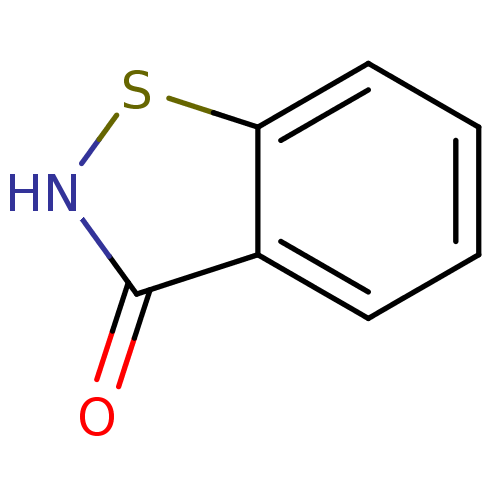
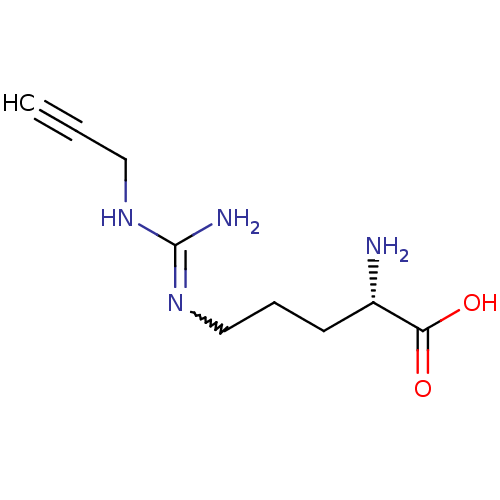
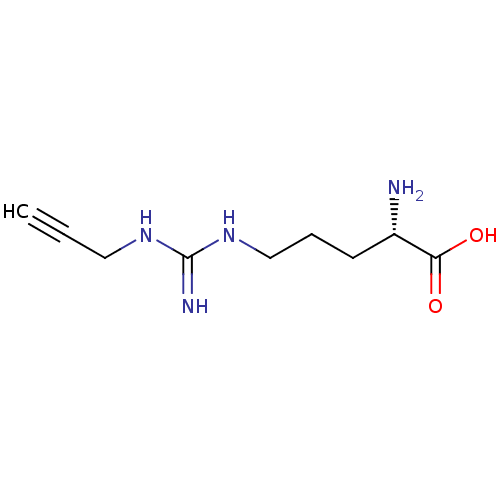
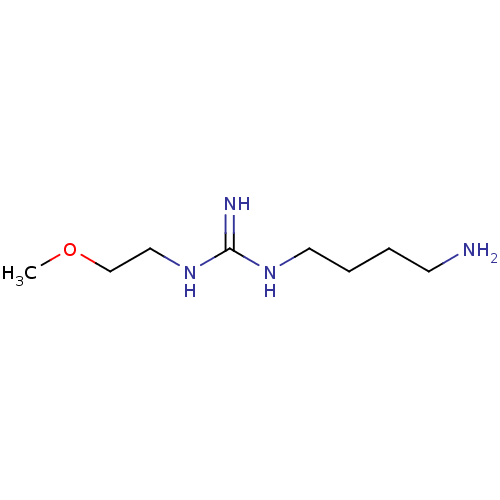
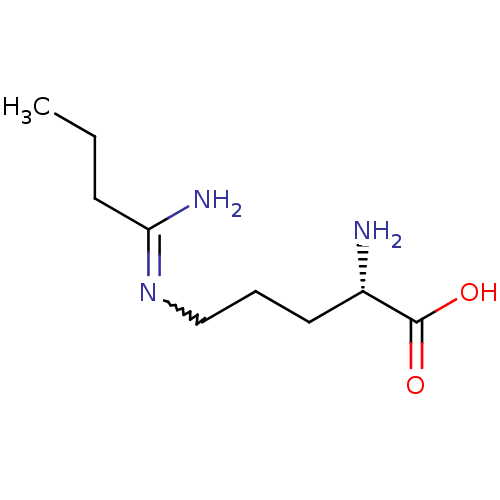
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
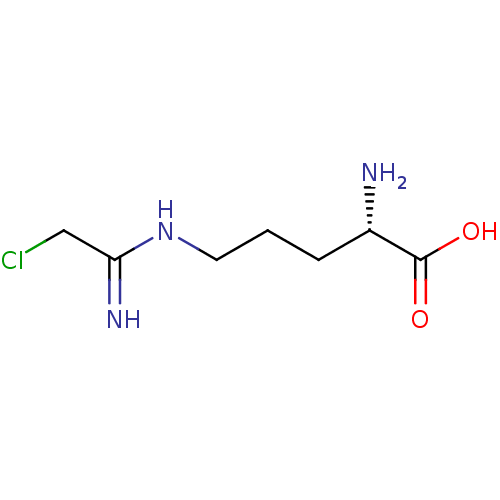
Affinity DataKi: 190nMAssay Description:Inhibition of N-terminal His6-tagged human DDAH1 expressed in Escherichia coli BL21 DE3 cells using L-Nva as substrate assessed as L-citrulline forma...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
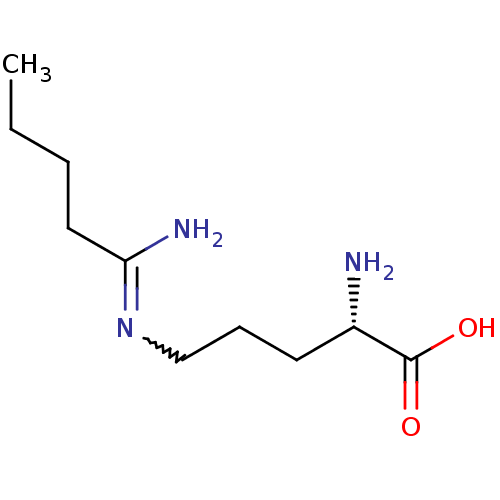
Affinity DataKi: 1.00E+3nMAssay Description:Inhibition of 6xHis-tagged recombinant human DDAH1 expressed in Escherichia coli BL21 cells using NMMA as substrate measured after 30 mins by colder ...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
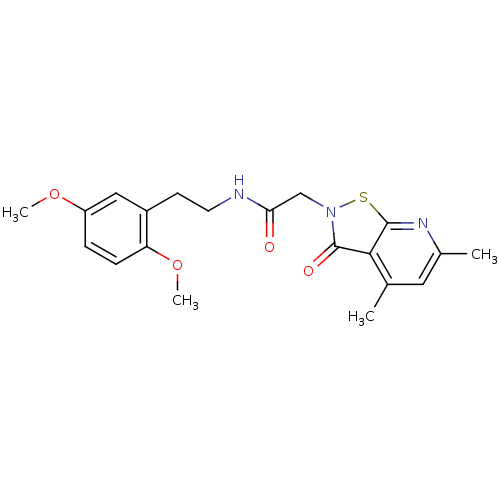
Affinity DataKi: 1.00E+3nMAssay Description:Inhibition of recombinant human DDAH1 expressed in HEK293 cells using ADMA as substrate assessed as inhibition constant preincubated for 5 minsMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
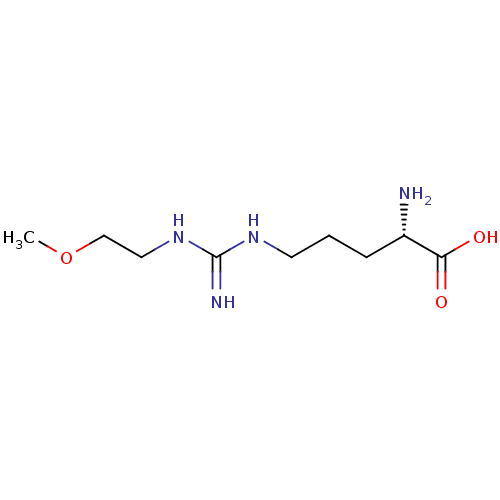
Affinity DataKi: 1.30E+3nMAssay Description:Inhibition of 6xHis-tagged recombinant human DDAH1 expressed in Escherichia coli BL21 cells using NMMA as substrate measured after 30 mins by colder ...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 1.30E+3nMAssay Description:Inhibition of N-terminal His6-tagged human DDAH1 transfected in HEK293T cells assessed as inhibition constant measured after 15 mins in presence of N...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 2.00E+3nMAssay Description:Inhibition of human DDAH1 expressed in Escherichia coli BL21 cells assessed as inhibition constant measured after 30 mins by calorimetric assayMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 2.00E+3nMAssay Description:Inhibition of human DDAH1 expressed using SMTC as substrate assessed as inhibition constant measured after 5 minsMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 2.00E+3nMAssay Description:Inhibition of 6xHis-tagged recombinant human DDAH1 expressed in Escherichia coli BL21 cells using NMMA as substrate measured after 30 mins by colder ...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 2.00E+3nM ΔG°: -33.8kJ/molepH: 7.4 T: 2°CAssay Description:A colorimetric assay was carried out in 150 μL 50 mM potassium phosphate buffer pH 7.4 containing 4 μg of recombinant hDDAH-1 and varying c...