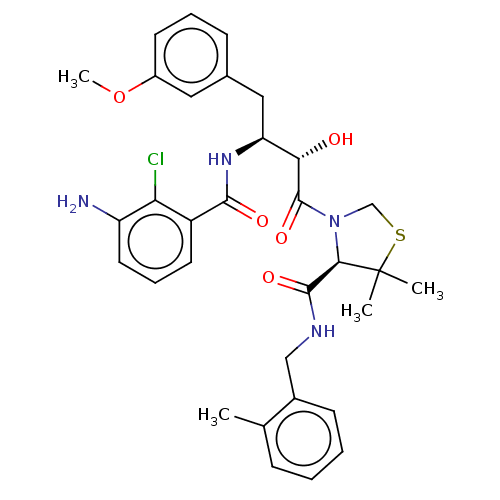
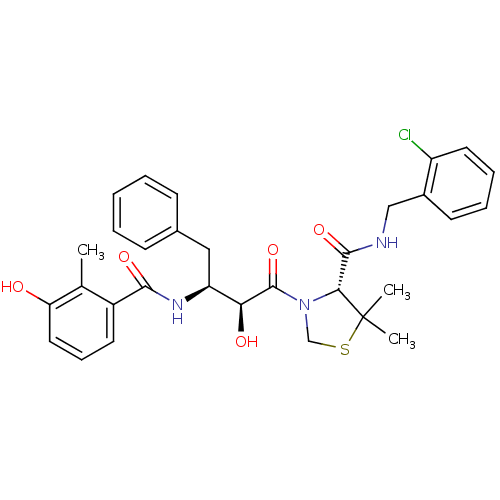
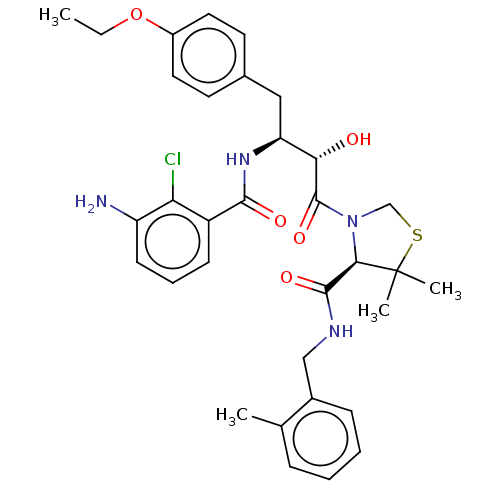
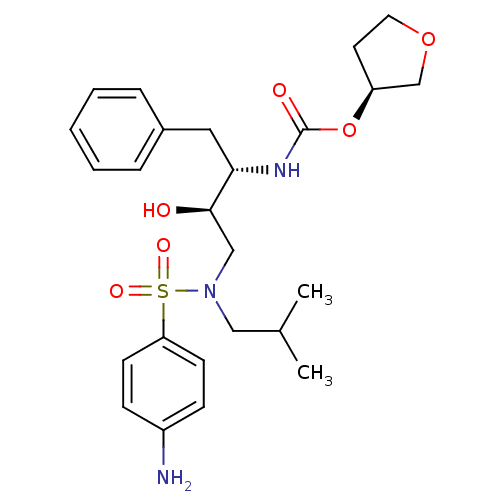
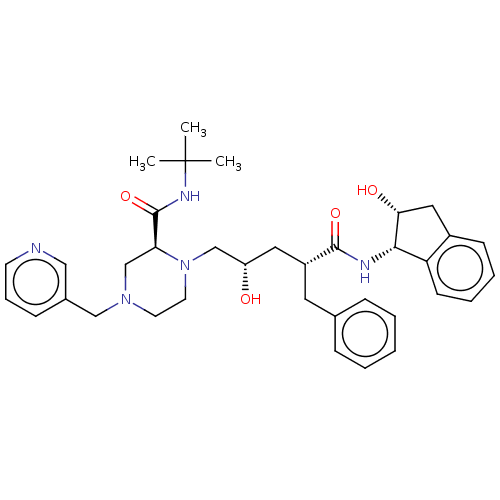
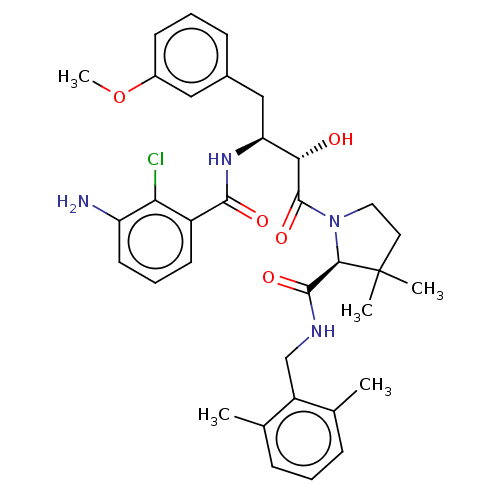
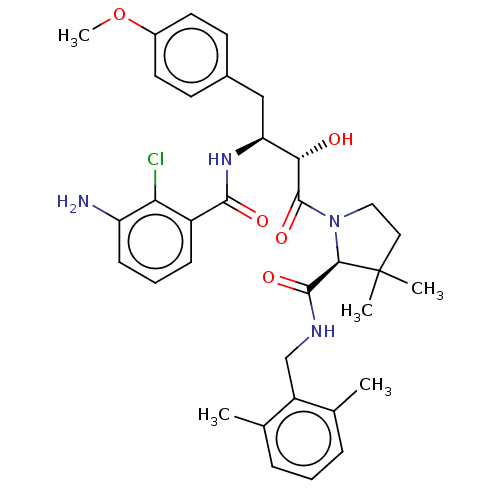
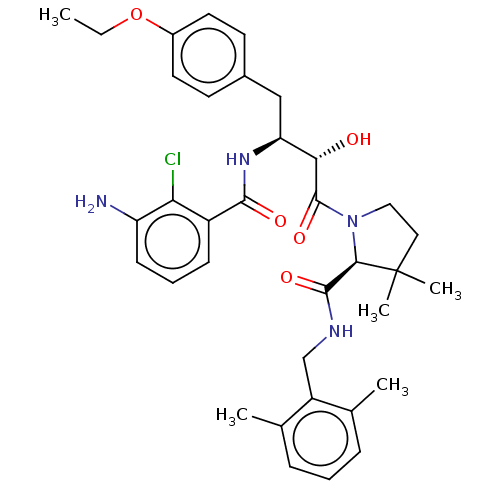
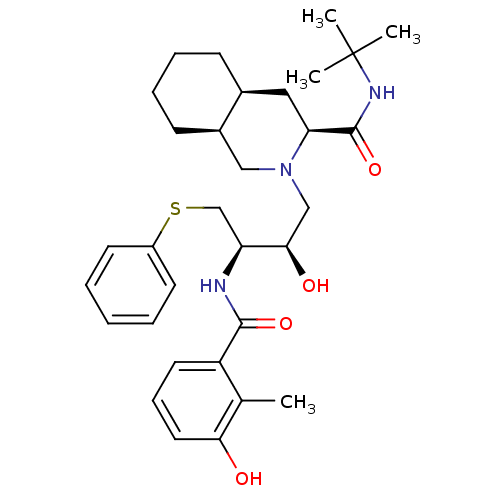
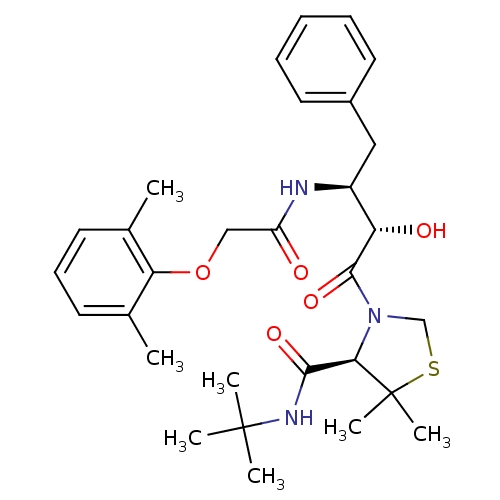
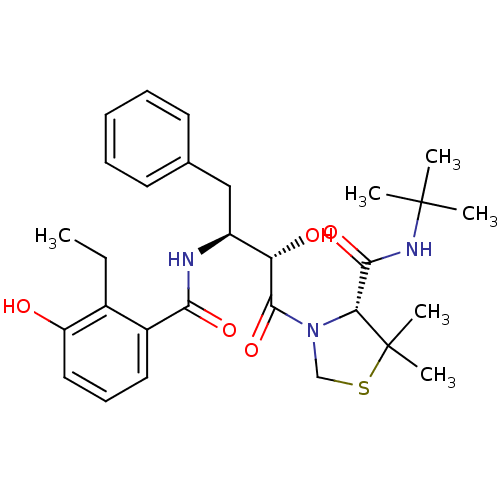
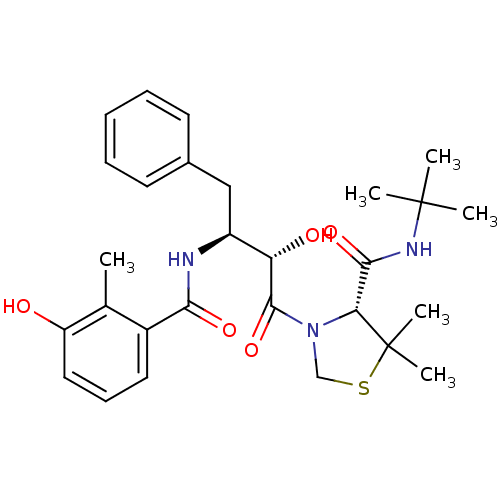
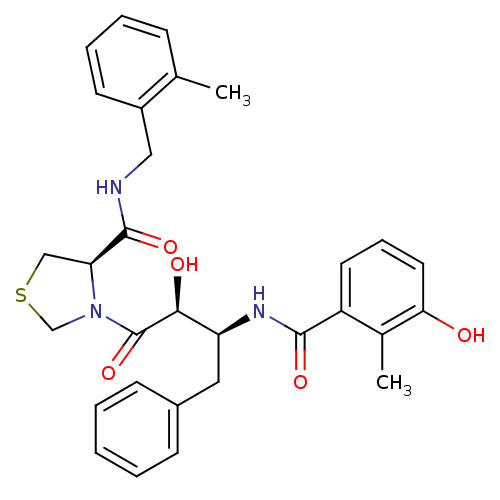
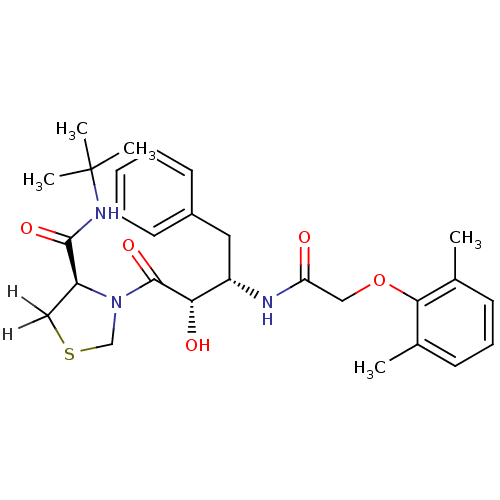
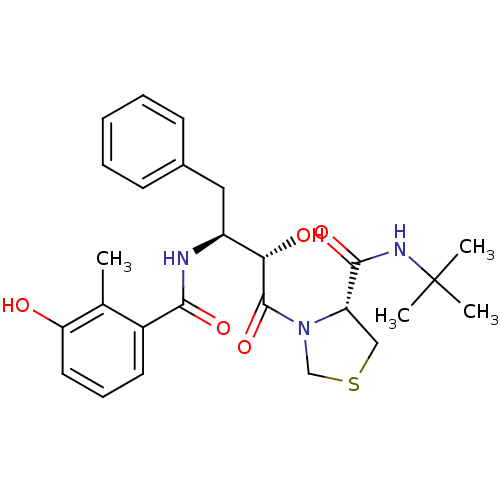
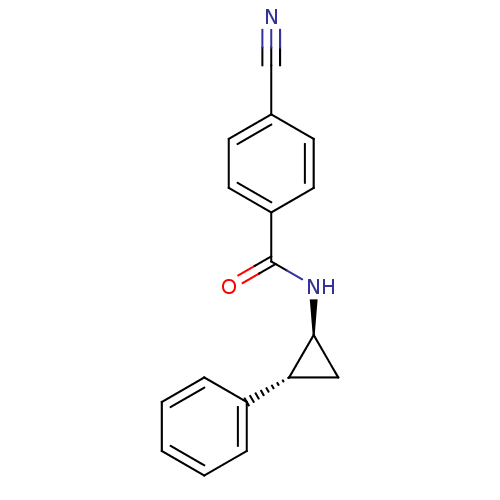
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 0.00230nM ΔG°: -69.1kJ/molepH: 6.2 T: 2°CAssay Description:Inhibition constants were determined by a fluorometric assay with the fluorogenic substrate Arg-Glu(EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Lys(DABCYL)-Ar...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 0.00550nM ΔG°: -66.9kJ/molepH: 6.2 T: 2°CAssay Description:Inhibition constants were determined by a fluorometric assay with the fluorogenic substrate Arg-Glu(EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Lys(DABCYL)-Ar...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 0.00680nM ΔG°: -66.3kJ/molepH: 6.2 T: 2°CAssay Description:Inhibition constants were determined by a fluorometric assay with the fluorogenic substrate Arg-Glu(EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Lys(DABCYL)-Ar...More data for this Ligand-Target Pair
