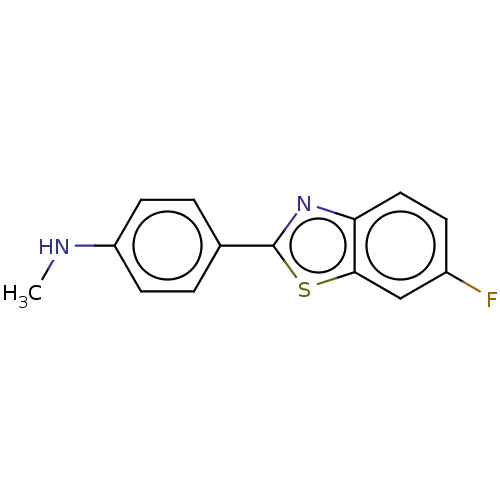
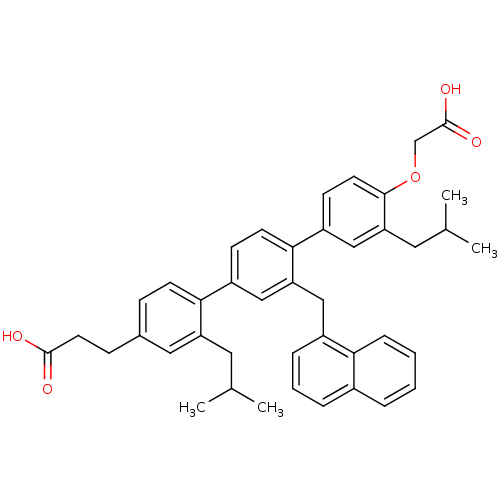
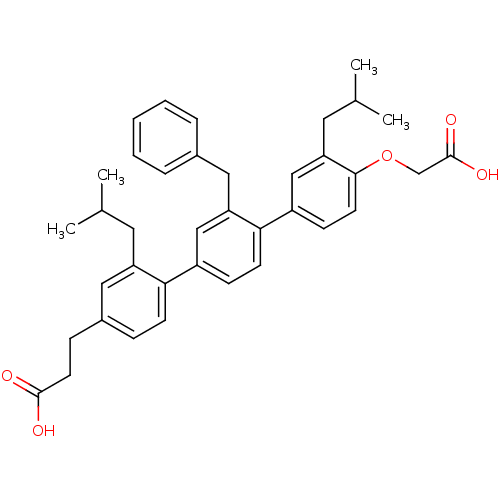
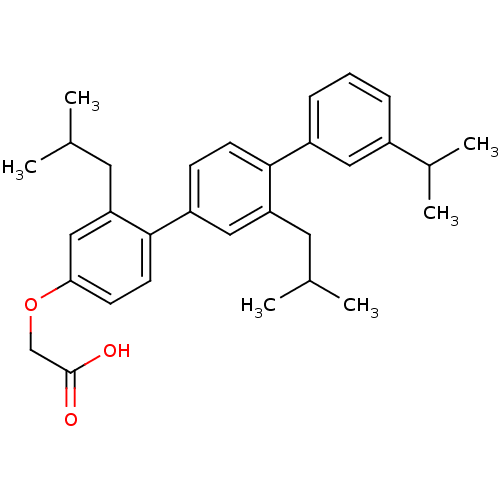
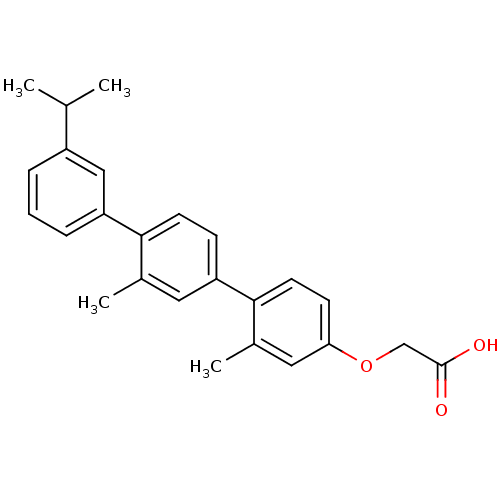
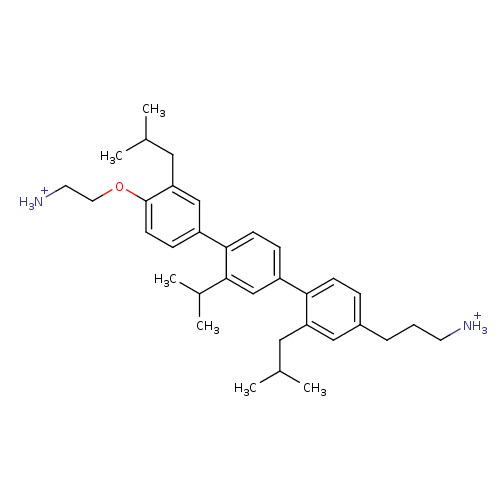
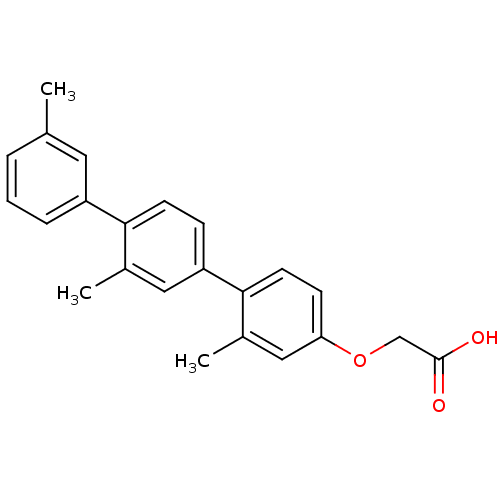
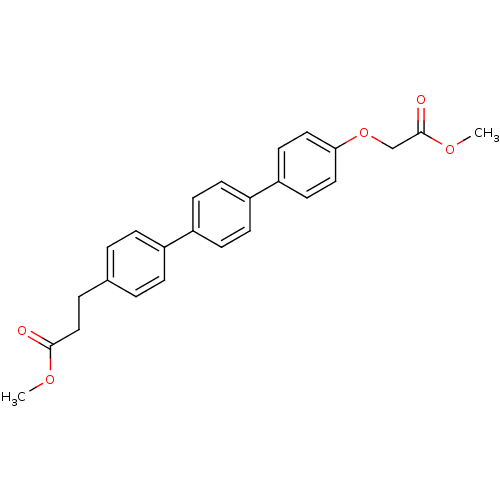
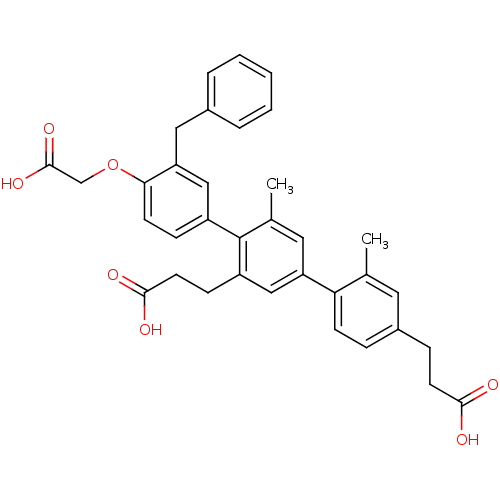
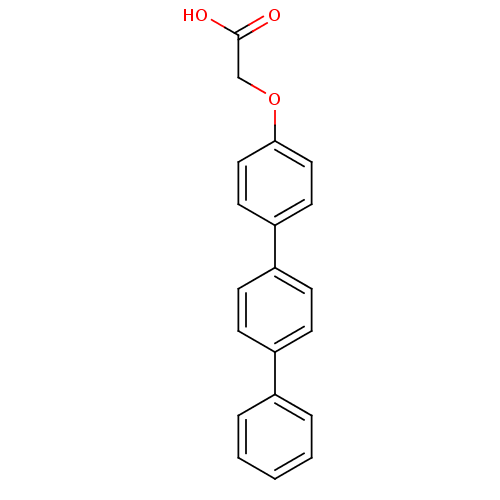
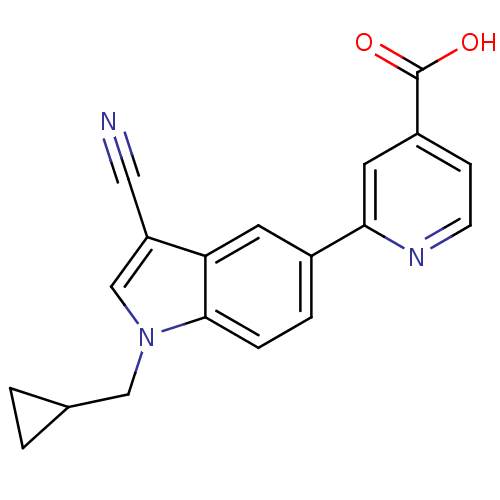
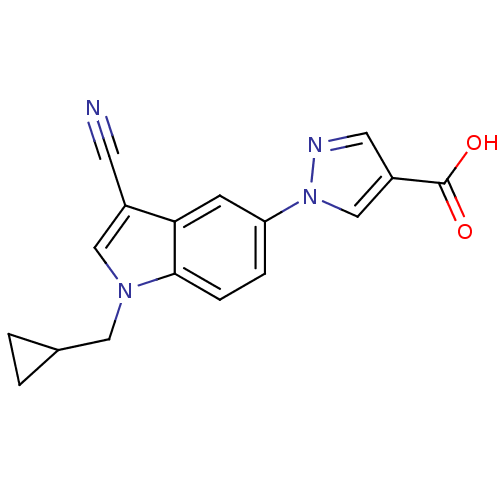
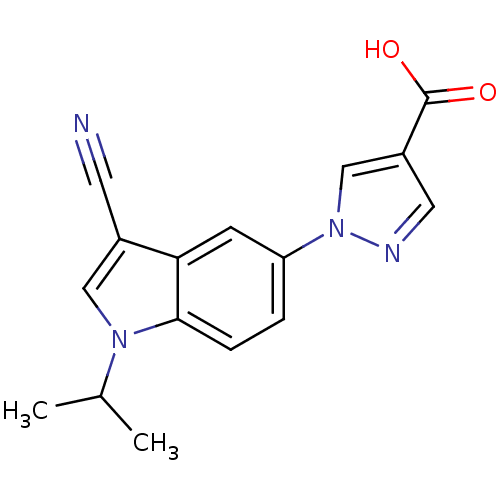
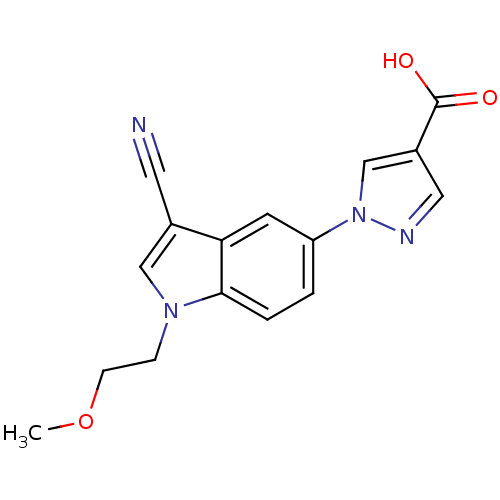
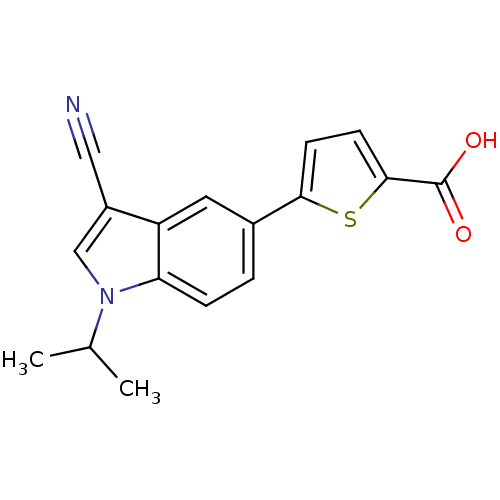