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 2.00E+3nMAssay Description:Inhibition of human DDAH1 by plate-reader assayMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of recombinant human DDAH1 expressed in HEK293 cells using ADMA as substrate preincubated for 5 minsMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 3.30E+3nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 5.10E+3nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 7.00E+3nMAssay Description:Inhibition of recombinant human DDAH1 expressed in HEK293 cells using ADMA as substrate assessed as inhibition constant preincubated for 5 minsMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 7.10E+3nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 7.50E+3nMAssay Description:Inhibition assay using DDAH-1 and nNOS.More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 7.50E+3nMAssay Description:Inhibition of N-terminal His6-tagged human DDAH1 expressed in Escherichia coli Rosetta 2 DE3 cells assessed as inhibition constant by calorimetric as...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 8.60E+3nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 9.00E+3nMAssay Description:Inhibition of 6xHis-tagged recombinant human DDAH1 expressed in Escherichia coli BL21 cells using NMMA as substrate measured after 30 mins by colder ...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 9.00E+3nMAssay Description:Inhibition of human DDAH1 using SMTC as substrate by fluorimetric assayMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 9.00E+3nMAssay Description:Inhibition of His-tagged human DDAH1 using SMTC as substrate assessed as inhibition constantMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 9.90E+3nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 9.90E+3nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 1.05E+4nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 1.10E+4nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 1.18E+4nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of human DDAH1 expressed in Escherichia coli BL21 cells measured after 30 mins by calorimetric assayMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 1.28E+4nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 1.30E+4nMAssay Description:Inhibition of human DDAH1 by plate-reader assayMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 1.30E+4nM ΔG°: -29.0kJ/molepH: 7.4 T: 2°CAssay Description:A colorimetric assay was carried out in 150 μL 50 mM potassium phosphate buffer pH 7.4 containing 4 μg of recombinant hDDAH-1 and varying c...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 1.30E+4nMAssay Description:Inhibition of human DDAH1 by plate-reader assayMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 1.30E+4nMAssay Description:Inhibition of 6xHis-tagged recombinant human DDAH1 expressed in Escherichia coli BL21 cells using NMMA as substrate measured after 30 mins by colder ...