Affinity DataKi: 0.0160nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.0350nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.0700nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 0.0880nM ΔG°: -59.7kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 0.0980nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.134nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.138nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.138nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.152nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 0.160nM ΔG°: -58.2kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 0.163nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.255nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 0.290nM ΔG°: -56.6kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 0.310nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.311nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 0.330nM ΔG°: -56.3kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 0.353nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.359nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.739nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 0.740nM ΔG°: -54.2kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 0.75nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.804nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.861nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.931nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 1.40nM ΔG°: -52.6kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 2.24nM ΔG°: -51.4kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 5.14nM ΔG°: -49.2kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 8.91nM ΔG°: -47.8kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 21.7nM ΔG°: -45.5kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Cancer Institute-Bethesda
National Cancer Institute-Bethesda
Affinity DataKi: 24.9nM ΔG°: -45.1kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Inhibition of rat soluble epoxide hydrolase in cell free systemMore data for this Ligand-Target Pair
TargetBifunctional epoxide hydrolase 2(Homo sapiens (Human))
Dainippon Sumitomo Pharma
Curated by ChEMBL
Dainippon Sumitomo Pharma
Curated by ChEMBL
Affinity DataIC50: 2.29nMAssay Description:Inhibition of human soluble epoxide hydrolase in cell free systemMore data for this Ligand-Target Pair
Affinity DataIC50: 3.02nMAssay Description:Inhibition of rat soluble epoxide hydrolase in cell free systemMore data for this Ligand-Target Pair
TargetBifunctional epoxide hydrolase 2(Homo sapiens (Human))
Dainippon Sumitomo Pharma
Curated by ChEMBL
Dainippon Sumitomo Pharma
Curated by ChEMBL
Affinity DataIC50: 4.68nMAssay Description:Inhibition of soluble epoxide hydrolase in human HepG2 cells after 30 minsMore data for this Ligand-Target Pair
TargetBifunctional epoxide hydrolase 2(Homo sapiens (Human))
Dainippon Sumitomo Pharma
Curated by ChEMBL
Dainippon Sumitomo Pharma
Curated by ChEMBL
