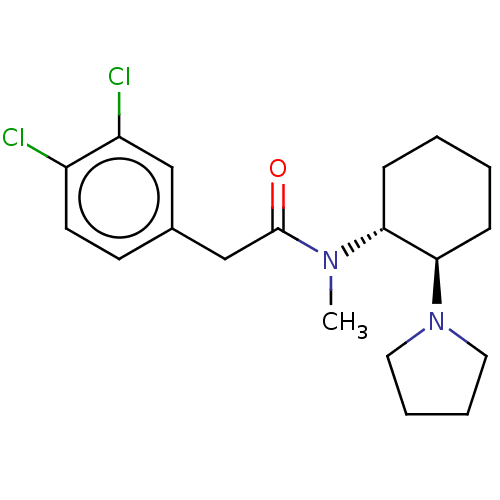
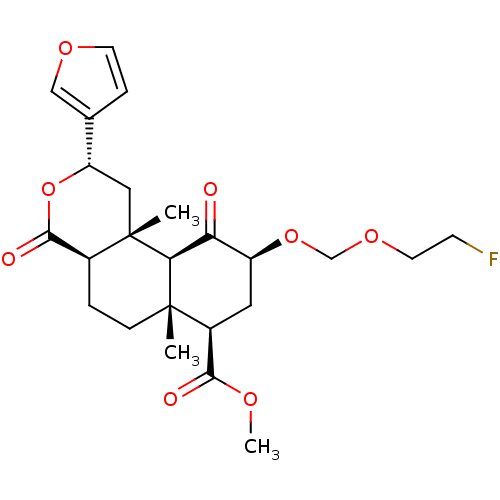
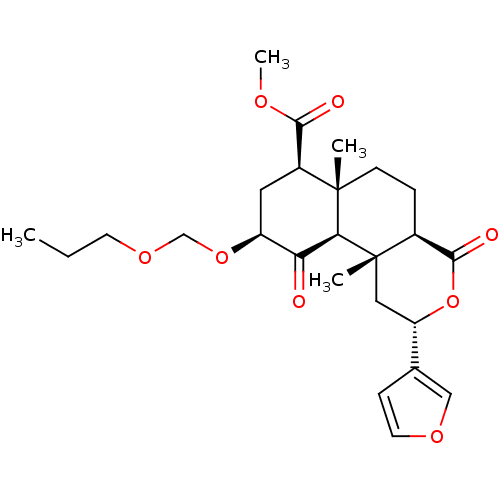
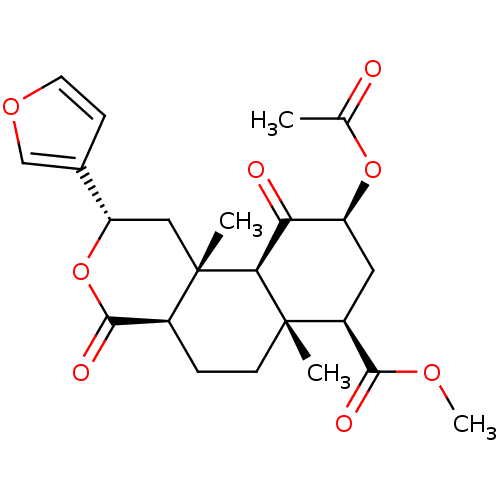
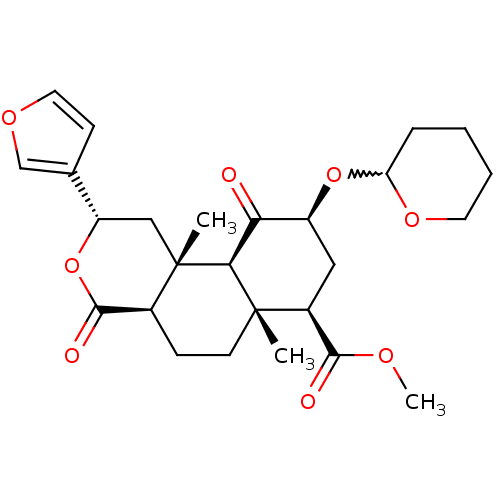
