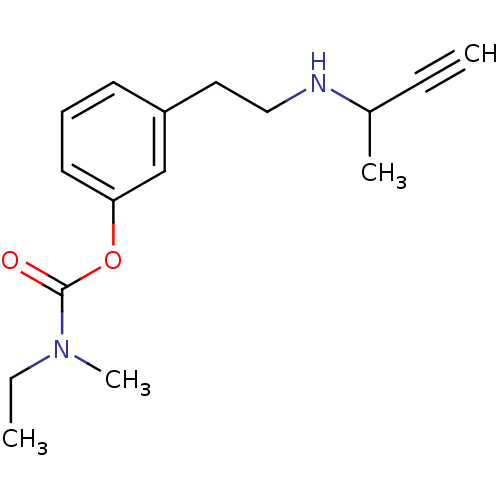
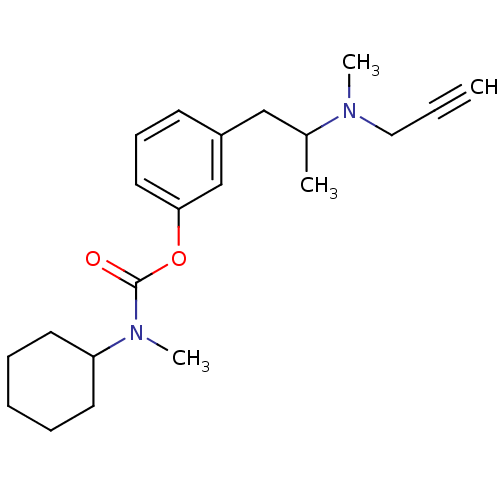
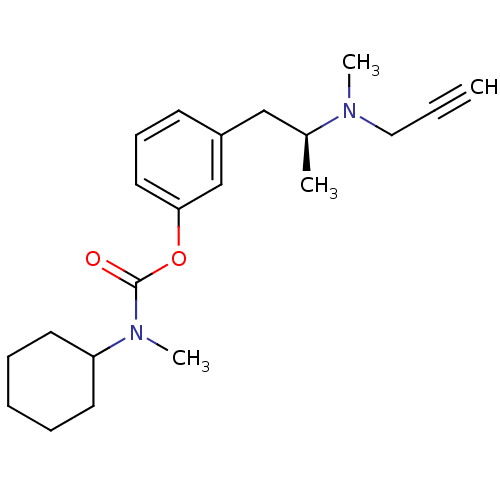
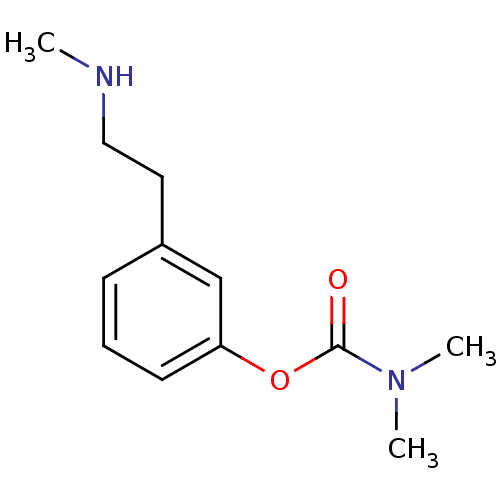
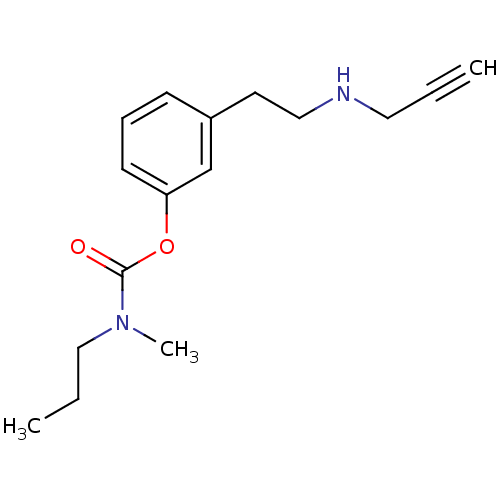
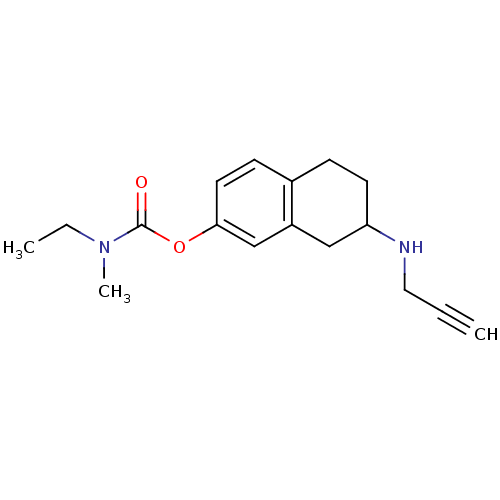
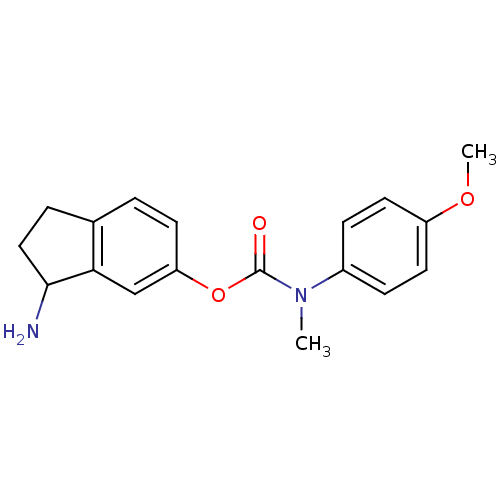
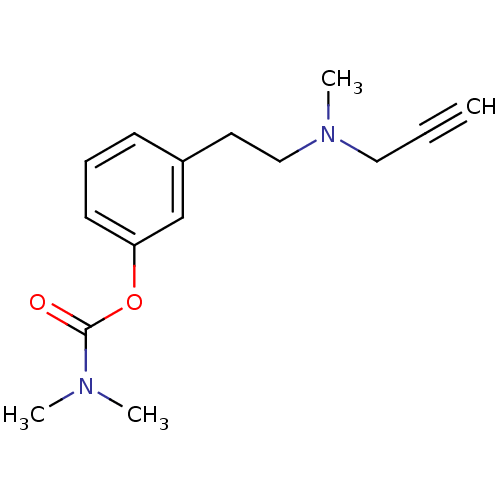
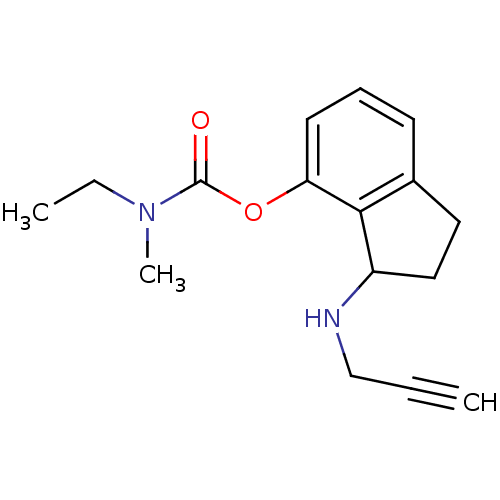
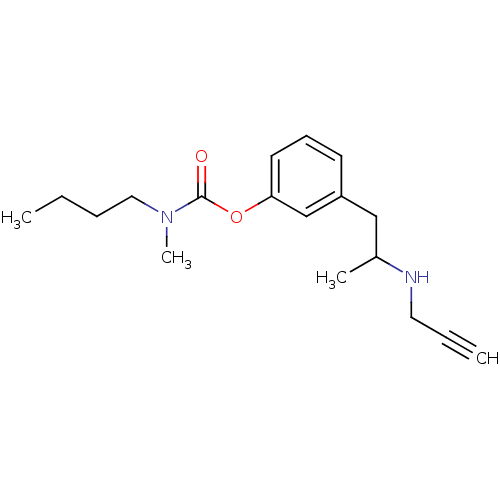
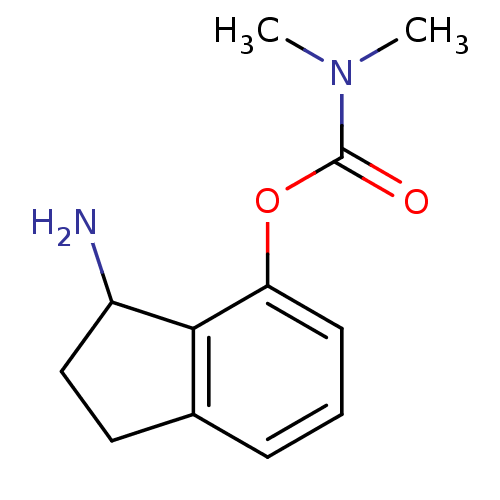
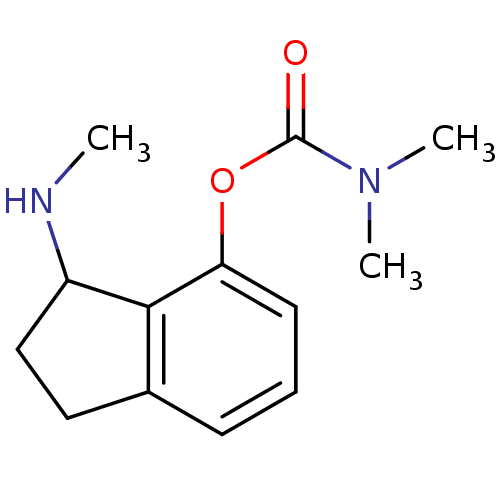
TargetMu-type opioid receptor(Rattus norvegicus (rat))
Hebrew University Of Jerusalem
Curated by ChEMBL
Hebrew University Of Jerusalem
Curated by ChEMBL
