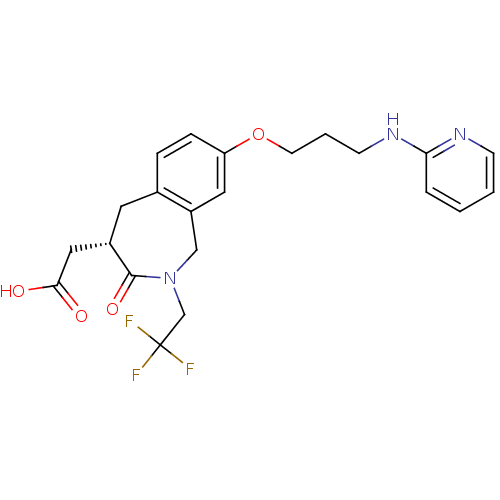
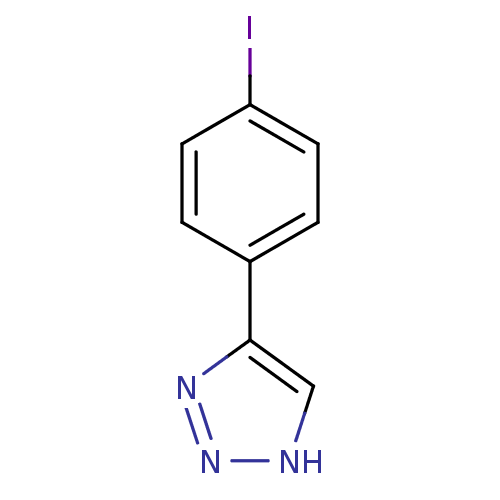
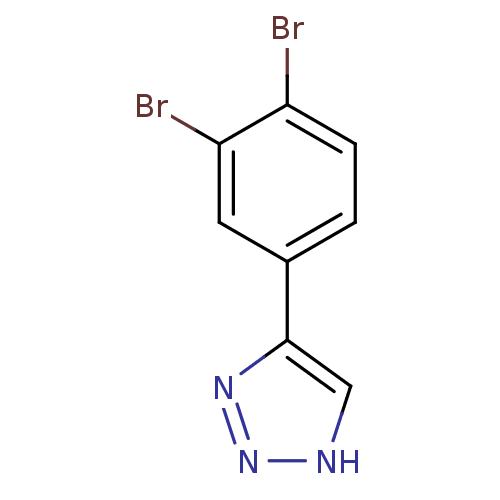
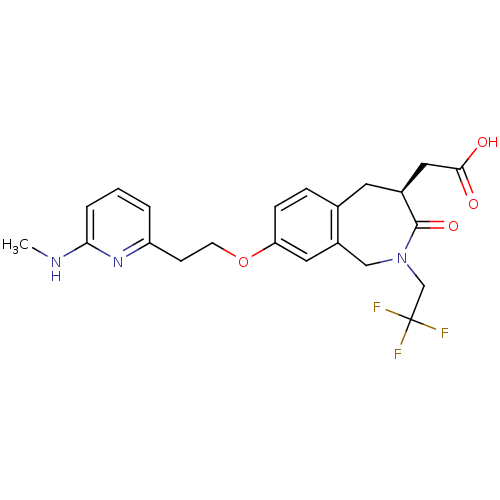
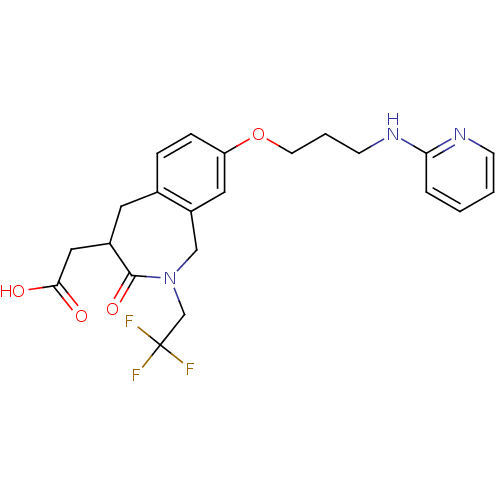
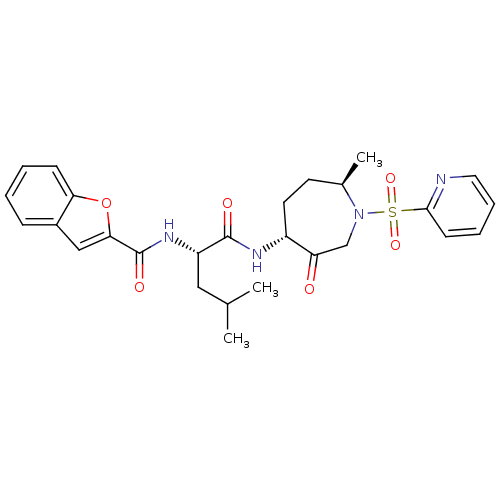
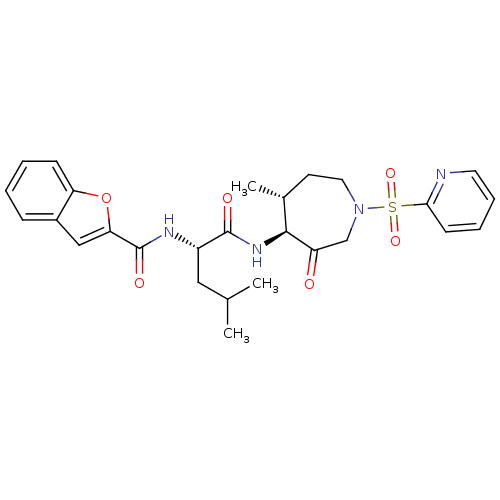
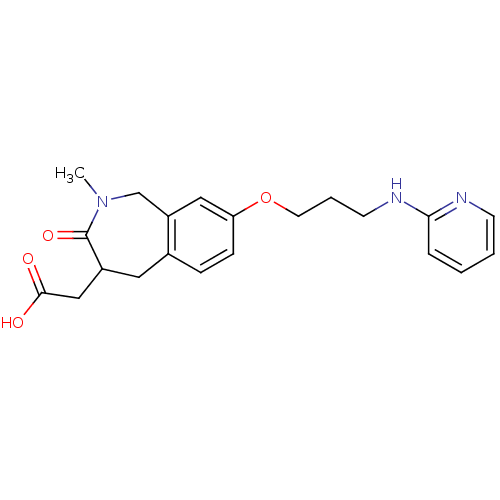
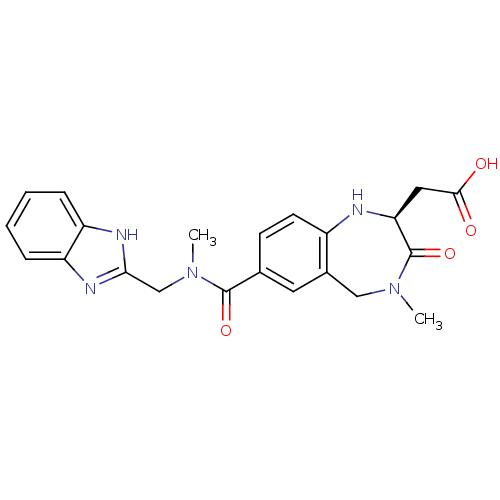
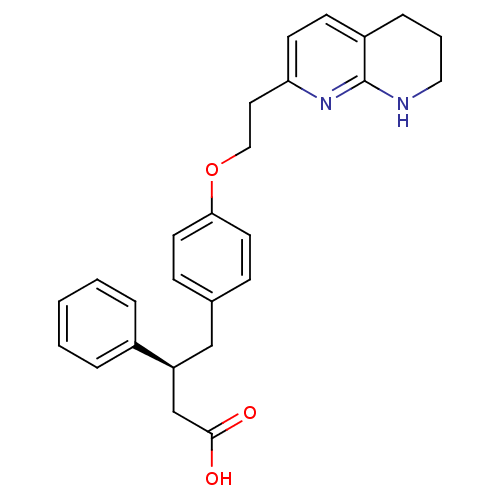
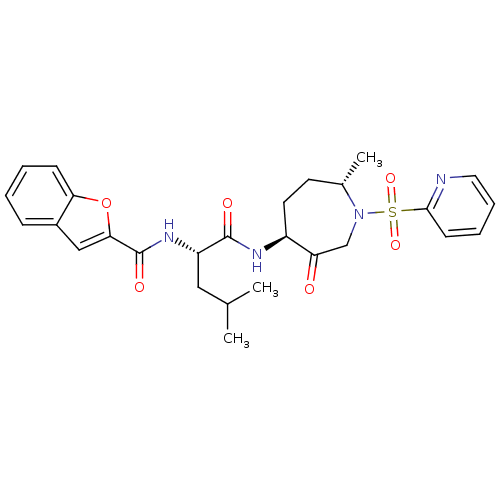
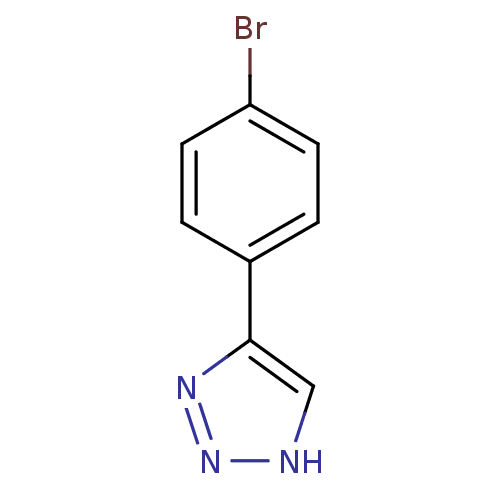
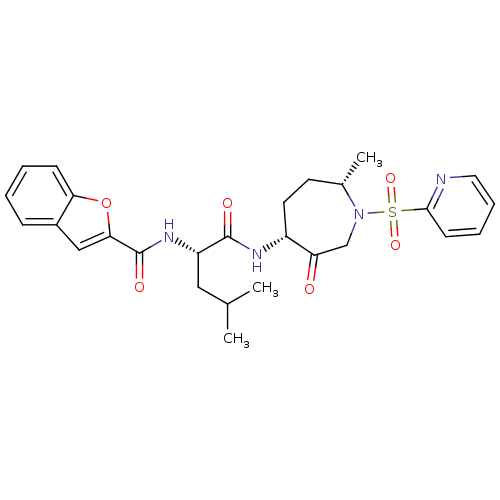
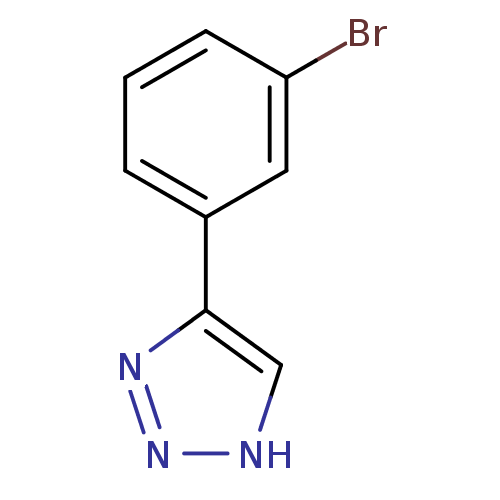
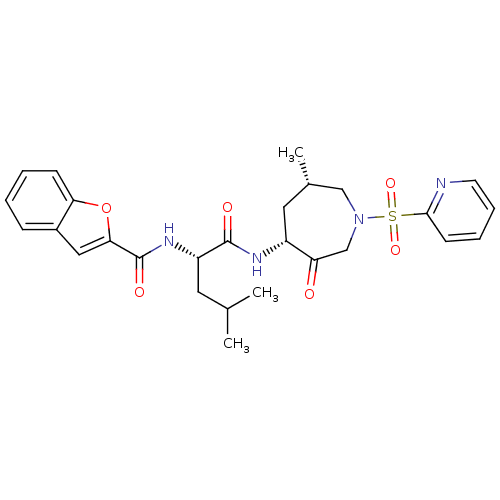
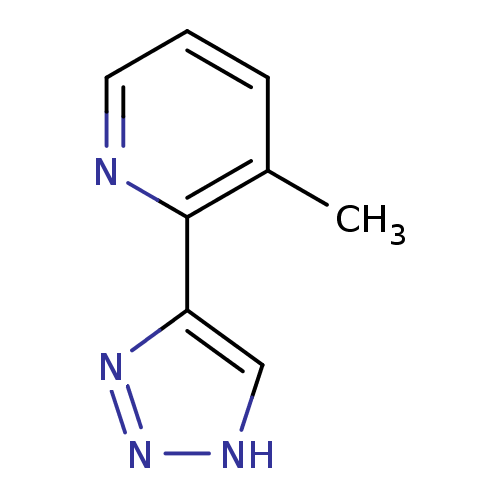
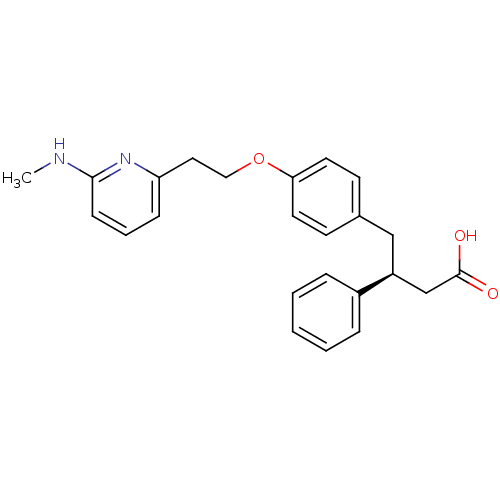
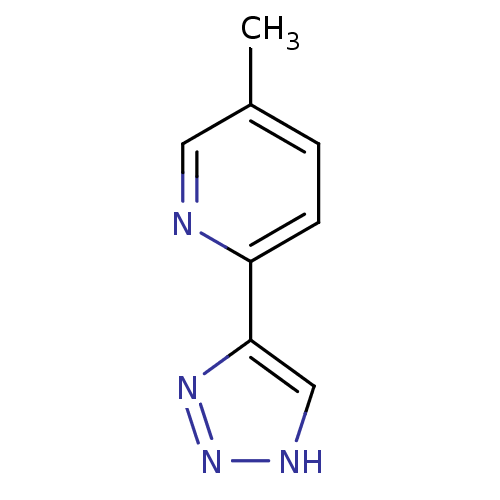
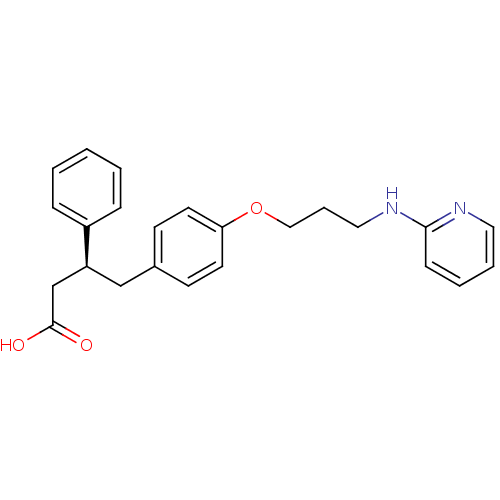
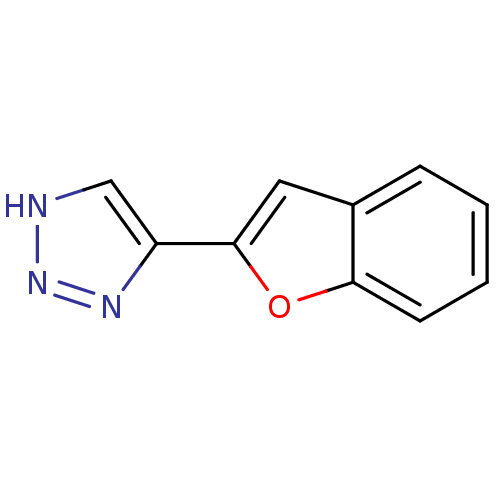
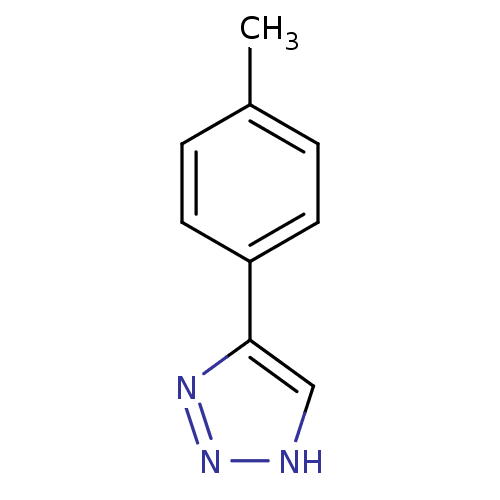
Affinity DataKi: 0.00990nM ΔG°: -62.2kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.0400nM ΔG°: -58.8kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.0410nM ΔG°: -58.7kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.0630nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.0680nM ΔG°: -57.4kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.140nM ΔG°: -55.7kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.160nM ΔG°: -55.3kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.630nM ΔG°: -52.0kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.900nMAssay Description:Binding affinity for Vitronectin receptor (alpha V beta 3)More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 1.20nMAssay Description:Binding affinity for Vitronectin receptor (alpha V beta 3)More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 1.30nMAssay Description:Binding affinity for Vitronectin receptor (alpha V beta 3)More data for this Ligand-Target Pair
Affinity DataKi: 1.40nM ΔG°: -50.0kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nM ΔG°: -49.9kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 1.90nMAssay Description:Binding affinity for Vitronectin receptor (alpha V beta 3)More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 2nMAssay Description:Binding affinity for Vitronectin receptor (alpha V beta 3)More data for this Ligand-Target Pair
Affinity DataKi: 2.20nM ΔG°: -48.9kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Binding affinity for alphav/beta 3 vitronectin receptor in HEK cellsMore data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 2.5nMAssay Description:Binding affinity for alphaV-beta5 vitronectin receptorMore data for this Ligand-Target Pair
Affinity DataKi: 2.5nM ΔG°: -48.6kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 2.60nMAssay Description:Binding affinity for alphaV-beta3 vitronectin receptorMore data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -48.2kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -48.2kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 3.90nM ΔG°: -47.5kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 4nM ΔG°: -47.5kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 4nMAssay Description:Binding affinity for alphaIIb-beta3 vitronectin receptorMore data for this Ligand-Target Pair