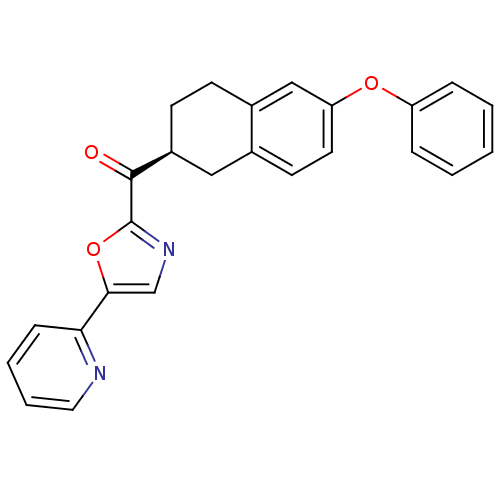
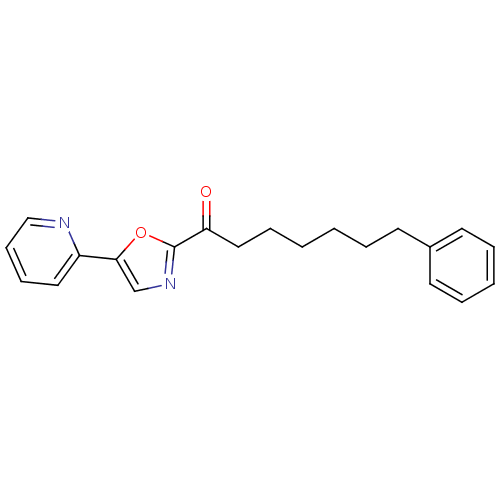
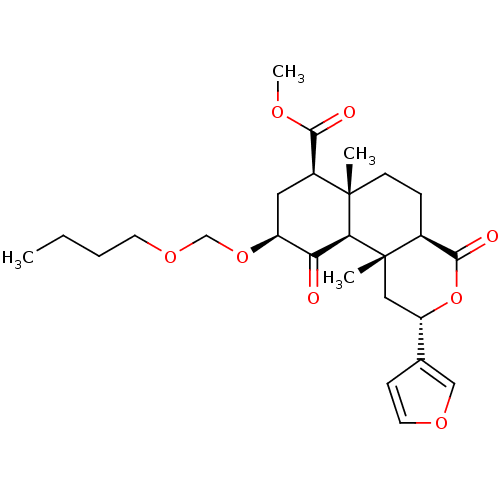
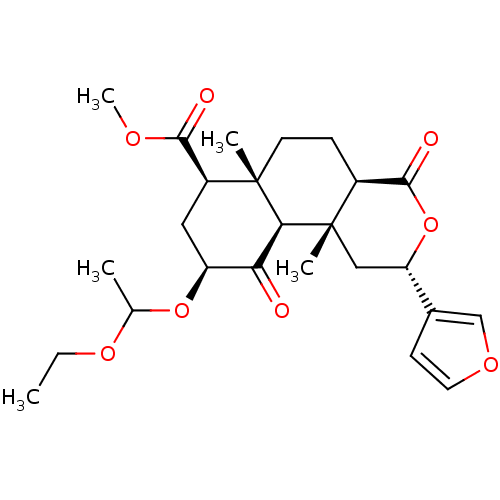
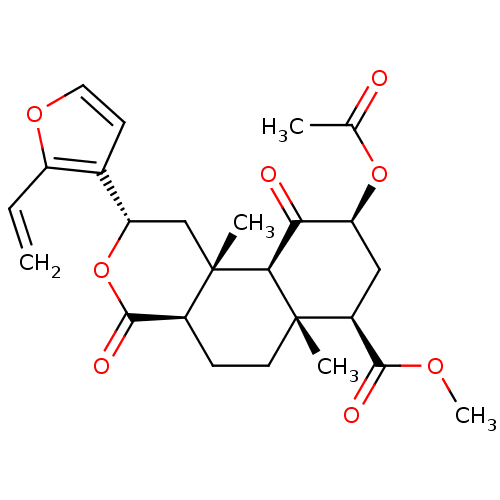
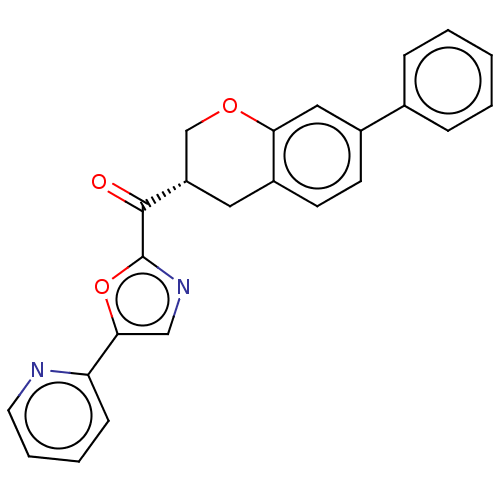