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 1.30E+4nM IC50: 2.90E+4nMpH: 7.4Assay Description:This assay was used to determine the inhibiting effect of the inhibitors in a concentration of 1 mM. A standard incubation mixture consisted of 75 &#...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 1.31E+4nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 1.32E+4nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 1.34E+4nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 1.37E+4nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 1.40E+4nMAssay Description:Inhibition of recombinant human DDAH1 expressed in HEK293 cells using ADMA as substrate assessed as inhibition constant preincubated for 5 minsMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 1.41E+4nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 1.70E+4nMAssay Description:Inhibition of human DDAH1 expressed in Escherichia coli BL21 cells assessed as inhibition constant measured after 30 mins by calorimetric assayMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 1.70E+4nM ΔG°: -28.3kJ/molepH: 7.4 T: 2°CAssay Description:A colorimetric assay was carried out in 150 μL 50 mM potassium phosphate buffer pH 7.4 containing 4 μg of recombinant hDDAH-1 and varying c...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 1.70E+4nMAssay Description:Inhibition of human DDAH1 by plate-reader assayMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 1.77E+4nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibition of recombinant human DDAH1 expressed in HEK293 cells using ADMA as substrate preincubated for 5 minsMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 1.80E+4nMAssay Description:Inhibition of 6xHis-tagged recombinant human DDAH1 expressed in Escherichia coli BL21 cells using NMMA as substrate measured after 30 mins by colder ...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 1.80E+4nM IC50: 7.00E+4nMpH: 7.4Assay Description:This assay was used to determine the inhibiting effect of the inhibitors in a concentration of 1 mM. A standard incubation mixture consisted of 75 &#...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 2.12E+4nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 2.33E+4nMAssay Description:The L-citrulline assay was based upon an original test-tube method developed by Prescott and Jones in 1969 (Prescott, L. M. & Jones, M. E. Modified m...More data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataIC50: 2.90E+4nMAssay Description:Inhibition of human DDAH1 by plate-reader assayMore data for this Ligand-Target Pair
TargetN(G),N(G)-dimethylarginine dimethylaminohydrolase 1(Human)
Flinders University
Curated by ChEMBL
Flinders University
Curated by ChEMBL
Affinity DataKi: 3.20E+4nMAssay Description:Inhibition of human DDAH1 by plate-reader assayMore data for this Ligand-Target Pair

 3D Structure (crystal)
3D Structure (crystal)