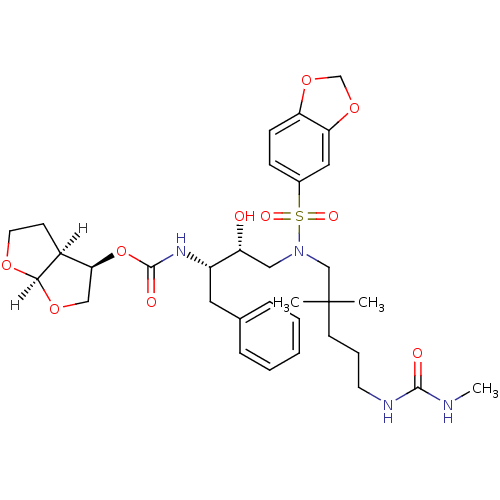
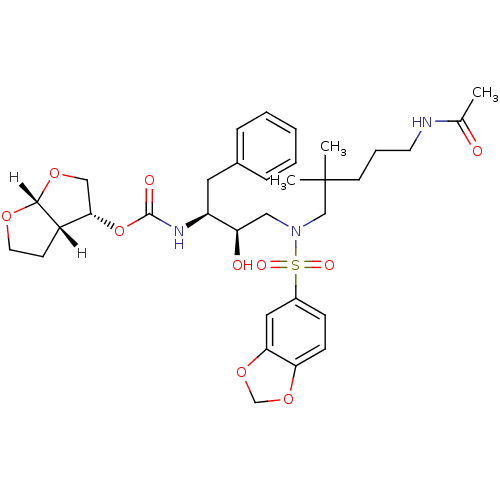
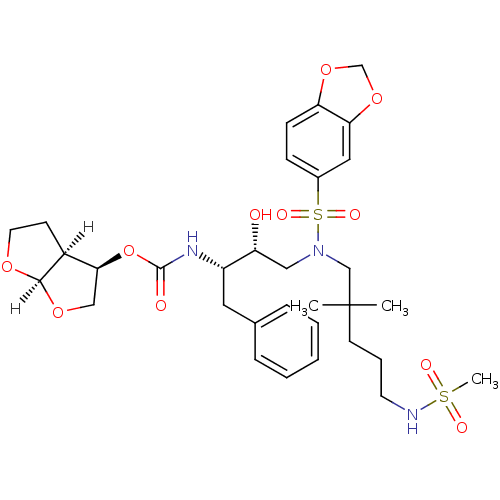
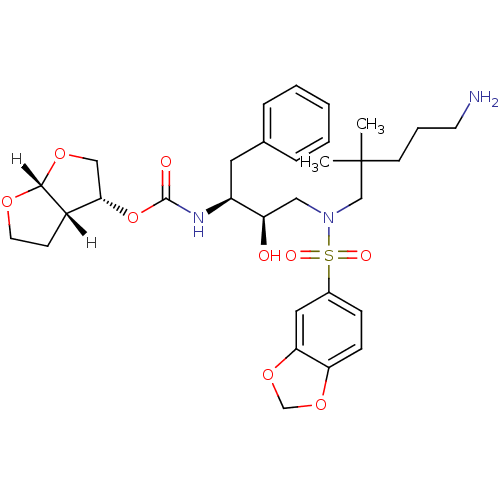
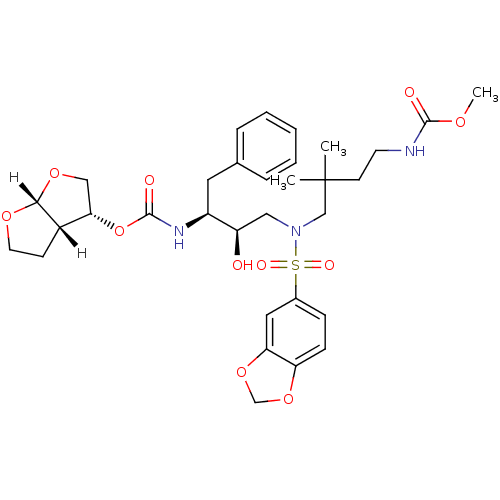
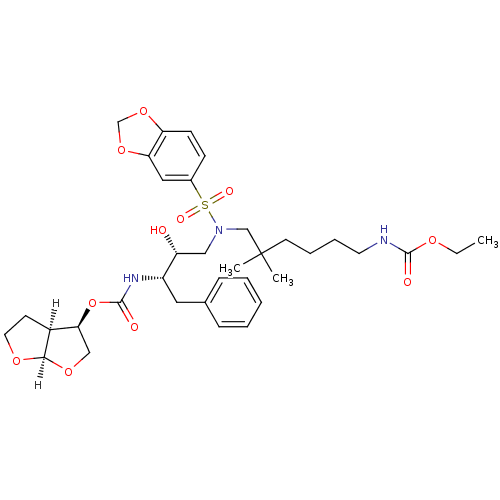
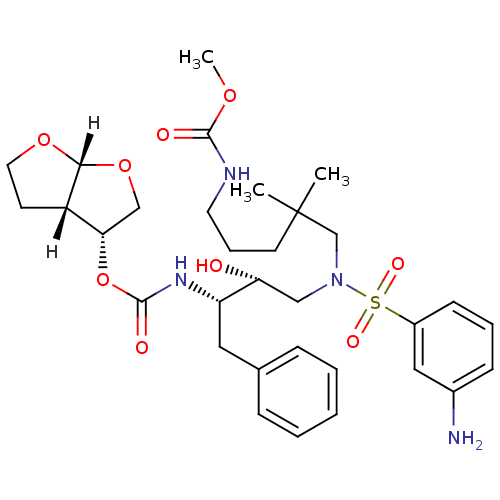
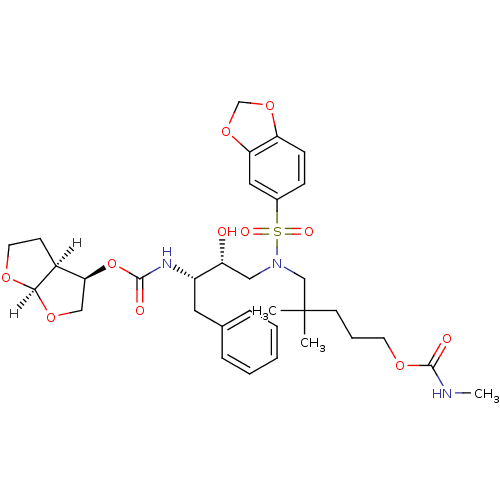
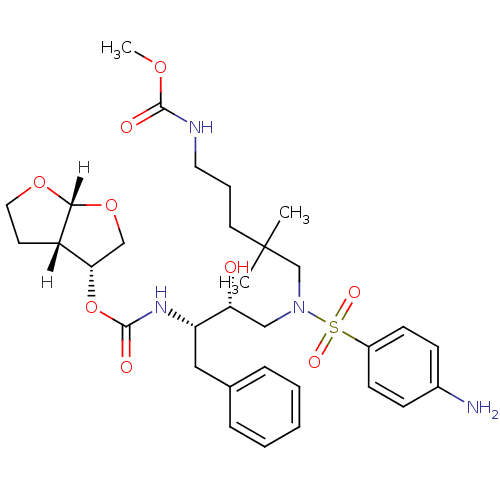
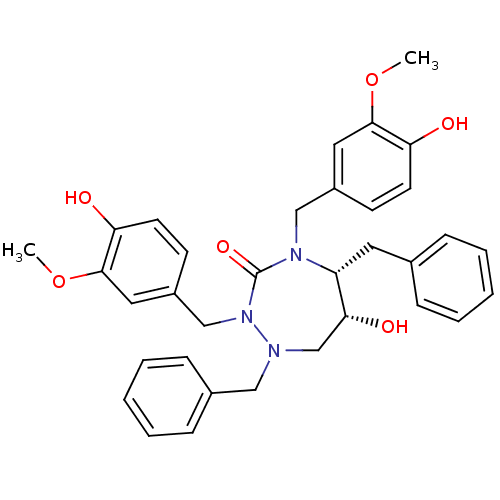
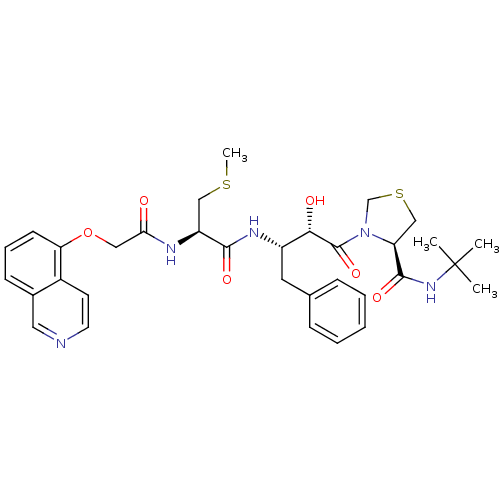
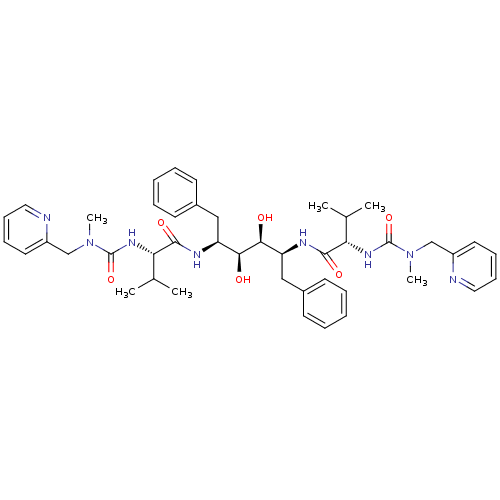
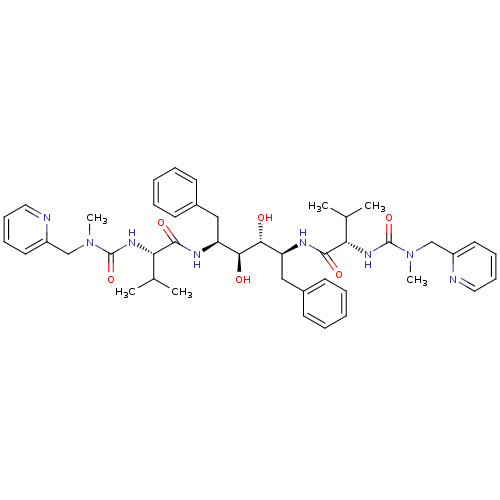
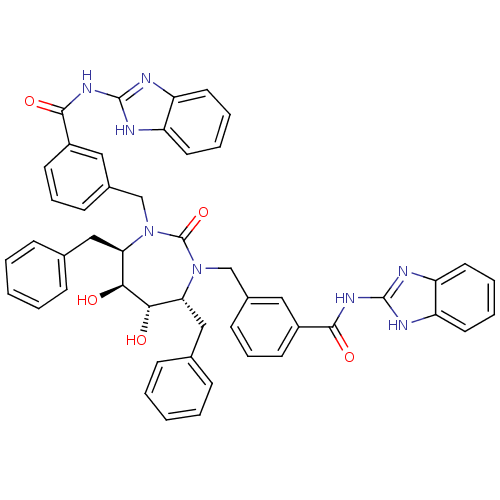
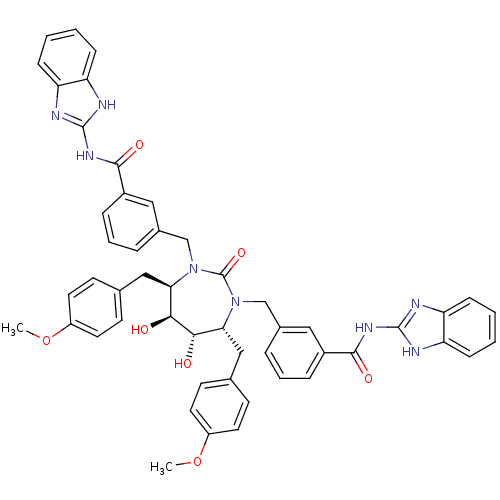
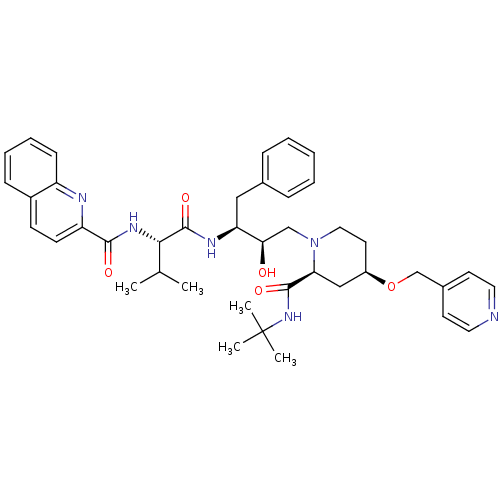
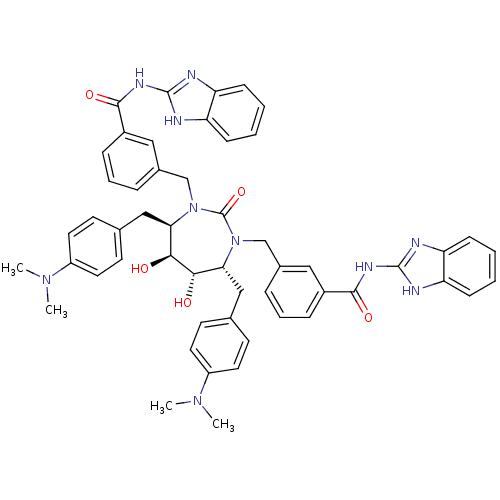
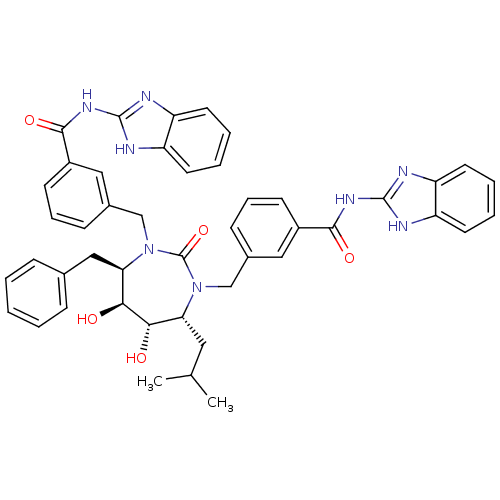
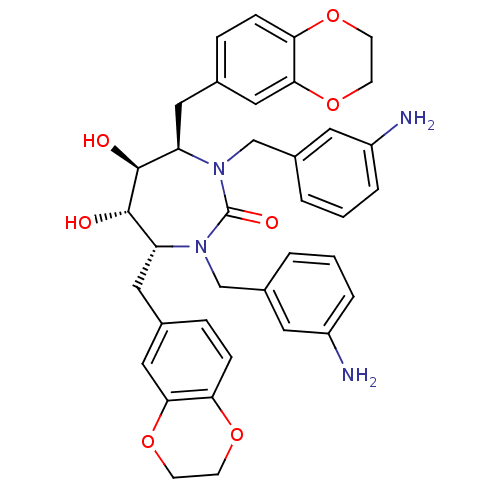
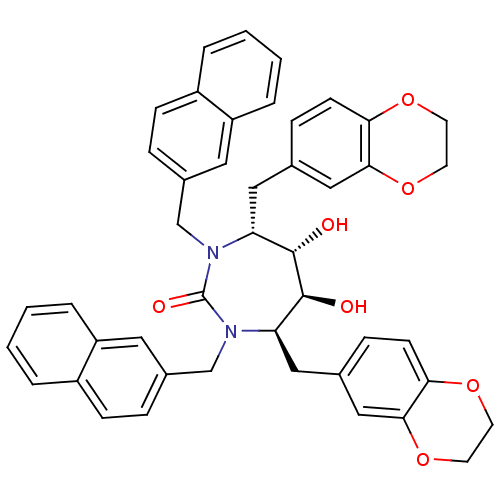
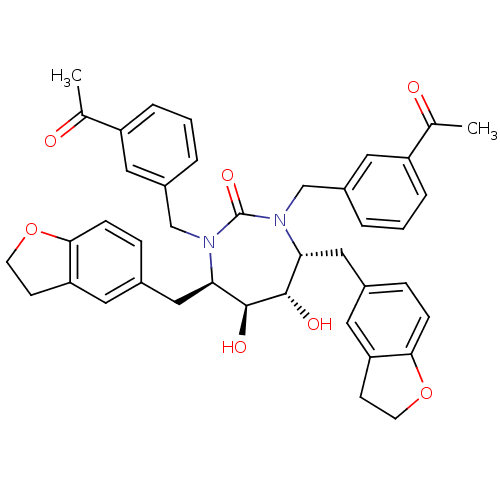
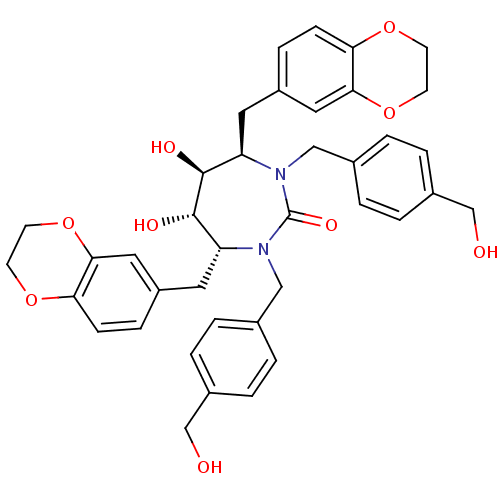
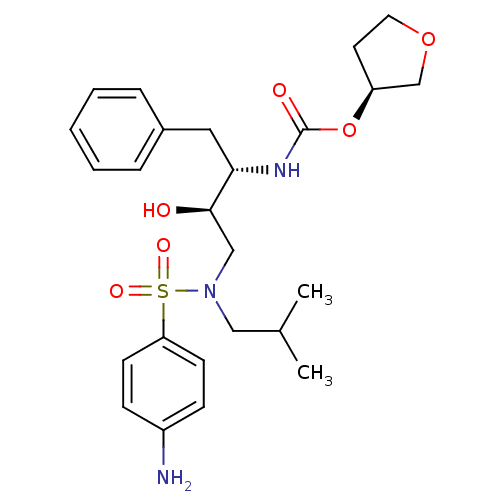
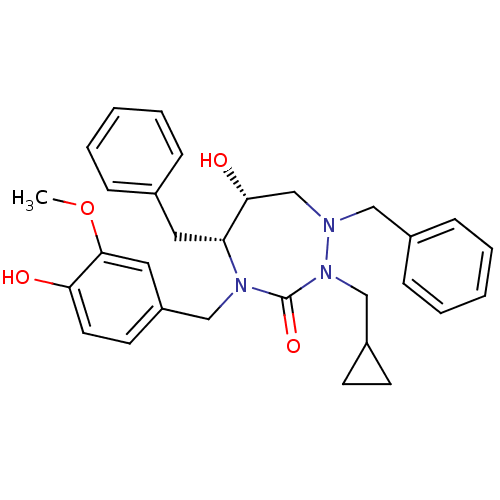
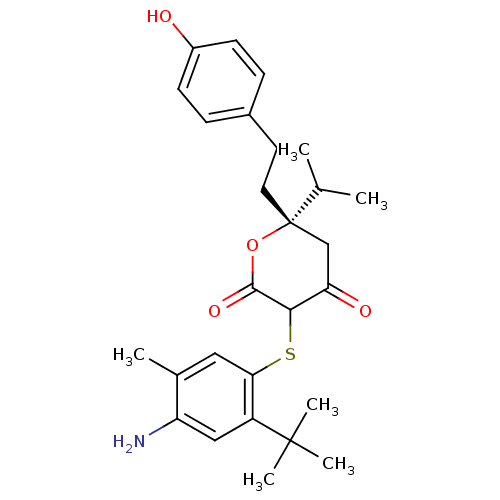
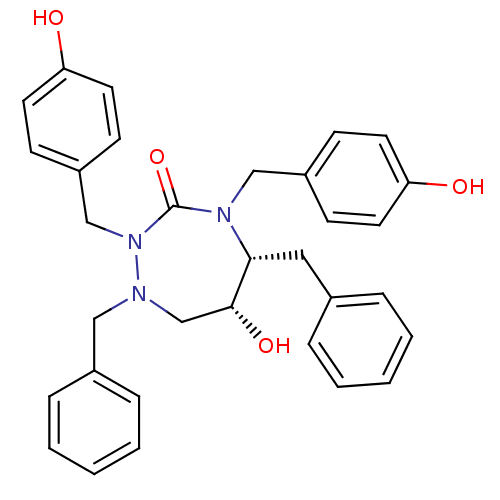
