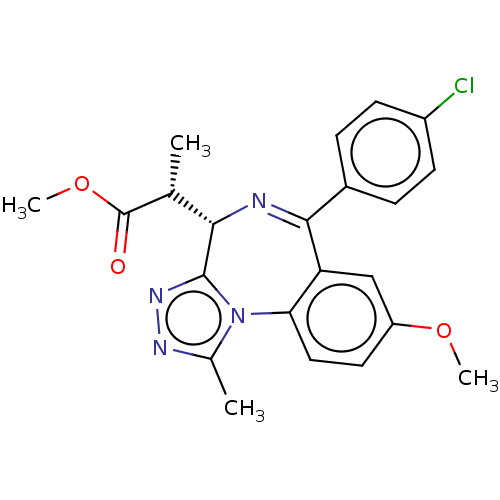
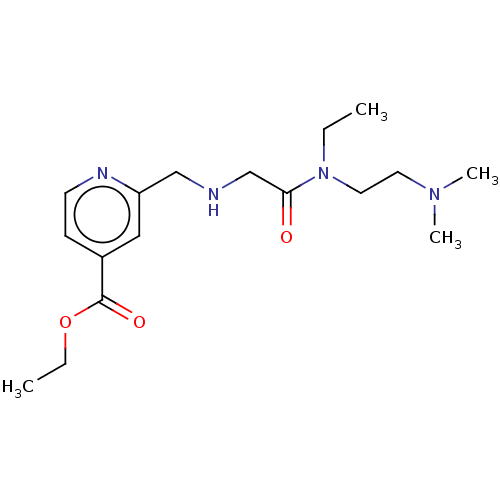
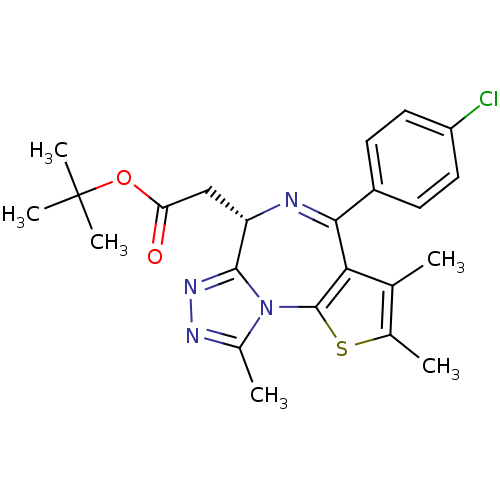
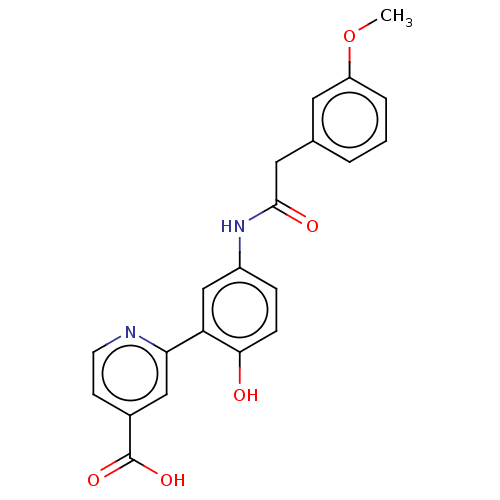
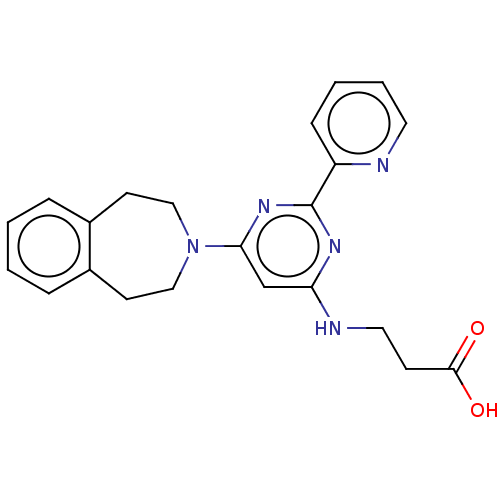
Affinity DataKi: 2.20nMAssay Description:Competitive inhibition of N-terminal His6-tagged KDM4C (1 to 352 residues) (unknown origin) expressed in Escherichia coli Rosetta 2(DE3)pLysS using A...More data for this Ligand-Target Pair
Affinity DataKi: 43nMAssay Description:Competitive inhibition of N-terminal His6-tagged KDM4C (1 to 352 residues) (unknown origin) expressed in Escherichia coli Rosetta 2(DE3)pLysS using A...More data for this Ligand-Target Pair
Affinity DataKi: 683nMAssay Description:Competitive inhibition of N-terminal His6-tagged KDM4C (1 to 352 residues) (unknown origin) expressed in Escherichia coli Rosetta 2(DE3)pLysS using A...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Induction of ALK degradation in human NCI-H3122 cells after 16 hrs by immunoblot methodMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Induction of ALK degradation in human NCI-H3122 cells after 16 hrs by immunoblot methodMore data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 24nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 26nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
TargetBromodomain-containing protein 4(Homo sapiens (Human))
Dana Farber Cancer Institute
Curated by ChEMBL
Dana Farber Cancer Institute
Curated by ChEMBL
Affinity DataIC50: 39nMAssay Description:Displacement of FITC-conjugated JQ1 from wildtype BRD4 BD1 (unknown origin) expressed in Escherichia coli BL21-DE3 rosetta cells incubated for 15 min...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Induction of ALK degradation in human KARPAS299 cells after 16 hrs by immunoblot methodMore data for this Ligand-Target Pair
Affinity DataIC50: 49nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
TargetBromodomain-containing protein 4(Homo sapiens (Human))
Dana Farber Cancer Institute
Curated by ChEMBL
Dana Farber Cancer Institute
Curated by ChEMBL
Affinity DataIC50: 49nMAssay Description:Displacement of FITC-conjugated JQ1 from wildtype BRD4 BD1 (unknown origin) expressed in Escherichia coli BL21-DE3 rosetta cells incubated for 15 min...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Induction of ALK degradation in human Kelly cells after 16 hrs by immunoblot methodMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Induction of ALK degradation in human Kelly cells after 16 hrs by immunoblot methodMore data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 64nMAssay Description:Inhibition of N-terminal His6-tagged KDM4C (1 to 352 residues) (unknown origin) expressed in Escherichia coli Rosetta 2(DE3)pLysS using biotinylated ...More data for this Ligand-Target Pair
Affinity DataIC50: 65nMAssay Description:Inhibition of N-terminal His6-tagged KDM4C (1 to 352 residues) (unknown origin) expressed in Escherichia coli Rosetta 2(DE3)pLysS using biotinylated ...More data for this Ligand-Target Pair
Affinity DataIC50: 65nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 69nMAssay Description:Inhibition of N-terminal His6-tagged KDM4C (1 to 352 residues) (unknown origin) expressed in Escherichia coli Rosetta 2(DE3)pLysS using biotinylated ...More data for this Ligand-Target Pair
Affinity DataIC50: 69nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 69nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 71nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 77nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 98nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
TargetBromodomain-containing protein 4(Homo sapiens (Human))
Dana Farber Cancer Institute
Curated by ChEMBL
Dana Farber Cancer Institute
Curated by ChEMBL
Affinity DataIC50: 123nMAssay Description:Displacement of FITC-conjugated JQ1 from BRD4 BD2 (unknown origin) expressed in Escherichia coli BL21-DE3 rosetta cells incubated for 15 mins by comp...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:Inhibition of N-terminal His6-tagged KDM4C (1 to 352 residues) (unknown origin) expressed in Escherichia coli Rosetta 2(DE3)pLysS using biotinylated ...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:Inhibition of N-terminal His6-tagged KDM4C (1 to 352 residues) (unknown origin) expressed in Escherichia coli Rosetta 2(DE3)pLysS using biotinylated ...More data for this Ligand-Target Pair
Affinity DataIC50: 160nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of N-terminal His6-tagged KDM4C (1 to 352 residues) (unknown origin) expressed in Escherichia coli Rosetta 2(DE3)pLysS using biotinylated ...More data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:Induction of ALK degradation in human KARPAS299 cells after 16 hrs by immunoblot methodMore data for this Ligand-Target Pair
Affinity DataIC50: 188nMAssay Description:Inhibition of N-terminal His6-tagged KDM4C (1 to 352 residues) (unknown origin) expressed in Escherichia coli Rosetta 2(DE3)pLysS using biotinylated ...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair

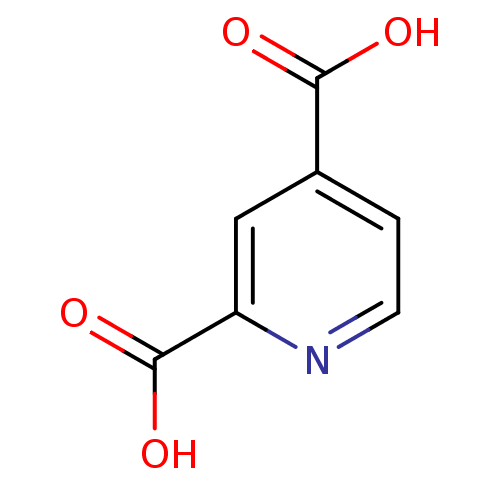
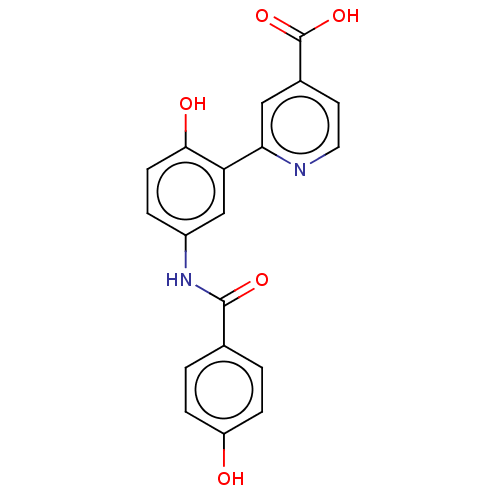

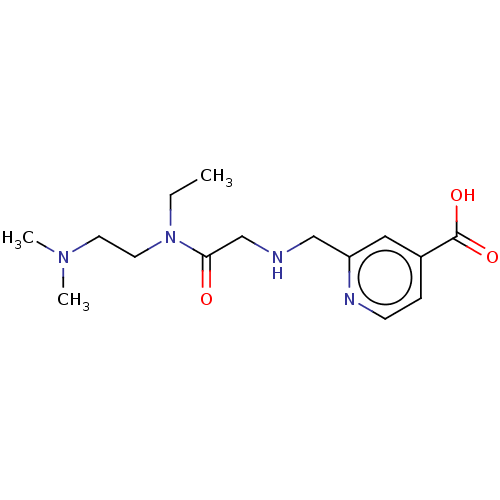



 3D Structure (crystal)
3D Structure (crystal)