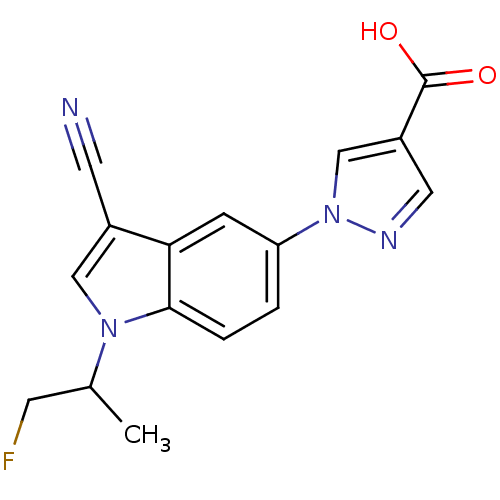
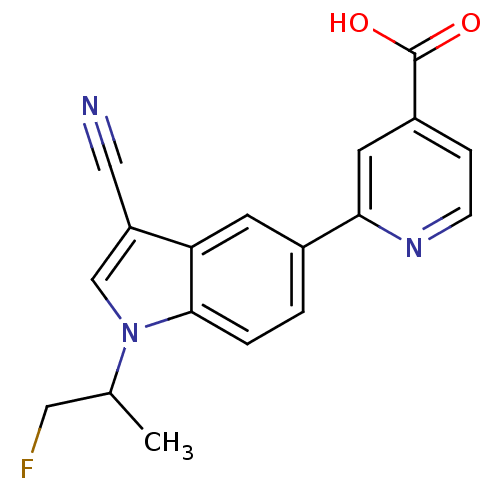
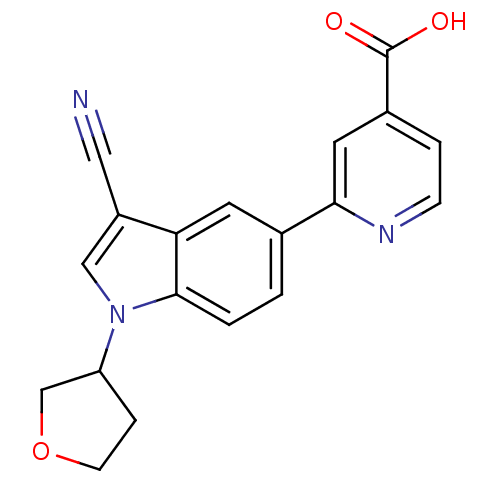
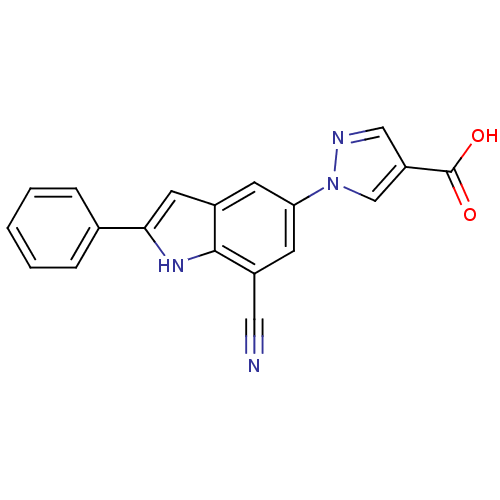
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]PIB from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
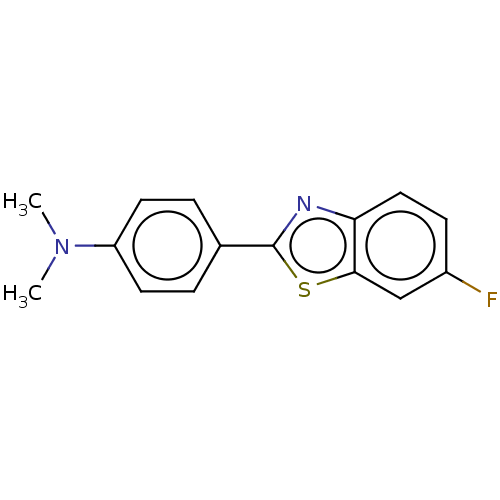
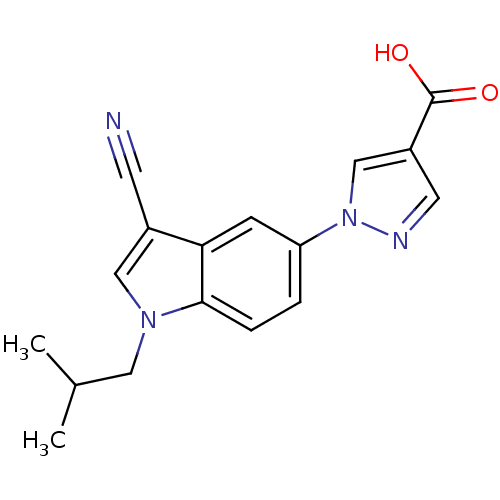
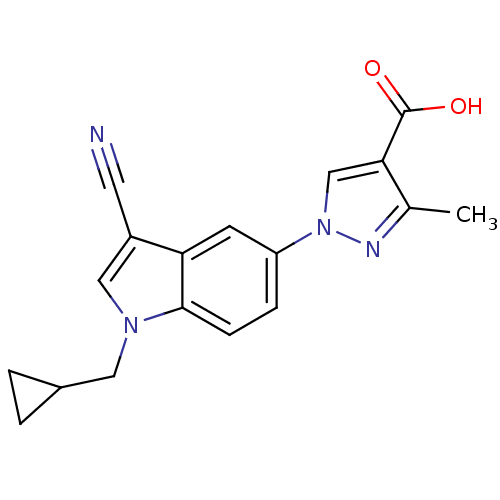
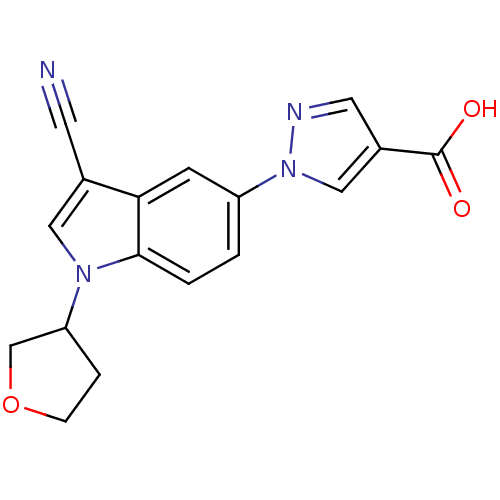
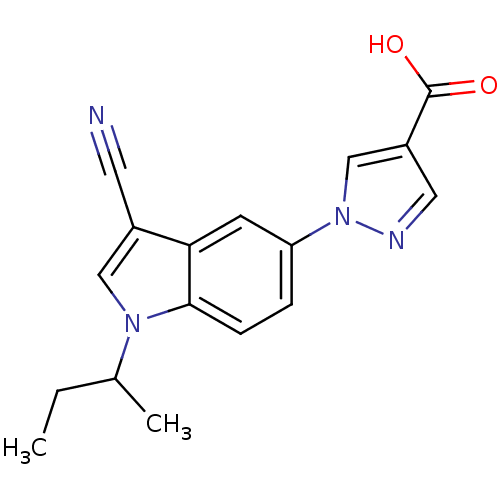
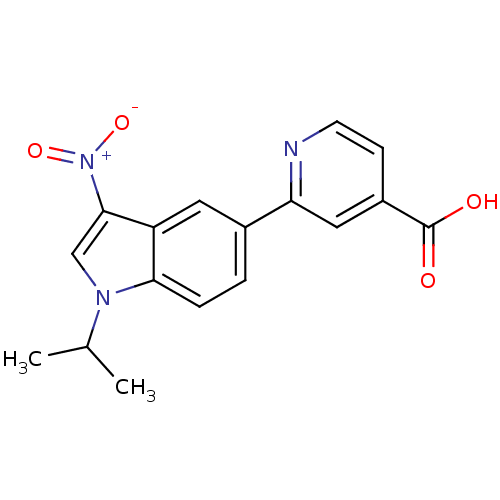
