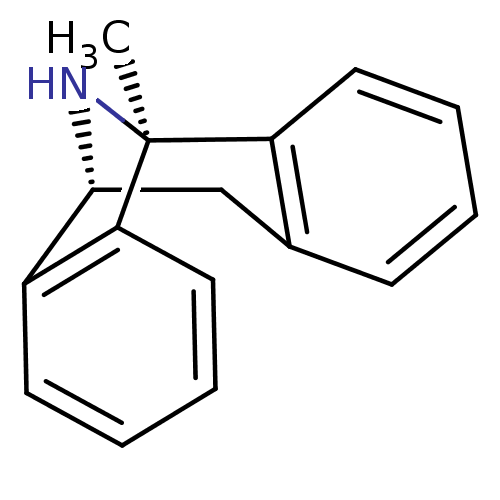TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 9nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Antagonist activity at NR2A transfected in oocytesMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 31nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 31nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
Affinity DataEC50: 91nMAssay Description:Agonist activity at rat GluN1/GluN2D NMDA receptor expressed in Xenopus oocytes measured after 4 days in presence of L-glutamte by two-electrode volt...More data for this Ligand-Target Pair
Affinity DataEC50: 210nMAssay Description:Agonist activity at rat GluN1/GluN2C NMDA receptor expressed in Xenopus oocytes measured after 4 days in presence of L-glutamte by two-electrode volt...More data for this Ligand-Target Pair
Affinity DataEC50: 240nMAssay Description:Agonist activity at rat GluN1/GluN2B NMDA receptor expressed in Xenopus oocytes measured after 4 days in presence of L-glutamte by two-electrode volt...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataIC50: 250nMAssay Description:In vitro inhibitory activity to inhibit [3H]glycine binding to NMDA receptorMore data for this Ligand-Target Pair
Affinity DataEC50: 990nMAssay Description:Agonist activity at rat GluN1/GluN2A NMDA receptor expressed in Xenopus oocytes measured after 4 days in presence of L-glutamte by two-electrode volt...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataIC50: 6.20E+3nMAssay Description:Inhibitory concentration required to inhibit [3H]strychnine binding to N-methyl-D-aspartate glutamate receptor 1 of rat spinal cord membranes.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
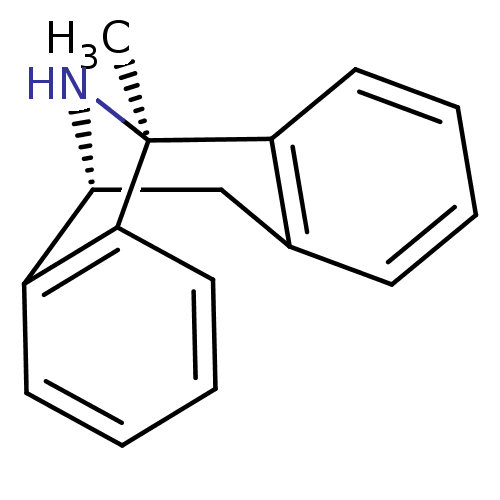
 3D Structure (crystal)
3D Structure (crystal)