Affinity DataIC50: 6.46nMAssay Description:Inhibition of human soluble epoxide hydrolase in cell free systemMore data for this Ligand-Target Pair
TargetBifunctional epoxide hydrolase 2(Homo sapiens (Human))
Dainippon Sumitomo Pharma
Curated by ChEMBL
Dainippon Sumitomo Pharma
Curated by ChEMBL
Affinity DataIC50: 7.94nMAssay Description:Inhibition of soluble epoxide hydrolase in human HepG2 cells after 30 minsMore data for this Ligand-Target Pair
TargetBifunctional epoxide hydrolase 2(Homo sapiens (Human))
Dainippon Sumitomo Pharma
Curated by ChEMBL
Dainippon Sumitomo Pharma
Curated by ChEMBL
Affinity DataIC50: 8.5nMAssay Description:Inhibition of human soluble epoxide hydrolase in cell free systemMore data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of rat soluble epoxide hydrolase in cell free systemMore data for this Ligand-Target Pair
TargetBifunctional epoxide hydrolase 2(Homo sapiens (Human))
Dainippon Sumitomo Pharma
Curated by ChEMBL
Dainippon Sumitomo Pharma
Curated by ChEMBL
Affinity DataIC50: 13nMAssay Description:Inhibition of soluble epoxide hydrolase in human HepG2 cells after 30 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 19.9nMAssay Description:Inhibition of rat soluble epoxide hydrolase in cell free systemMore data for this Ligand-Target Pair
TargetBifunctional epoxide hydrolase 2(Homo sapiens (Human))
Dainippon Sumitomo Pharma
Curated by ChEMBL
Dainippon Sumitomo Pharma
Curated by ChEMBL
Affinity DataIC50: 30.2nMAssay Description:Inhibition of human soluble epoxide hydrolase in cell free systemMore data for this Ligand-Target Pair
