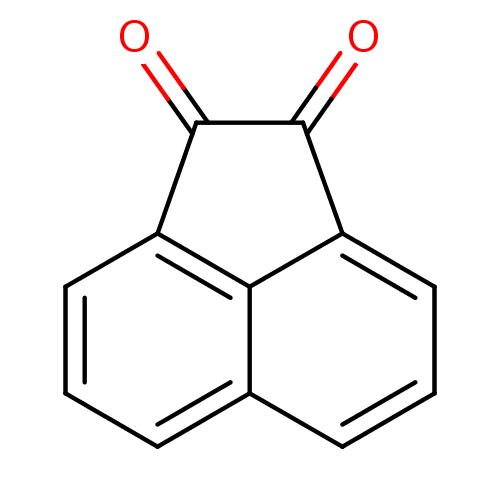
Affinity DataKi: 31nM ΔG°: -42.9kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:Inhibition of human CE1 using o-NPA as substrate by spectrophotometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 170nM ΔG°: -38.6kJ/molepH: 7.4 T: 2°CAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 170nMAssay Description:Inhibition of human iCE using o-NPA as substrate by spectrophotometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.97E+3nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Estimates of the competitive inhibition constants (Ki) were ob...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Dalian University Of Technology
Curated by ChEMBL
Dalian University Of Technology
Curated by ChEMBL
Affinity DataKi: 5.70E+4nMAssay Description:Inhibition of FAM-Bid peptide binding to Mcl1 (unknown origin) expressed in Escherichia coli BL21 by fluorescence polarization-based binding assayMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Estimates of the competitive inhibition constants (Ki) were ob...More data for this Ligand-Target Pair