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 0.300nMAssay Description:Displacement of [3H]PIB from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
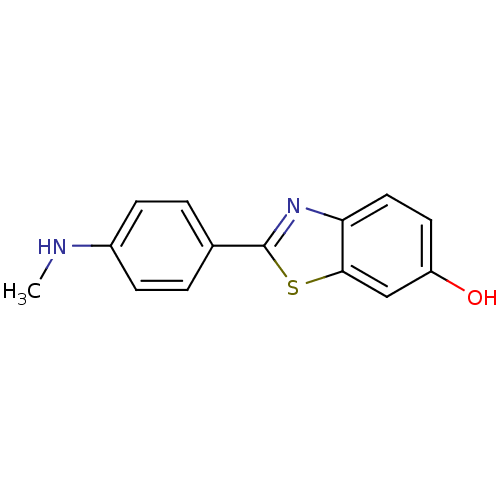
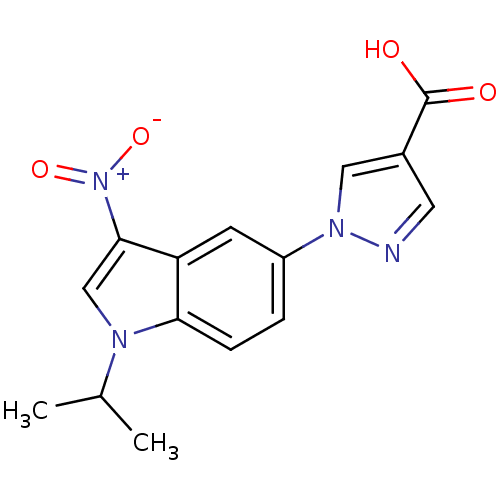
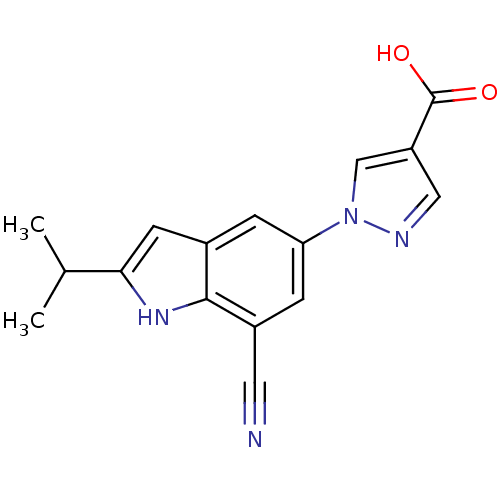
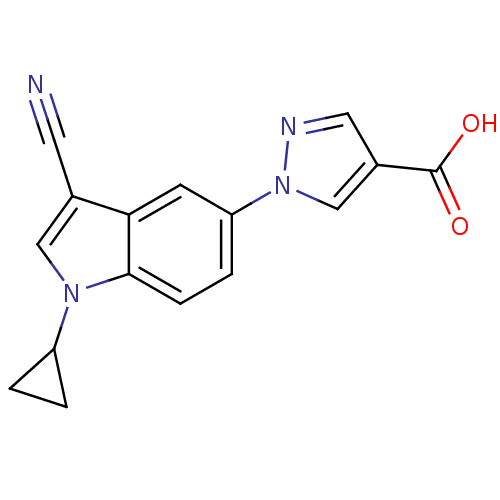
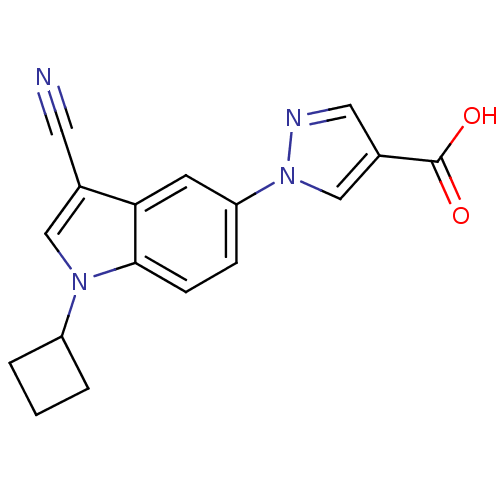
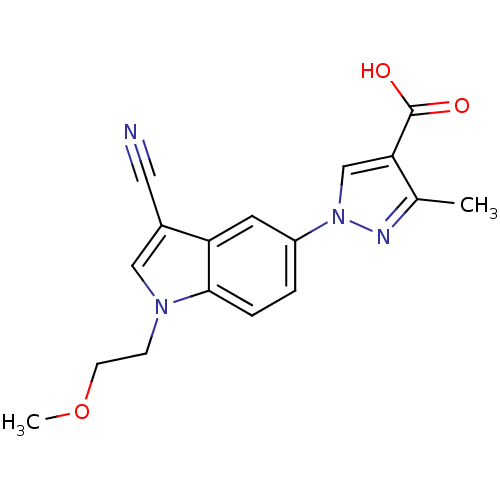
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 1.60nMAssay Description:Displacement of [3H]PIB from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
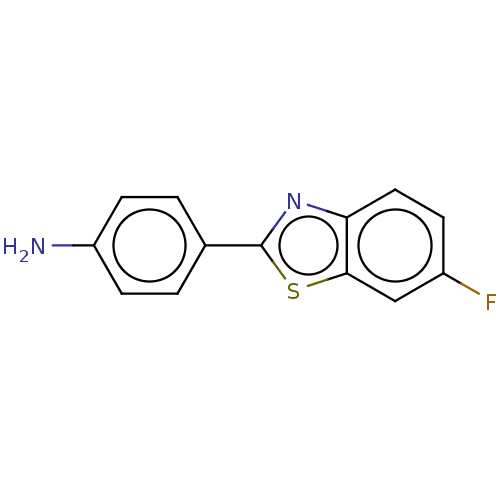
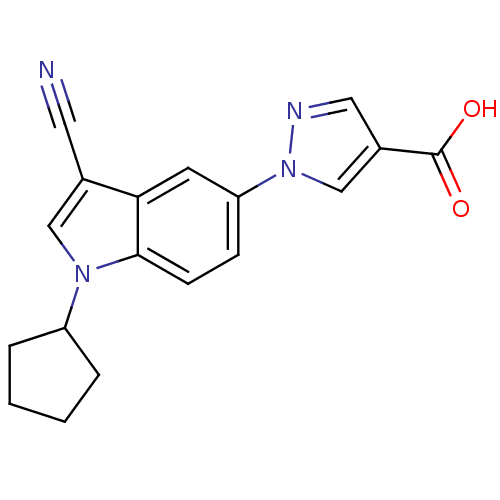
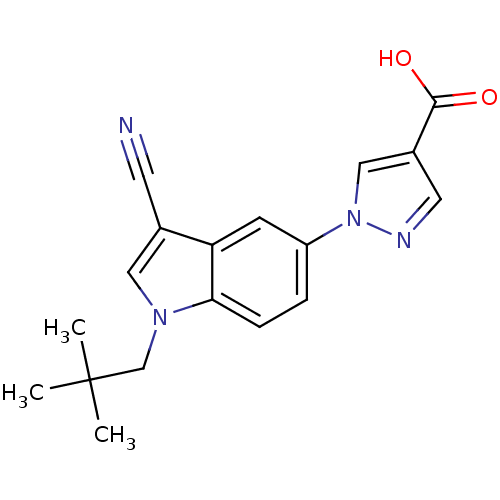