TargetBifunctional epoxide hydrolase 2(Homo sapiens (Human))
Dainippon Sumitomo Pharma
Curated by ChEMBL
Dainippon Sumitomo Pharma
Curated by ChEMBL
Affinity DataIC50: 39.8nMAssay Description:Inhibition of human soluble epoxide hydrolase in cell free systemMore data for this Ligand-Target Pair
TargetBifunctional epoxide hydrolase 2(Homo sapiens (Human))
Dainippon Sumitomo Pharma
Curated by ChEMBL
Dainippon Sumitomo Pharma
Curated by ChEMBL
Affinity DataIC50: 61.7nMAssay Description:Inhibition of human soluble epoxide hydrolase in cell free systemMore data for this Ligand-Target Pair
TargetBifunctional epoxide hydrolase 2(Homo sapiens (Human))
Dainippon Sumitomo Pharma
Curated by ChEMBL
Dainippon Sumitomo Pharma
Curated by ChEMBL
Affinity DataIC50: 81.3nMAssay Description:Inhibition of soluble epoxide hydrolase in human HepG2 cells after 30 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 240nMAssay Description:Inhibition of rat soluble epoxide hydrolase in cell free systemMore data for this Ligand-Target Pair
TargetBifunctional epoxide hydrolase 2(Homo sapiens (Human))
Dainippon Sumitomo Pharma
Curated by ChEMBL
Dainippon Sumitomo Pharma
Curated by ChEMBL
Affinity DataIC50: 565nMAssay Description:Inhibition of human soluble epoxide hydrolase in cell free systemMore data for this Ligand-Target Pair

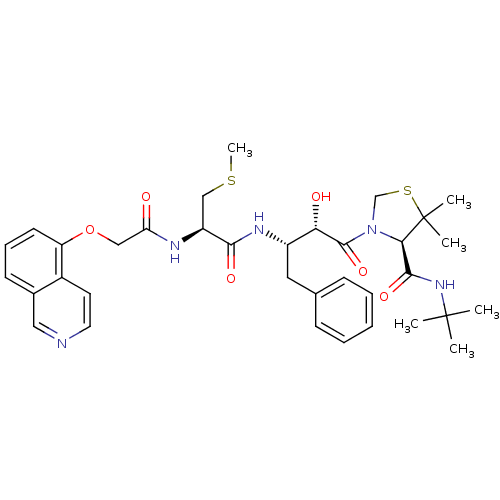
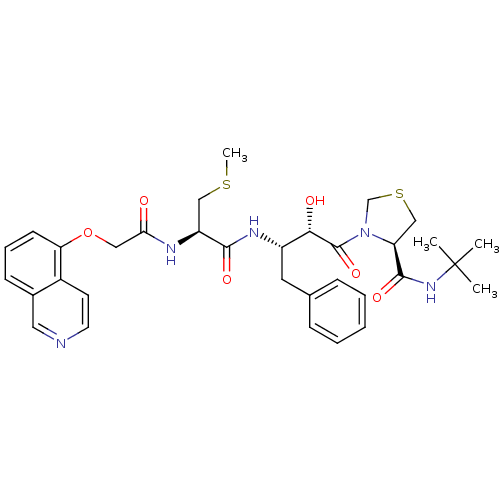
 3D Structure (crystal)
3D Structure (crystal)