Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Glycogen debranching enzyme
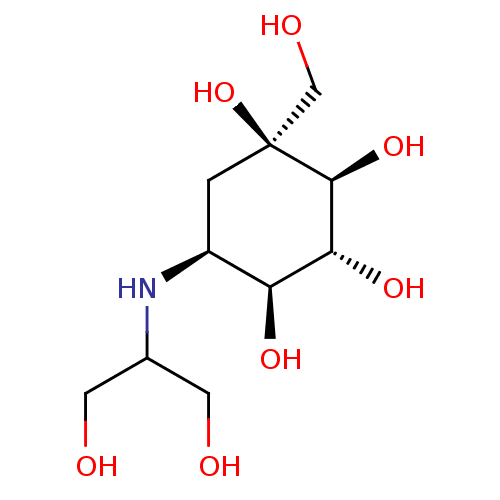
Ligand
BDBM50263044
Substrate
n/a
Meas. Tech.
ChEMBL_490834 (CHEMBL993573)
IC50
70000±n/a nM
Citation
 Kuriyama, C; Kamiyama, O; Ikeda, K; Sanae, F; Kato, A; Adachi, I; Imahori, T; Takahata, H; Okamoto, T; Asano, N In vitro inhibition of glycogen-degrading enzymes and glycosidases by six-membered sugar mimics and their evaluation in cell cultures. Bioorg Med Chem 16:7330-6 (2008) [PubMed] Article
Kuriyama, C; Kamiyama, O; Ikeda, K; Sanae, F; Kato, A; Adachi, I; Imahori, T; Takahata, H; Okamoto, T; Asano, N In vitro inhibition of glycogen-degrading enzymes and glycosidases by six-membered sugar mimics and their evaluation in cell cultures. Bioorg Med Chem 16:7330-6 (2008) [PubMed] Article More Info.:
Target
Name:
Glycogen debranching enzyme
Synonyms:
AGL | GDE_RABIT
Type:
PROTEIN
Mol. Mass.:
177662.68
Organism:
Oryctolagus cuniculus
Description:
ChEMBL_490834
Residue:
1555
Sequence:
MGNSFDFGVLLILLKYFKSSRSQNGHSKQIRILLLNEMEKLEKTLFRLEQGFELQFRLGPTLQGKPVTVFTNYPFPGETFNREKFRSLEWENPTEREDDSDKYCKLNLQQSGSFQYYFLQGNEKSGGGYIVVDPILRVGADNHMLHLDCVTLQTFLAKCLGPFDEWESRLRVAKESGYNMIHFTPLQTLGLSRSCYSLADQLELNPDFSRPHKKYTWSDVGQLVEKLKREWNVLCITDVVYNHTAANSKWIQEHPECAYNLVNSPHLKPAWVLDRALWHFSCDVAEGKYKNRGVPALIENDHHLNCIRKVIWEDIFPKLHLWEFFQVDVYKAVEKFRGLLTQETWRVIKSDPKQHLKIIQDPEYRRFGCTVDMNIALATFIPHDNGPAAIEECCNWFRKRIEELNSEKHQLMNYHQEQAVNCLLGNVFYERLAGHGPKLGPVTRKYPLVTRYFTFPFEEMPVSTEETMIHLPNKACFFMAHNGWVMGDDPLRNFAEPGSDVYLRRELICWGDSVKLRYGTKPEDCPYLWAHMRKYTEIIATYFQGVRLDNCHSTPLHVAEYMLDAARKLQPNLYVVAELFTGSEDLDNIFVTRLGISSLIREAMSAYNSHEEGRLVYRYGGEPVGSFVQPCLRPLMPAIAHALFMDITHDNECPIVHRSVYDALPSTTIVSMACCASGSTRGYDELVPHQISVVSEERFYTKWNPEALPSNAGEVNFQSGIIAARCAINKLHQELGAKGFIQVYVDQVDEDIVAVTRHSPSIHQSFVAVSRTAFRNPKTSFYSKDVPQMCIPGKIEEVVLEARTIERNISPYRKDENSINGMPNITVEIREHIQLNESRIVKQAGVTTKGPNEYIQEIEFENLSPGSVIIFRVSLDPHAQVAVGILRNHLTQFSAHFKAGSLAVDNSDPILKIPFASIASKLTLAEINQILYRCESEEQEDGGGCYDIPNWSSLKYAGLQGLMSVLAEIRPKNDLGHPFCDNLRSGDWMIDYVSGRLISRSGTIAEVGKWLQAMFFYLKQIPRYLIPCYFDAILIGAYTTLLDIAWKQMSSFVQTGSTFVKHLSLGSVQMCGVGKFPSLPLLSPSLTDVPYRLNEITKEKEQCCVSLAAGLPHFSSGIFRCWGRDTFIALRGLLLITGRYLEARNIILAFAGTLRHGLIPNLLGEGTYARYNCRDAVWWWLQCIQDYCKMVPNGLDILKCPVSRMYPTDDSAPLPAGTLDQPLFDVIQEAMQRHMQGIQFRERNAGPQIDRNMKDEGFTVIAGVNEETGFVYGGNRFNCGTWMDKMGESDRARNRGIPATPRDGSAVEIVGLCKSTVRWLLELSKKNIFPYHEVRVKRHGKVVTVSYEEWNRKIQDNFEKRFHVSEDPSASNEEHPNLVHKRGIYKDSYGASSPWCDYQLRPNFTIAMVVAPELFTAEKAWKALEIAEKKLLGPLGMKTLDPDDMVYCGIYDNALDNDNYNLAKGFNYHQGPEWLWPVGYFLRAKLYFSKLMDRETNARTIFLVKNVLSRHYVHLERSPWKGLPELTNENGQYCPFSCETQAWSIATILETLYDL