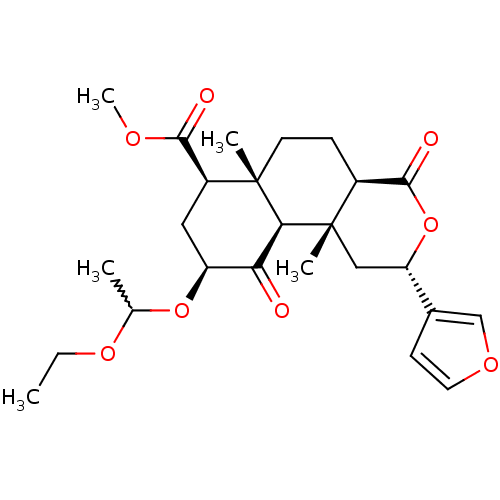
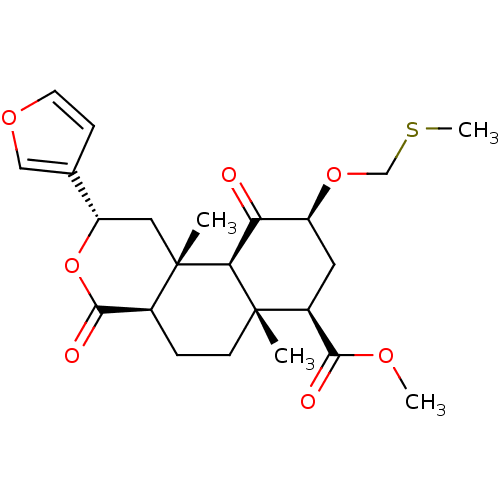
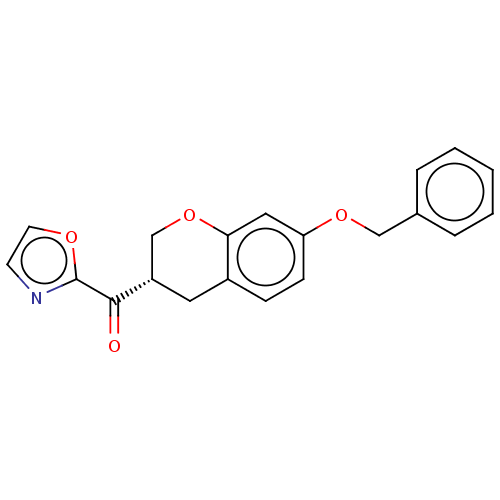
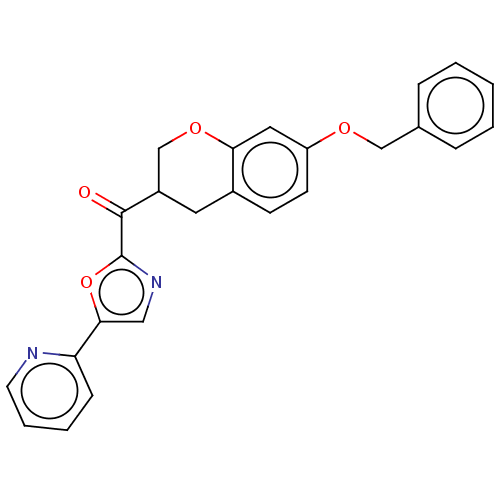
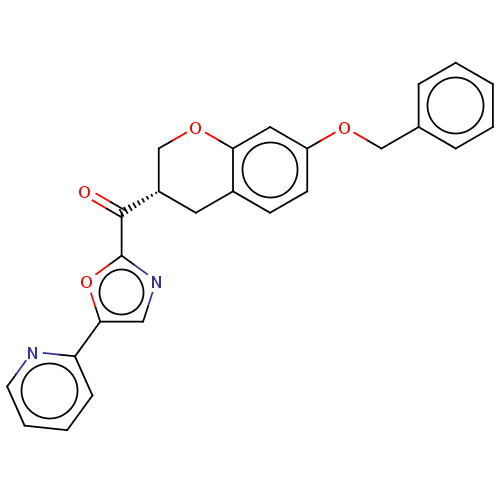