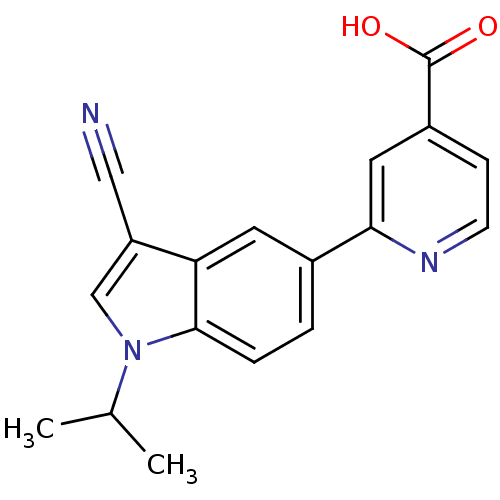
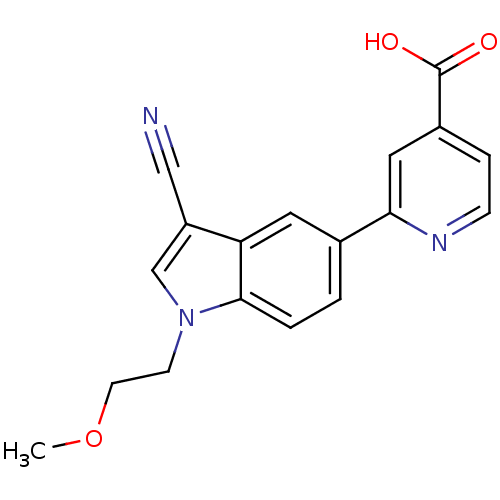
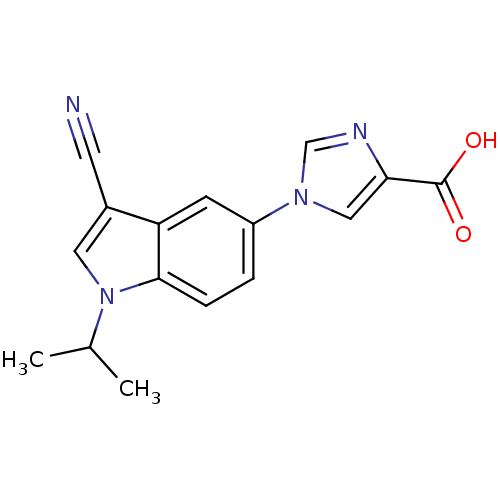
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 5.5nMAssay Description:Displacement of [125I]TZDM from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 5.80nMAssay Description:Displacement of [125I]TZDM from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 5.90nMAssay Description:Displacement of [125I]TZDM from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
TargetAmyloid-beta precursor protein(Homo sapiens (Human))
Seoul National University Bundang Hospital
