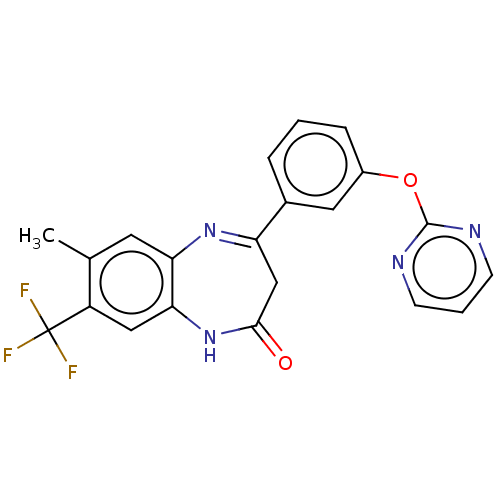
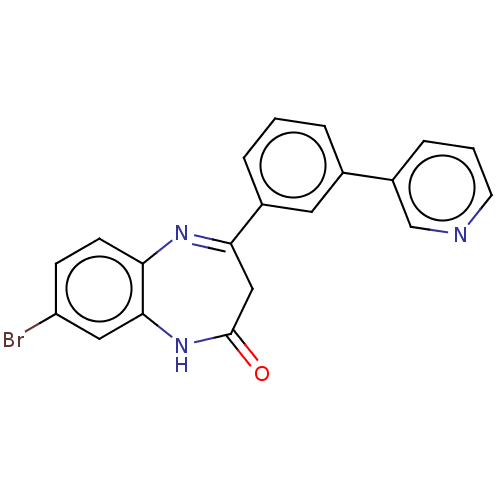
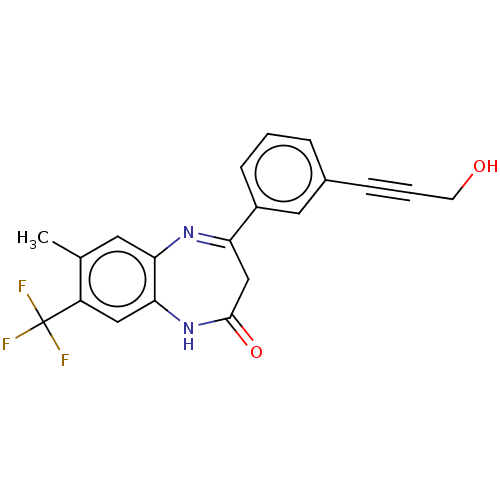
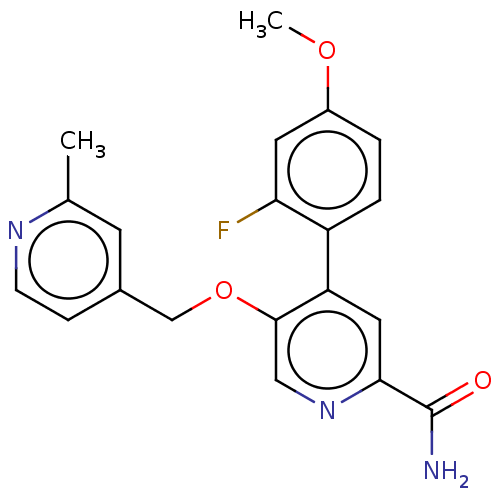
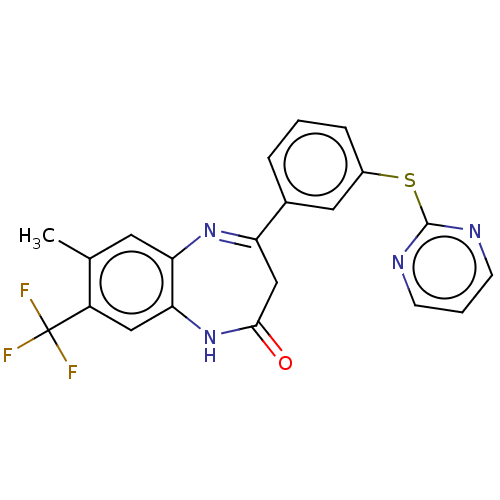
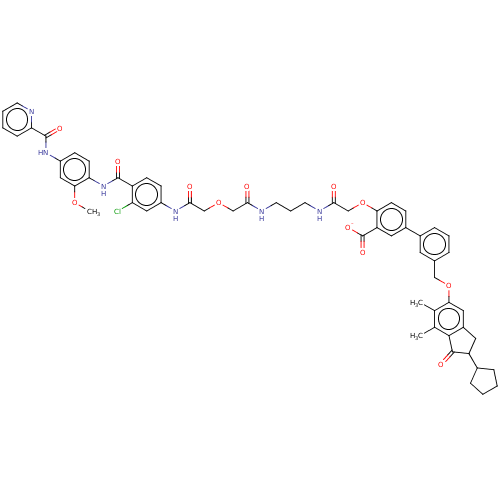
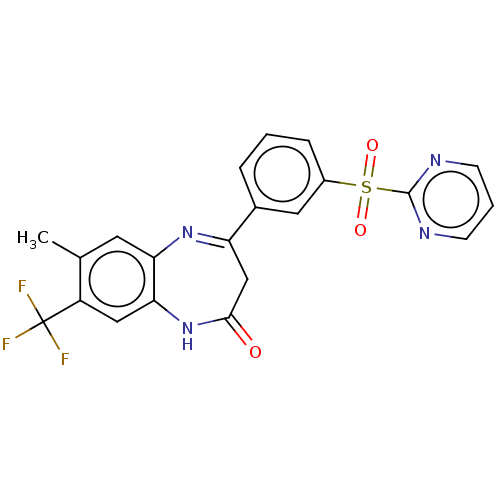
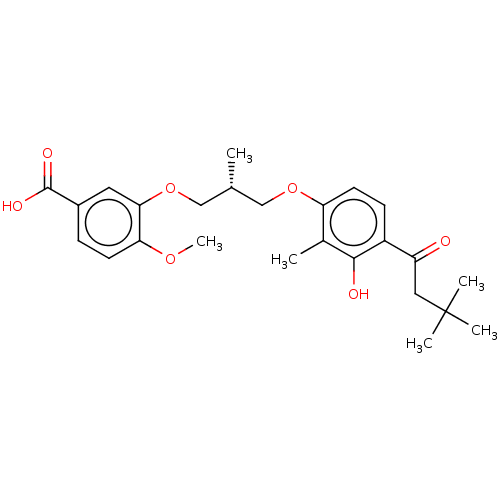
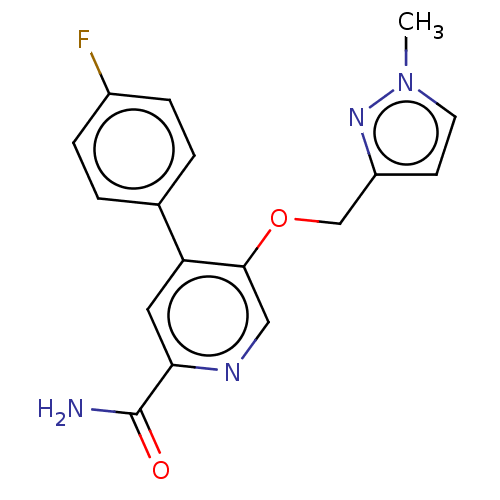
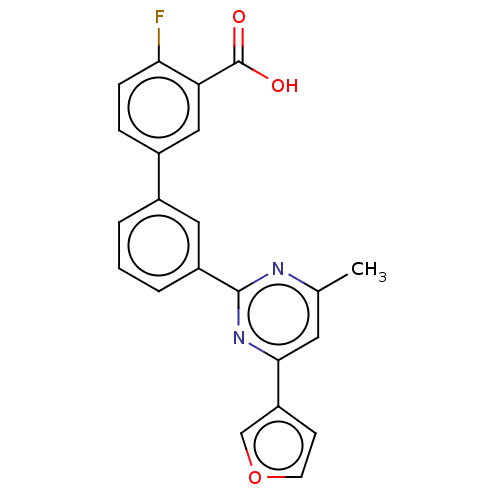
Affinity DataIC50: 8.90nMAssay Description:Negative allosteric modulation of rat mGluR2 by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 9.80nMAssay Description:Agonist activity at mGlu2/3 receptor in Sprague-Dawley rat forebrain assessed as inhibition of spontaneous Ca2+ oscillations in cortical neurons afte...More data for this Ligand-Target Pair
Affinity DataEC50: 10nMAssay Description:Positive allosteric modulation of recombinant rat Gi-coupled mGluR2 expressed in HEK293 Flp-in-T-REx cells with Galpha16 assessed as increase in glut...More data for this Ligand-Target Pair
Affinity DataEC50: 13nMAssay Description:Positive allosteric modulator activity at rat mGlu2 receptor transfected in GIRK channel overexpressing cells assessed as potentiation of glutamate-i...More data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Positive allosteric modulation of recombinant rat Gi-coupled mGluR2 expressed in HEK293 Flp-in-T-REx cells with Galpha16 assessed as increase in glut...More data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 28nMAssay Description:Positive allosteric modulator activity at rat mGlu2 receptor transfected in GIRK channel overexpressing cells assessed as potentiation of glutamate-i...More data for this Ligand-Target Pair
Affinity DataEC50: 28nMAssay Description:Positive allosteric modulator activity at rat mGlu2 receptor transfected in GIRK channel overexpressing cells assessed as potentiation of glutamate-i...More data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:G ±15 HEK293 cells stably expressing rat mGlu2 were plated in black-walled, clear-bottomed, poly-D-lysine coated 384-well plates in 20 L of assay med...More data for this Ligand-Target Pair
Affinity DataEC50: 30nMAssay Description:Positive allosteric modulation of recombinant rat Gi-coupled mGluR2 expressed in HEK293 Flp-in-T-REx cells with Galpha16 assessed as increase in glut...More data for this Ligand-Target Pair
Affinity DataEC50: 30nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 30nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 35nMAssay Description:Tested for the agonistic activity against Metabotropic glutamate receptor 2More data for this Ligand-Target Pair
Affinity DataIC50: 39nMAssay Description:G ±15 HEK293 cells stably expressing rat mGlu2 were plated in black-walled, clear-bottomed, poly-D-lysine coated 384-well plates in 20 L of assay med...More data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 50nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 50nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 50nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 50nMAssay Description:Positive allosteric modulator activity at rat mGlu2 receptor transfected in GIRK channel overexpressing cells assessed as potentiation of glutamate-i...More data for this Ligand-Target Pair
Affinity DataEC50: 50nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 50nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 50nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataIC50: 54nMAssay Description:Negative allosteric modulation of rat mGluR2 by GTPgammaS binding assayMore data for this Ligand-Target Pair
Affinity DataEC50: 54nMAssay Description:Positive allosteric modulation of rat mGlu2 receptor expressed in HEK293 cells coexpressing GIRK assessed as thallium influx in presence of EC20 glut...More data for this Ligand-Target Pair
Affinity DataEC50: 60nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 60nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 60nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 60nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 60nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 60nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 62nMAssay Description:Positive allosteric modulation of rat mGlu2 receptor expressed in HEK293 cells coexpressing GIRK assessed as thallium influx in presence of EC20 glut...More data for this Ligand-Target Pair
Affinity DataEC50: 62nMAssay Description:Positive allosteric modulation of rat mGlu2 receptor expressed in HEK293 cells coexpressing GIRK assessed as thallium influx in presence of EC20 glut...More data for this Ligand-Target Pair
Affinity DataEC50: 63nMAssay Description:Positive allosteric modulator activity at rat mGlu2 receptor transfected in GIRK channel overexpressing cells assessed as potentiation of glutamate-i...More data for this Ligand-Target Pair
Affinity DataIC50: 66nMAssay Description:G ±15 HEK293 cells stably expressing rat mGlu2 were plated in black-walled, clear-bottomed, poly-D-lysine coated 384-well plates in 20 L of assay med...More data for this Ligand-Target Pair
Affinity DataEC50: 69nMAssay Description:Positive allosteric modulation of rat mGlu2 receptor expressed in HEK293 cells coexpressing GIRK assessed as thallium influx in presence of EC20 glut...More data for this Ligand-Target Pair
Affinity DataEC50: 70nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 70nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
Affinity DataEC50: 70nMAssay Description:Human Embryonic Kidney (HEK-293) cell lines co-expressing rat mGlu receptors 2, 3, 4, 6, 7 or 8 and G protein-coupled inwardly-rectifying potassium (...More data for this Ligand-Target Pair
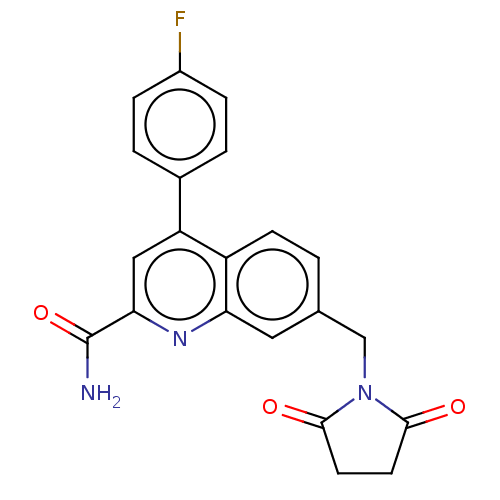
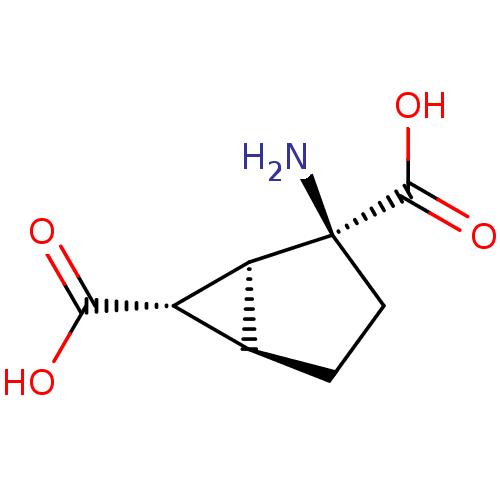
 3D Structure (crystal)
3D Structure (crystal)