Affinity DataKi: 3.70nMAssay Description:Binding affinity of the compound to opioid receptor mu was determined in rat brain using [3H]-DAGO as radioligandMore data for this Ligand-Target Pair
TargetMu-type opioid receptor(Rattus norvegicus (rat))
Hebrew University Of Jerusalem
Curated by ChEMBL
Hebrew University Of Jerusalem
Curated by ChEMBL
Affinity DataKi: 3.90nMAssay Description:Binding affinity of the compound to opioid receptor mu was determined in rat brain using [3H]-DAGO as radioligandMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor(Rattus norvegicus (rat))
Hebrew University Of Jerusalem
Curated by ChEMBL
Hebrew University Of Jerusalem
Curated by ChEMBL
Affinity DataKi: 7.80nMAssay Description:Binding affinity of the compound to opioid receptor delta was determined in rat brain using [3H]-DSTBULET as radioligandMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor(Rattus norvegicus (rat))
Hebrew University Of Jerusalem
Curated by ChEMBL
Hebrew University Of Jerusalem
Curated by ChEMBL
Affinity DataKi: 629nMAssay Description:Binding affinity of the compound to opioid receptor delta was determined in rat brain using [3H]-DSTBULET as radioligandMore data for this Ligand-Target Pair
TargetMu-type opioid receptor(Rattus norvegicus (rat))
Hebrew University Of Jerusalem
Curated by ChEMBL
Hebrew University Of Jerusalem
Curated by ChEMBL
