Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Bromodomain-containing protein 4
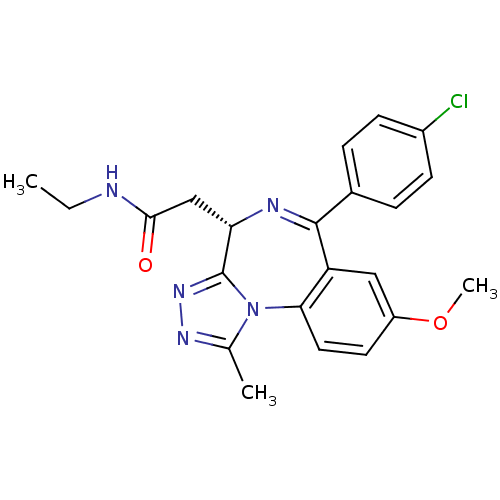
Ligand
BDBM50365463
Substrate
n/a
Meas. Tech.
ChEMBL_1832478 (CHEMBL4332486)
Ki
200±n/a nM
Citation
 Sperandio, D; Aktoudianakis, V; Babaoglu, K; Chen, X; Elbel, K; Chin, G; Corkey, B; Du, J; Jiang, B; Kobayashi, T; Mackman, R; Martinez, R; Yang, H; Zablocki, J; Kusam, S; Jordan, K; Webb, H; Bates, JG; Lad, L; Mish, M; Niedziela-Majka, A; Metobo, S; Sapre, A; Hung, M; Jin, D; Fung, W; Kan, E; Eisenberg, G; Larson, N; Newby, ZER; Lansdon, E; Tay, C; Neve, RM; Shevick, SL; Breckenridge, DG Structure-guided discovery of a novel, potent, and orally bioavailable 3,5-dimethylisoxazole aryl-benzimidazole BET bromodomain inhibitor. Bioorg Med Chem 27:457-469 (2019) [PubMed] Article
Sperandio, D; Aktoudianakis, V; Babaoglu, K; Chen, X; Elbel, K; Chin, G; Corkey, B; Du, J; Jiang, B; Kobayashi, T; Mackman, R; Martinez, R; Yang, H; Zablocki, J; Kusam, S; Jordan, K; Webb, H; Bates, JG; Lad, L; Mish, M; Niedziela-Majka, A; Metobo, S; Sapre, A; Hung, M; Jin, D; Fung, W; Kan, E; Eisenberg, G; Larson, N; Newby, ZER; Lansdon, E; Tay, C; Neve, RM; Shevick, SL; Breckenridge, DG Structure-guided discovery of a novel, potent, and orally bioavailable 3,5-dimethylisoxazole aryl-benzimidazole BET bromodomain inhibitor. Bioorg Med Chem 27:457-469 (2019) [PubMed] Article More Info.:
Target
Name:
Bromodomain-containing protein 4
Synonyms:
BRD4 | BRD4_HUMAN | Bromodomain-containing protein 4 (BRD4) | HUNK1 | Protein HUNK1
Type:
Protein
Mol. Mass.:
152264.84
Organism:
Homo sapiens (Human)
Description:
O60885
Residue:
1362
Sequence:
MSAESGPGTRLRNLPVMGDGLETSQMSTTQAQAQPQPANAASTNPPPPETSNPNKPKRQTNQLQYLLRVVLKTLWKHQFAWPFQQPVDAVKLNLPDYYKIIKTPMDMGTIKKRLENNYYWNAQECIQDFNTMFTNCYIYNKPGDDIVLMAEALEKLFLQKINELPTEETEIMIVQAKGRGRGRKETGTAKPGVSTVPNTTQASTPPQTQTPQPNPPPVQATPHPFPAVTPDLIVQTPVMTVVPPQPLQTPPPVPPQPQPPPAPAPQPVQSHPPIIAATPQPVKTKKGVKRKADTTTPTTIDPIHEPPSLPPEPKTTKLGQRRESSRPVKPPKKDVPDSQQHPAPEKSSKVSEQLKCCSGILKEMFAKKHAAYAWPFYKPVDVEALGLHDYCDIIKHPMDMSTIKSKLEAREYRDAQEFGADVRLMFSNCYKYNPPDHEVVAMARKLQDVFEMRFAKMPDEPEEPVVAVSSPAVPPPTKVVAPPSSSDSSSDSSSDSDSSTDDSEEERAQRLAELQEQLKAVHEQLAALSQPQQNKPKKKEKDKKEKKKEKHKRKEEVEENKKSKAKEPPPKKTKKNNSSNSNVSKKEPAPMKSKPPPTYESEEEDKCKPMSYEEKRQLSLDINKLPGEKLGRVVHIIQSREPSLKNSNPDEIEIDFETLKPSTLRELERYVTSCLRKKRKPQAEKVDVIAGSSKMKGFSSSESESSSESSSSDSEDSETEMAPKSKKKGHPGREQKKHHHHHHQQMQQAPAPVPQQPPPPPQQPPPPPPPQQQQQPPPPPPPPSMPQQAAPAMKSSPPPFIATQVPVLEPQLPGSVFDPIGHFTQPILHLPQPELPPHLPQPPEHSTPPHLNQHAVVSPPALHNALPQQPSRPSNRAAALPPKPARPPAVSPALTQTPLLPQPPMAQPPQVLLEDEEPPAPPLTSMQMQLYLQQLQKVQPPTPLLPSVKVQSQPPPPLPPPPHPSVQQQLQQQPPPPPPPQPQPPPQQQHQPPPRPVHLQPMQFSTHIQQPPPPQGQQPPHPPPGQQPPPPQPAKPQQVIQHHHSPRHHKSDPYSTGHLREAPSPLMIHSPQMSQFQSLTHQSPPQQNVQPKKQELRAASVVQPQPLVVVKEEKIHSPIIRSEPFSPSLRPEPPKHPESIKAPVHLPQRPEMKPVDVGRPVIRPPEQNAPPPGAPDKDKQKQEPKTPVAPKKDLKIKNMGSWASLVQKHPTTPSSTAKSSSDSFEQFRRAAREKEEREKALKAQAEHAEKEKERLRQERMRSREDEDALEQARRAHEEARRRQEQQQQQRQEQQQQQQQQAAAVAAAATPQAQSSQPQSMLDQQRELARKREQERRRREAMAATIDMNFQSDLLSIFEENLF