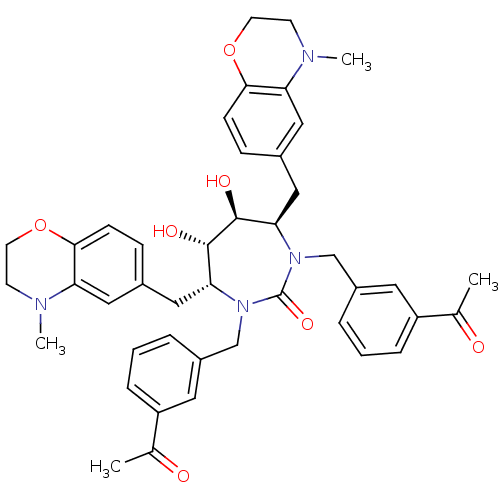
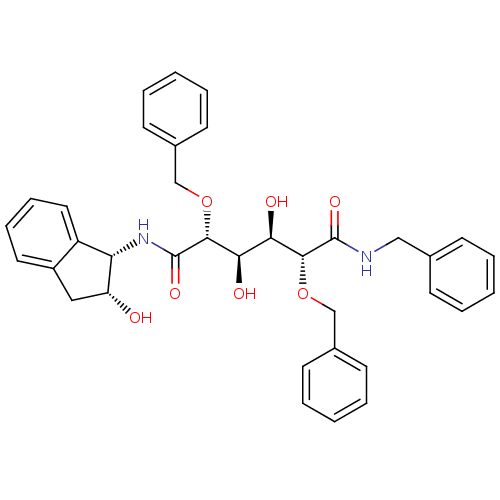
Affinity DataKi: 0.00000600nM ΔG°: -19.7kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
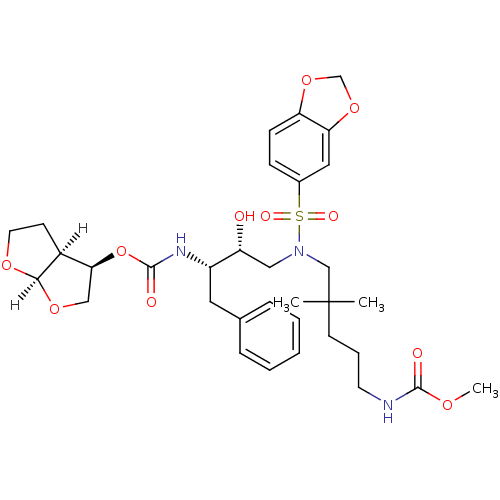
Affinity DataKi: 0.0000130nM ΔG°: -19.2kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
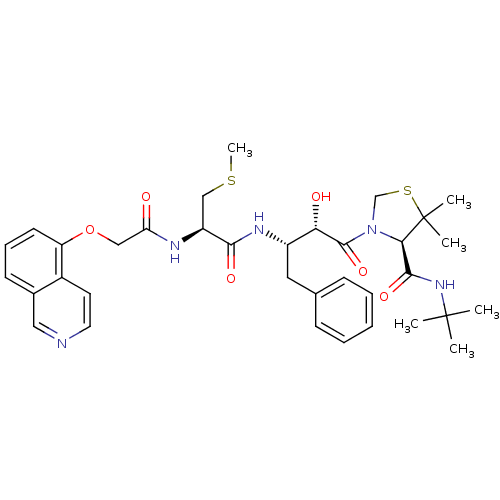
Affinity DataKi: 0.0000140nM ΔG°: -19.2kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
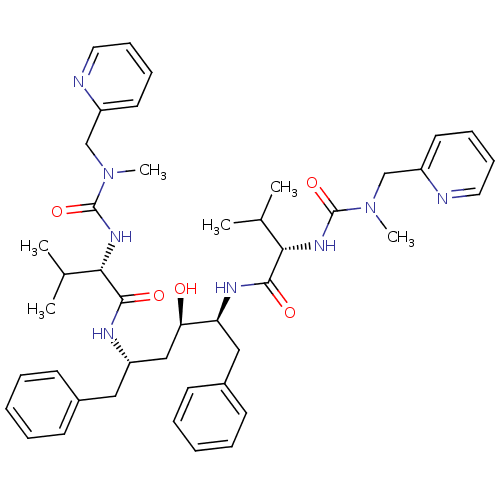
Affinity DataKi: 0.0000150nM ΔG°: -19.2kcal/molepH: 5.5 T: 2°CAssay Description:The assay method employed kinetic determinations of values for k1 and k-1, from which value of inhibition constant (Ki ) was determined (k-1/k1). The...More data for this Ligand-Target Pair
Affinity DataKi: 0.0000480nM ΔG°: -18.5kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
Affinity DataKi: 0.0000540nM ΔG°: -18.4kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
Affinity DataKi: 0.0000580nM ΔG°: -18.3kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
Affinity DataKi: 0.0000610nM ΔG°: -18.3kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