Affinity DataKi: 1.38E+3nMAssay Description:Binding affinity of the compound to opioid receptor mu was determined in rat brain using [3H]-DAGO as radioligandMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor(Rattus norvegicus (rat))
Hebrew University Of Jerusalem
Curated by ChEMBL
Hebrew University Of Jerusalem
Curated by ChEMBL
Affinity DataKi: 9.20E+3nMAssay Description:Binding affinity of the compound to opioid receptor delta was determined in rat brain using [3H]-DSTBULET as radioligandMore data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 27nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 30nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 53nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 54nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 70nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 70nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 240nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 240nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 240nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 260nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 280nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 280nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 290nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 290nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 310nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 400nMAssay Description:Inhibition of MAO activity was determined by a radiometric procedure from Tipton and Youdim. Homogenized rat brain was used as the source of enzymes....More data for this Ligand-Target Pair
Affinity DataIC50: 460nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair
Affinity DataIC50: 510nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The absorbance changes at 412 nm were recorded for 5 min with ...More data for this Ligand-Target Pair

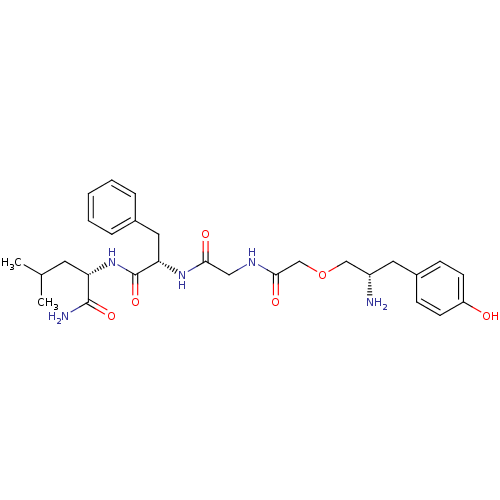
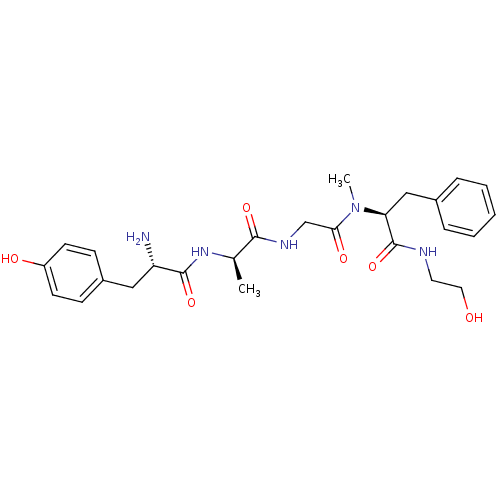
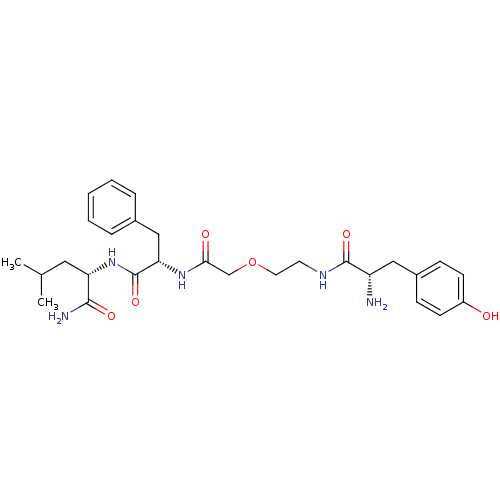
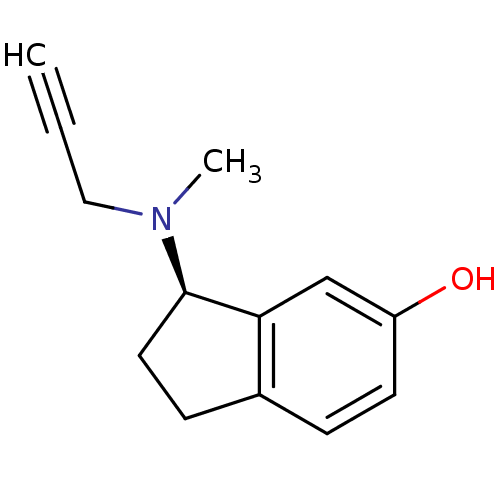
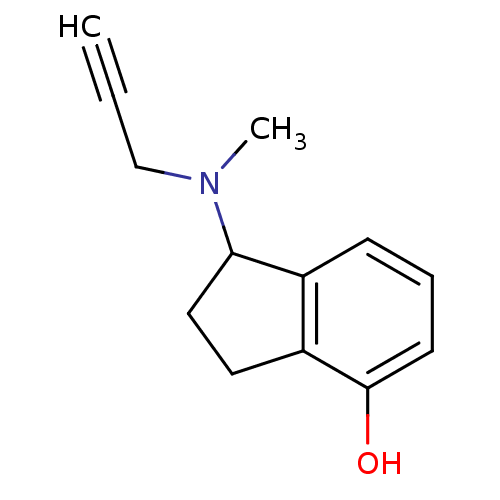
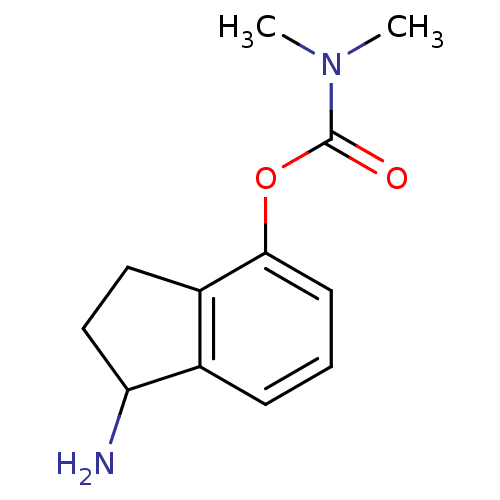
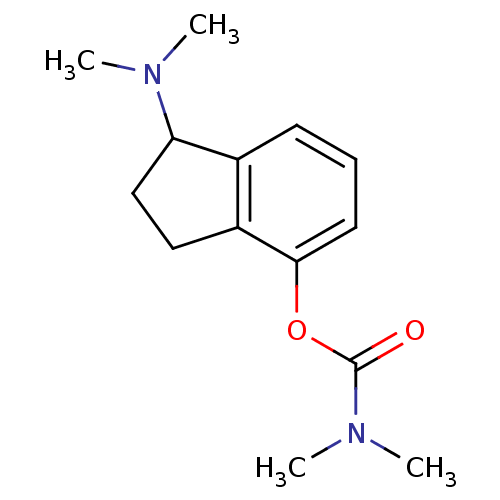

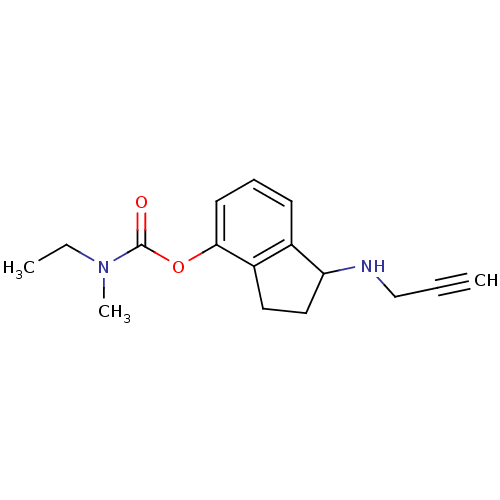
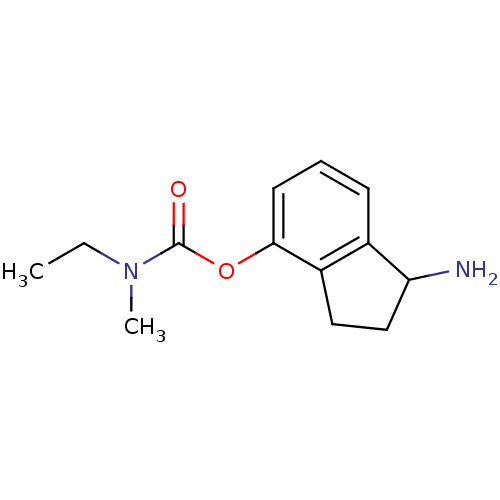
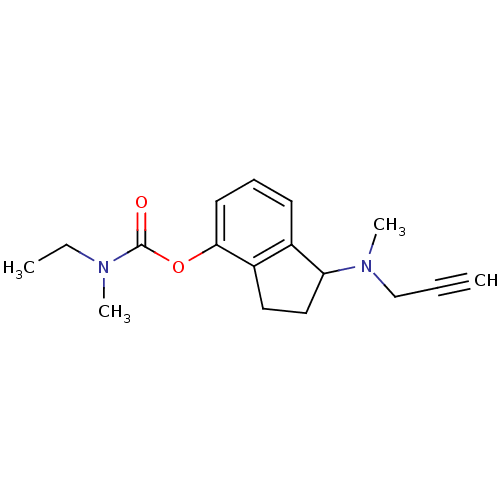


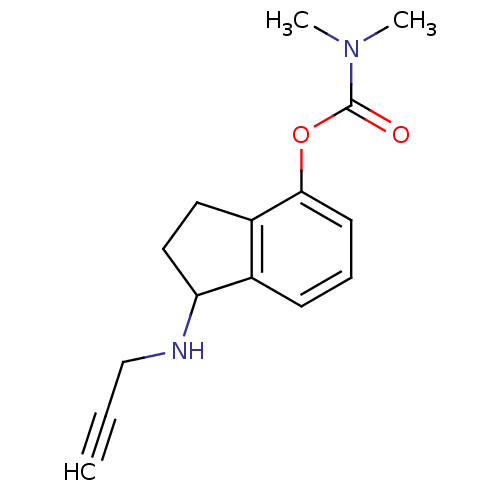
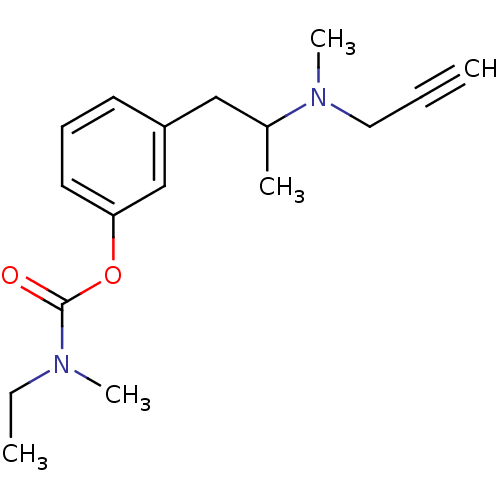
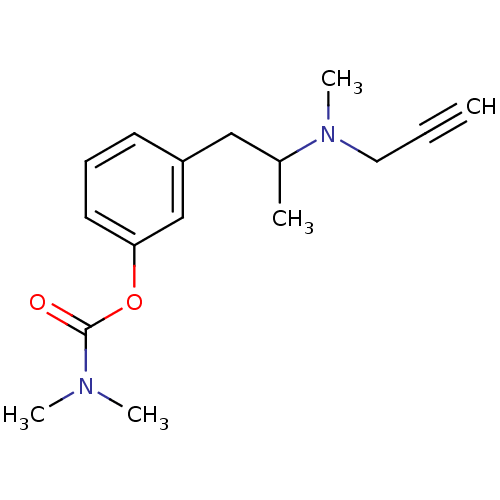
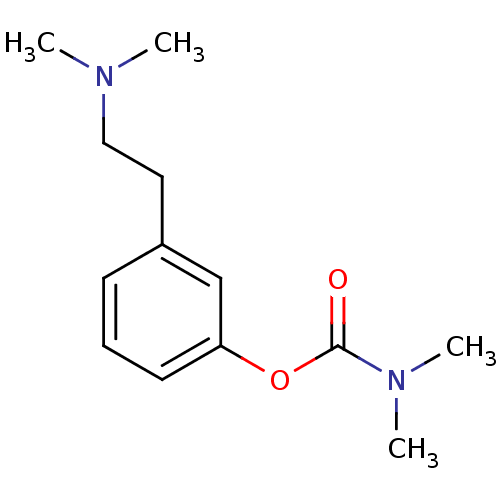
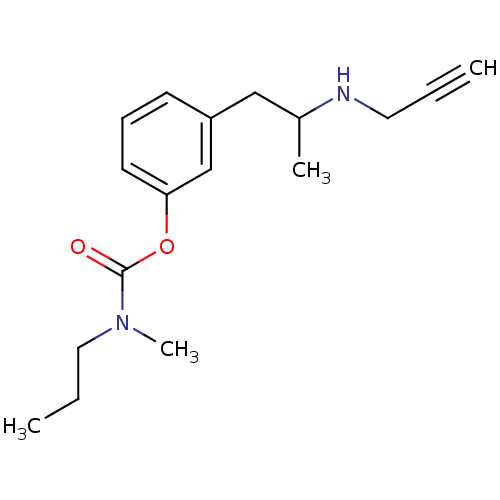
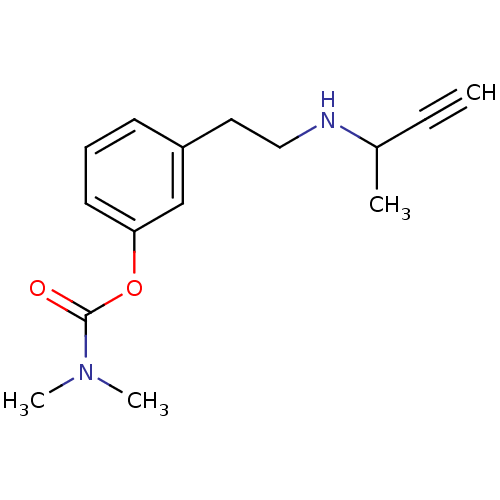
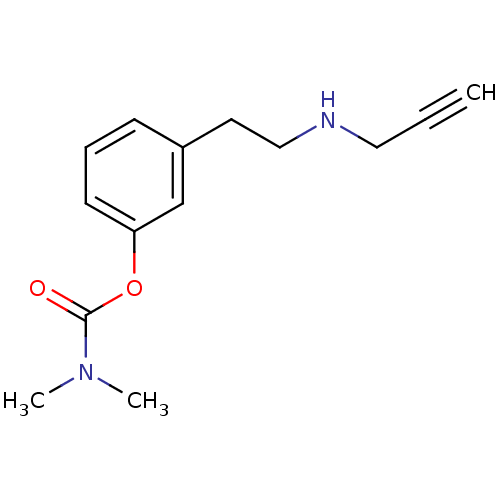
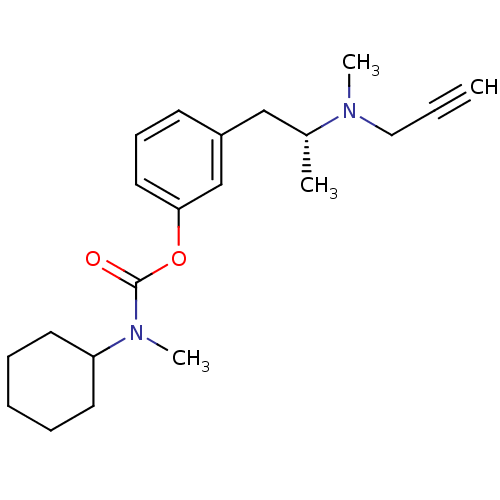
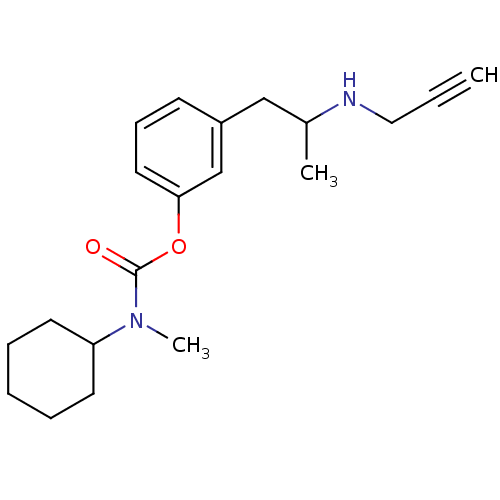
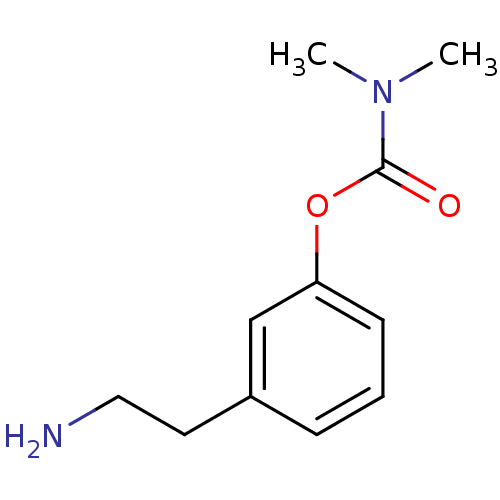
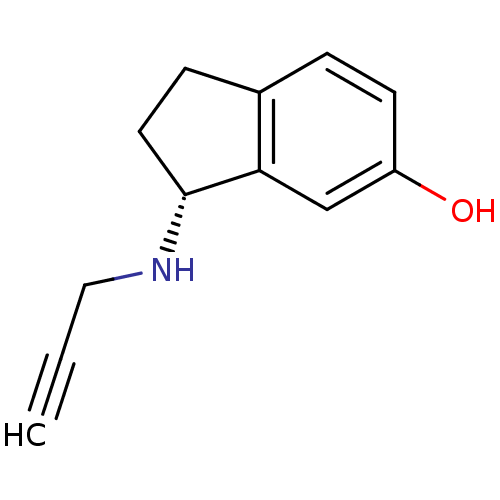
 3D Structure (crystal)
3D Structure (crystal)