Curated by ChEMBL
Seoul National University Bundang Hospital
Curated by ChEMBL
Affinity DataKi: 26nMAssay Description:Displacement of [125I]TZDM from amyloid beta (1 to 42) aggregates in Alzheimer's disease-human brain homogenate after 3 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 114nM ΔG°: -39.6kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: 1.82E+3nM ΔG°: -32.8kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: 2.09E+3nM ΔG°: -32.4kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: 2.34E+3nM ΔG°: -32.1kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: 2.50E+3nM ΔG°: -32.0kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: 2.70E+3nM ΔG°: -31.8kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: 2.73E+3nM ΔG°: -31.8kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: 3.27E+3nM ΔG°: -31.3kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: 4.82E+3nM ΔG°: -30.3kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: 6.88E+3nM ΔG°: -29.5kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: 1.04E+4nM ΔG°: -28.4kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: 1.36E+4nM ΔG°: -27.8kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: 1.82E+4nM ΔG°: -27.1kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nM ΔG°: >-25.8kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nM ΔG°: >-25.8kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nM ΔG°: >-25.8kJ/molepH: 7.4 T: 2°CAssay Description:Fluorescence polarization experiments were conducted on a Photon Technology International instrument using a 0.3 cm path length cuvette. Spectra were...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 3.40nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 3.80nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 4.5nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 4.70nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 5.20nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.20nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.20nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.70nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.90nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 8.5nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 11.6nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 24nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 24nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair