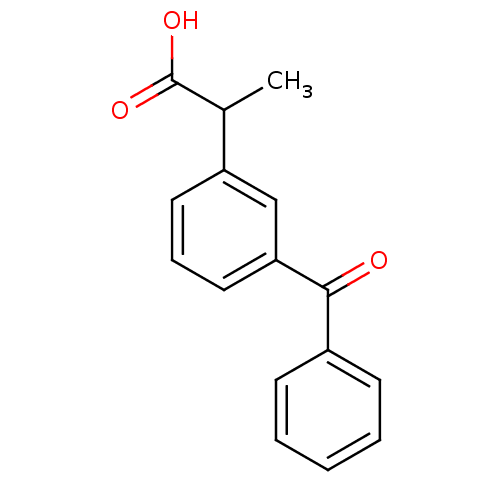
BDBM50022271 2-(3-Benzoylphenyl)propionic acid::2-(3-benzoylphenyl)propanoic acid::3-Benzoyl-alpha-methylbenzeneacetic acid::3-Benzoylhydratropic acid::CHEMBL571::Dexketoprofen trometamol::KETOPROFEN::L'Acide (benzoyl-3-phenyl)-2-propionique::Orudis (TN)::m-Benzoylhydratropic acid
SMILES CC(C(O)=O)c1cccc(c1)C(=O)c1ccccc1
InChI Key InChIKey=DKYWVDODHFEZIM-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 49 hits for monomerid = 50022271
Found 49 hits for monomerid = 50022271
Affinity DataKi: 30nMAssay Description:University of New Mexico Assay Overview: Assay Support: NIH I RO3 MH081231-01 HTS to identify specific small molecule inhibitors of Ras and Ras-rela...More data for this Ligand-Target Pair
TargetProstaglandin G/H synthase 1(Homo sapiens (Human))
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
TargetProstaglandin G/H synthase 1(Homo sapiens (Human))
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
TargetProstaglandin G/H synthase 2(Homo sapiens (Human))
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
TargetProstaglandin G/H synthase 2(Homo sapiens (Human))
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
TargetProstaglandin G/H synthase 2(Homo sapiens (Human))
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
Affinity DataKi: 8.24E+4nM ΔG°: -10.6kcal/moleT: 2°CAssay Description:Briefly, steady-state fluorescence spectra of ANS binding was monitored by measuring the increase in fluorescence signal between 450?550 nm following...More data for this Ligand-Target Pair
Affinity DataKi: 2.08E+5nMAssay Description:Binding affinity towards human serum albuminMore data for this Ligand-Target Pair
Affinity DataKi: 4.14E+5nMAssay Description:Inhibition of glyoxalase 1More data for this Ligand-Target Pair
Affinity DataKi: 8.43E+5nMAssay Description:Inhibition of human recombinant His-tagged glyoxalase 1 expressed in Escherichia coli BL21 (DE3) preincubated for 20 mins by Dixon plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 8.43E+5nMAssay Description:Inhibition of human glyoxalase 1More data for this Ligand-Target Pair
Affinity DataKi: 1.16E+6nMAssay Description:TP_TRANSPORTER: inhibition of MTX uptake in OAT3-expressing S2 cellsMore data for this Ligand-Target Pair
TargetSolute carrier organic anion transporter family member 1A3(Rattus norvegicus)
Kyoto University Hospital
Curated by ChEMBL
Kyoto University Hospital
Curated by ChEMBL
Affinity DataKi: 1.90E+6nMAssay Description:TP_TRANSPORTER: inhibition of MTX uptake in OAT-K1-expressing LLC-PK1 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 1.30E+7nM ΔG°: -2.57kcal/mole IC50: 6.23E+6nMpH: 10.5 T: 2°CAssay Description:The enzyme assay was based on the estimation of p-nitrophenol at 412nm. The molar extinction coefficient of p-nitrophenol was used to calculate enzy...More data for this Ligand-Target Pair
Affinity DataIC50: 9.50E+5nMpH: 10.0Assay Description:CA activity was measured by the Maren method that is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.45E+6nMpH: 10.0Assay Description:CA activity was measured by the Maren method that is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.85E+6nMpH: 10.0Assay Description:CA activity was measured by the Maren method that is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 h...More data for this Ligand-Target Pair
TargetProstaglandin G/H synthase 1(Homo sapiens (Human))
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