Affinity DataKi: 0.000120nM ΔG°: -17.9kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
Affinity DataKi: 0.000160nM ΔG°: -17.7kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
Affinity DataKi: 0.000165nM ΔG°: -17.7kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
Affinity DataKi: 0.000210nM ΔG°: -17.6kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
Affinity DataKi: 0.000240nM ΔG°: -17.5kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
Affinity DataKi: 0.000260nM ΔG°: -17.4kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
Affinity DataKi: 0.000520nM ΔG°: -17.0kcal/molepH: 5.5 T: 2°CAssay Description:The method was a competitive displacement assay used to determine binding affinities of other inhibitors relative to that of GW0385. The inhibitor of...More data for this Ligand-Target Pair
Affinity DataKi: 0.00230nM ΔG°: -16.5kcal/molepH: 6.2 T: 2°CAssay Description:Inhibition constants were determined by a fluorometric assay with the fluorogenic substrate Arg-Glu(EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Lys(DABCYL)-Ar...More data for this Ligand-Target Pair
Affinity DataKi: 0.00400nM ΔG°: -15.8kcal/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease activity was measured by a continuous fluorometric assay using the internally quenched fluorogenic substrate DABCYL-GABA-Ser-Gln-Tyr-P...More data for this Ligand-Target Pair
Affinity DataKi: 0.00500nM ΔG°: -15.7kcal/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease activity was measured by a continuous fluorometric assay using the internally quenched fluorogenic substrate DABCYL-GABA-Ser-Gln-Tyr-P...More data for this Ligand-Target Pair
Affinity DataKi: 0.00550nM ΔG°: -16.0kcal/molepH: 6.2 T: 2°CAssay Description:Inhibition constants were determined by a fluorometric assay with the fluorogenic substrate Arg-Glu(EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Lys(DABCYL)-Ar...More data for this Ligand-Target Pair
Affinity DataKi: 0.00680nM ΔG°: -15.8kcal/molepH: 6.2 T: 2°CAssay Description:Inhibition constants were determined by a fluorometric assay with the fluorogenic substrate Arg-Glu(EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Lys(DABCYL)-Ar...More data for this Ligand-Target Pair
Affinity DataKi: 0.0110nM ΔG°: -15.2kcal/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease activity was measured by a continuous fluorometric assay using the internally quenched fluorogenic substrate DABCYL-GABA-Ser-Gln-Tyr-P...More data for this Ligand-Target Pair
Affinity DataKi: 0.0120nM ΔG°: -15.1kcal/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease activity was measured by a continuous fluorometric assay using the internally quenched fluorogenic substrate DABCYL-GABA-Ser-Gln-Tyr-P...More data for this Ligand-Target Pair
Affinity DataKi: 0.0160nM ΔG°: -15.3kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0240nM ΔG°: -15.1kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0260nM ΔG°: -15.0kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0300nMAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
Affinity DataKi: 0.0300nMAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
Affinity DataKi: 0.0300nM ΔG°: -14.9kcal/molepH: 6.2 T: 2°CAssay Description:For determination of IC50 values, HIV-1 protease was added to assay buffer containing inhibitor and the substrate (H-His-Lys-Ala-Arg-Val-Leu- (p-NO2)...More data for this Ligand-Target Pair
Affinity DataKi: 0.0300nM ΔG°: -14.9kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0310nM ΔG°: -14.9kcal/mole IC50: 4nMpH: 5.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Cleavage products and substrate w...More data for this Ligand-Target Pair
Affinity DataKi: 0.0350nM ΔG°: -14.8kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0380nM ΔG°: -14.8kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0380nM ΔG°: -14.8kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0400nM ΔG°: -14.7kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0400nM ΔG°: -14.7kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0400nM ΔG°: -14.7kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0430nM ΔG°: -14.7kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0460nM ΔG°: -14.7kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0480nM ΔG°: -14.6kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nM ΔG°: -14.6kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0570nM ΔG°: -14.2kcal/molepH: 5.5 T: 2°CAssay Description:Enzymatic activity was determined with fluorogenic substrate in the presence and absence of inhibitor. The fluorescence increase due to hydrolysis of...More data for this Ligand-Target Pair
Affinity DataKi: 0.0620nM ΔG°: -14.1kcal/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease activity was measured by a continuous fluorometric assay using the internally quenched fluorogenic substrate DABCYL-GABA-Ser-Gln-Tyr-P...More data for this Ligand-Target Pair
Affinity DataKi: 0.0690nM ΔG°: -14.4kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0700nMAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
Affinity DataKi: 0.0700nM ΔG°: -14.1kcal/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease activity was measured by a continuous fluorometric assay using the internally quenched fluorogenic substrate DABCYL-GABA-Ser-Gln-Tyr-P...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nM ΔG°: -14.3kcal/molepH: 5.5 T: 2°CAssay Description:Inhibition of HIV protease was measured by assay of the cleavage of a fluorescent peptide substrate. The fluorescent product (2-aminobenzoyl-Ala-Thr-...More data for this Ligand-Target Pair
Affinity DataKi: 0.0880nM ΔG°: -14.3kcal/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 0.0900nMAssay Description:For determination of IC50 values, HIV-1 protease was added to assay buffer containing inhibitor and the substrate (H-His-Lys-Ala-Arg-Val-Leu- (p-NO2)...More data for this Ligand-Target Pair
Affinity DataKi: 0.0900nMAssay Description:For determination of IC50 values, HIV-1 protease was added to assay buffer containing inhibitor and the substrate (H-His-Lys-Ala-Arg-Val-Leu- (p-NO2)...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -13.9kcal/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair

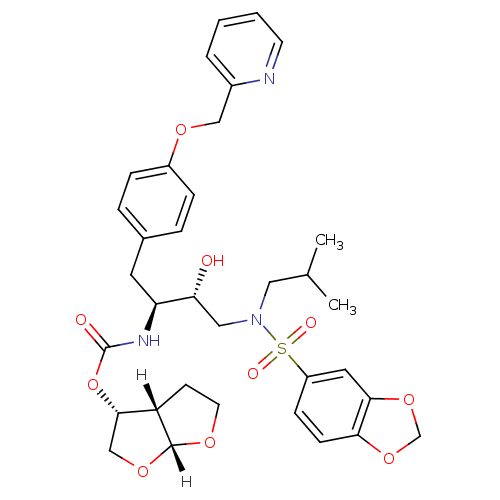
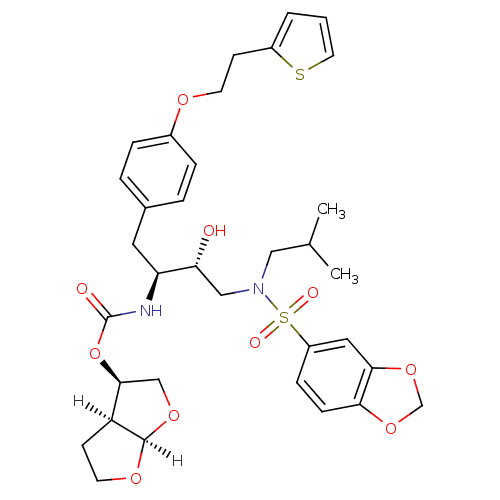
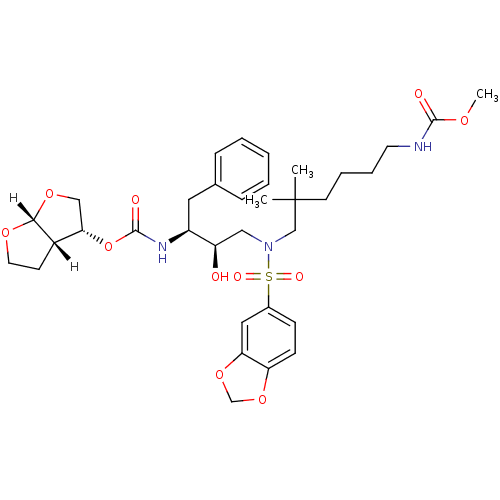
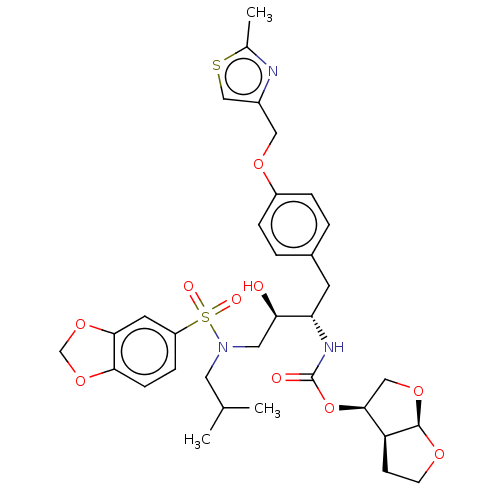
 3D Structure (crystal)
3D Structure (crystal)