Affinity DataKi: 4nM ΔG°: -47.5kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 4nMAssay Description:Binding affinity for Vitronectin receptor (alpha V beta 3)More data for this Ligand-Target Pair
Affinity DataKi: 4.20nM ΔG°: -47.3kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 4.5nM ΔG°: -47.2kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 4.5nM ΔG°: -47.2kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 5.80nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Binding affinity for alphaV-beta3 vitronectin receptorMore data for this Ligand-Target Pair
Affinity DataKi: 6nM ΔG°: -46.5kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 6nM ΔG°: -46.5kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 7nMAssay Description:Binding affinity for alphaV-beta3 vitronectin receptorMore data for this Ligand-Target Pair
Affinity DataKi: 7.90nM ΔG°: -45.8kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 8nM ΔG°: -45.7kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 8.30nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -45.2kJ/molepH: 5.5 T: 2°CAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
TargetIntegrin alpha-V/beta-3(Homo sapiens (Human))
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Smithkline Beecham Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 12nMAssay Description:Binding affinity for alphav/beta 3 vitronectin receptor in HEK cellsMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 13nM ΔG°: -44.6kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 15nM ΔG°: -44.2kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair
Affinity DataKi: 16nM ΔG°: -44.0kJ/molepH: 7.5 T: 2°CAssay Description:MetAP2 activity was monitored by measuring the initial velocity of turnover of the artificial substrate Met-AMC. Assays were performed in 96-well mi...More data for this Ligand-Target Pair

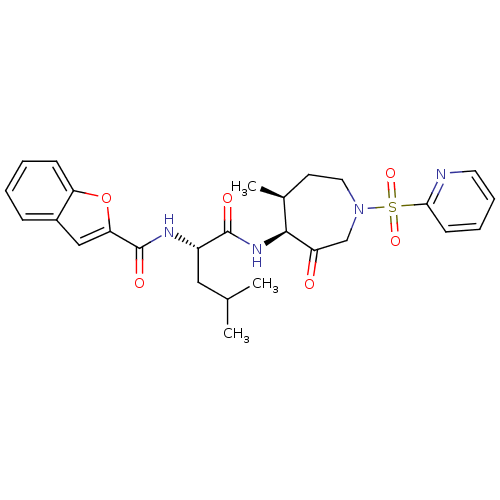
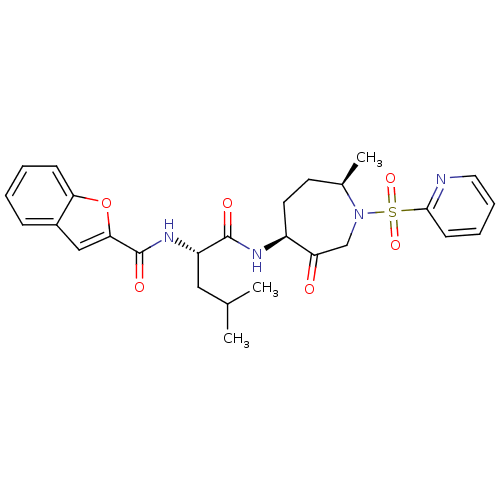
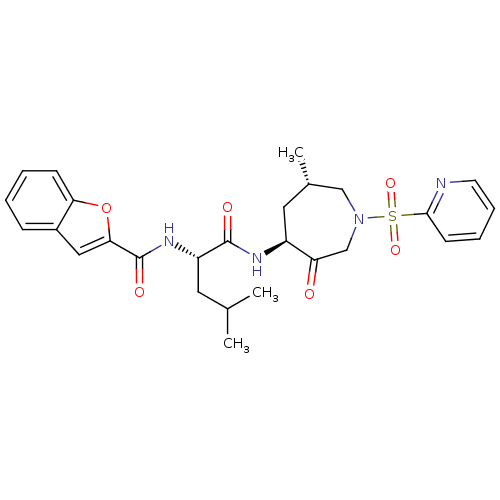
 3D Structure (crystal)
3D Structure (crystal)