Affinity DataIC50: 2nMAssay Description:In vitro inhibition of cyclooxygenase-1 via inhibition of TXB2 generation in the presence of 1 uM arachidonic acid in human plateletMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+5nMAssay Description:In vitro inhibition of rabbit lens aldose reductase.More data for this Ligand-Target Pair
TargetProstaglandin G/H synthase 1(Homo sapiens (Human))
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
Affinity DataIC50: 330nMAssay Description:Inhibition of COX1 in human whole bloodMore data for this Ligand-Target Pair
TargetProstaglandin G/H synthase 2(Homo sapiens (Human))
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
Affinity DataIC50: 26nMAssay Description:In vitro inhibition of PGE-2 generation by LPS-stimulated monocytes isolated from human blood.More data for this Ligand-Target Pair
Affinity DataIC50: 800nMAssay Description:Inhibition of purified ovine COX1 pre-treated for 1 hr before 10-acetyl-3,7-dihydroxyphenoxazin substrate addition in absence of porcine liver estera...More data for this Ligand-Target Pair
TargetProstaglandin G/H synthase 2(Homo sapiens (Human))
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
Affinity DataIC50: 3.80E+3nMAssay Description:Inhibition of human recombinant COX2 pre-treated for 1 hr before 10-acetyl-3,7-dihydroxyphenoxazin substrate addition in absence of porcine liver est...More data for this Ligand-Target Pair
Affinity DataIC50: >5.00E+4nMAssay Description:Inhibition of human recombinant CYP2J2 assessed as reduction in astemizole O-demethylation by LC-MS/MS methodMore data for this Ligand-Target Pair
TargetProstaglandin G/H synthase 2(Homo sapiens (Human))
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
Affinity DataIC50: 690nMAssay Description:Inhibition of COX2 in human whole bloodMore data for this Ligand-Target Pair
TargetSolute carrier family 22 member 6(Rattus norvegicus)
Kyoto University Hospital
Curated by ChEMBL
Kyoto University Hospital
Curated by ChEMBL
Affinity DataIC50: 500nMAssay Description:TP_TRANSPORTER: inhibition of MTX uptake in Xenopus laevis oocytesMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:TP_TRANSPORTER: inhibition of Adefovir uptake in OAT1-expressing CHO cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:Inhibition of ovine COX2 assessed as reduction in PGH2 production by enzyme immunoassayMore data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:Inhibition of ovine COX1 assessed as reduction in PGH2 production by enzyme immunoassayMore data for this Ligand-Target Pair
TargetProstaglandin G/H synthase 2(Homo sapiens (Human))
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
St. Bartholomew'S And The Royal London School Of Medicine And Dentistry
Curated by PDSP Ki Database
Affinity DataIC50: 61nMAssay Description:Inhibition of recombinant human COX-2 preincubated for 15 mins by fluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 67nMAssay Description:Inhibition of Ovine COX-1 preincubated for 15 mins by fluorescence analysisMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Homo sapiens (Human))
Guangzhou Institutes Of Biomedicine And Health
Curated by ChEMBL
Guangzhou Institutes Of Biomedicine And Health
Curated by ChEMBL
Affinity DataIC50: >1.00E+4nMAssay Description:Inhibition of EGFR (unknown origin) using tyrosine 4 as substrate by fluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+5nMAssay Description:Inhibition of Glycine max (soybean) lipoxygenase using sodium linoleate as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+5nMAssay Description:Inhibition of Glycine max (soybean) lipoxygenase assessed as formation of 13-hydroperoxylinoleic acid using sodium linoleate as substrate after 7 minMore data for this Ligand-Target Pair
Affinity DataIC50: 6.83E+5nMpH: 10.0Assay Description:CA activity was measured by the Maren method that is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 h...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.4Assay Description:Experiments were performed in a 384-well format (Greiner no. 781207) as per the conditions noted here. Caspase-1: 2.5 nM enzyme, 6.5 mM WEHD substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.4Assay Description:Experiments were performed in a 384-well format (Greiner no. 781207) as per the conditions noted here. Caspase-1: 2.5 nM enzyme, 6.5 mM WEHD substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.4Assay Description:Experiments were performed in a 384-well format (Greiner no. 781207) as per the conditions noted here. Caspase-1: 2.5 nM enzyme, 6.5 mM WEHD substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.4Assay Description:Experiments were performed in a 384-well format (Greiner no. 781207) as per the conditions noted here. Caspase-1: 2.5 nM enzyme, 6.5 mM WEHD substrat...More data for this Ligand-Target Pair
Affinity DataIC50: >6.60E+4nMpH: 7.4Assay Description:Experiments were performed in a 384-well format (Greiner no. 781207) as per the conditions noted here. Caspase-1: 2.5 nM enzyme, 6.5 mM WEHD substrat...More data for this Ligand-Target Pair
Affinity DataEC50: 3.00E+4nMAssay Description:University of New Mexico Assay Overview: Assay Support: NIH I RO3 MH081231-01 HTS to identify specific small molecule inhibitors of Ras and Ras-rela...More data for this Ligand-Target Pair
Affinity DataEC50: 3.00E+4nMAssay Description:University of New Mexico Assay Overview: Assay Support: NIH I RO3 MH081231-01 HTS to identify specific small molecule inhibitors of Ras and Ras-rela...More data for this Ligand-Target Pair
Affinity DataEC50: 3.00E+4nMAssay Description:University of New Mexico Assay Overview: Assay Support: NIH I RO3 MH081231-01 HTS to identify specific small molecule inhibitors of Ras and Ras-rela...More data for this Ligand-Target Pair
Affinity DataEC50: 3.00E+4nMAssay Description:University of New Mexico Assay Overview: Assay Support: NIH I RO3 MH081231-01 HTS to identify specific small molecule inhibitors of Ras and Ras-rela...More data for this Ligand-Target Pair
TargetCell division control protein 42 homolog(Homo sapiens (Human))
Nmmlsc
Curated by PubChem BioAssay
Nmmlsc
Curated by PubChem BioAssay
Affinity DataEC50: 3.00E+4nMAssay Description:University of New Mexico Assay Overview: Assay Support: NIH I RO3 MH081231-01 HTS to identify specific small molecule inhibitors of Ras and Ras-rela...More data for this Ligand-Target Pair
TargetCell division control protein 42 homolog(Homo sapiens (Human))
Nmmlsc
Curated by PubChem BioAssay
Nmmlsc
Curated by PubChem BioAssay
Affinity DataEC50: 3.00E+4nMAssay Description:University of New Mexico Assay Overview: Assay Support: NIH I RO3 MH081231-01 HTS to identify specific small molecule inhibitors of Ras and Ras-rela...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:TP_TRANSPORTER: inhibition of 6-Carboxyfluorescein uptake in OAT1-expressing CHO cellsMore data for this Ligand-Target Pair
Affinity DataEC50: 377nMAssay Description:University of New Mexico Assay Overview: Assay Support: NIH I RO3 MH081231-01 HTS to identify specific small molecule inhibitors of Ras and Ras-rela...More data for this Ligand-Target Pair
Affinity DataEC50: 3.00E+4nMAssay Description:University of New Mexico Assay Overview: Assay Support: NIH I RO3 MH081231-01 HTS to identify specific small molecule inhibitors of Ras and Ras-rela...More data for this Ligand-Target Pair
 3D Structure (crystal)
3D Structure (crystal)