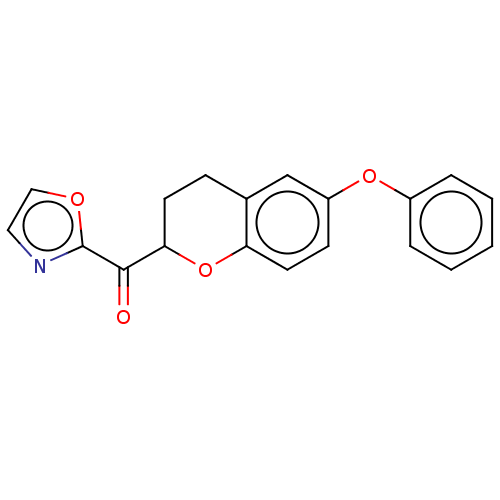
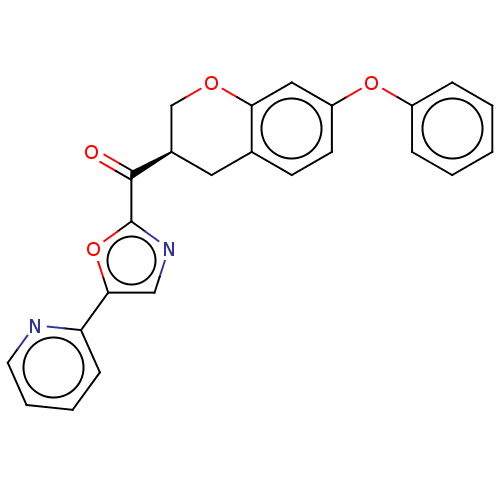
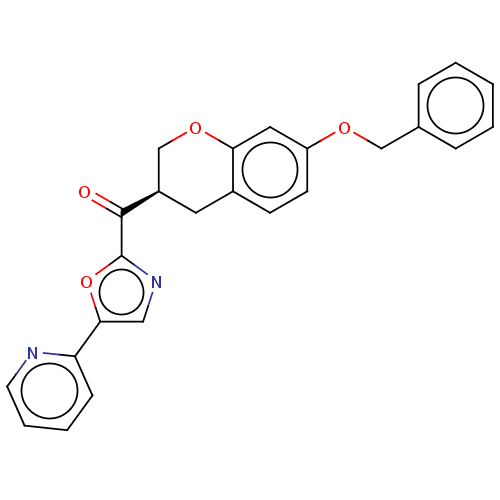
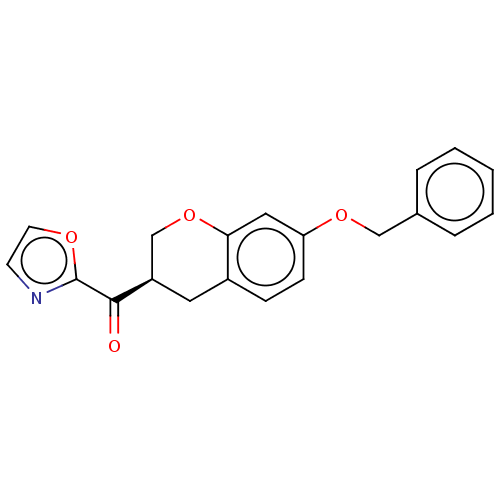
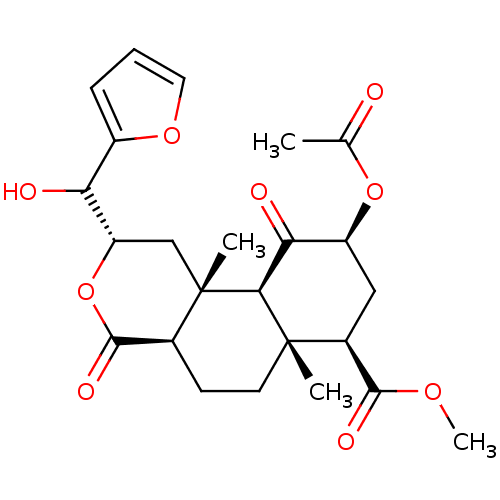
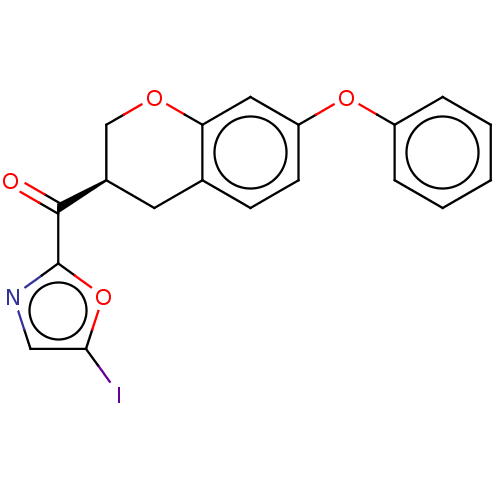
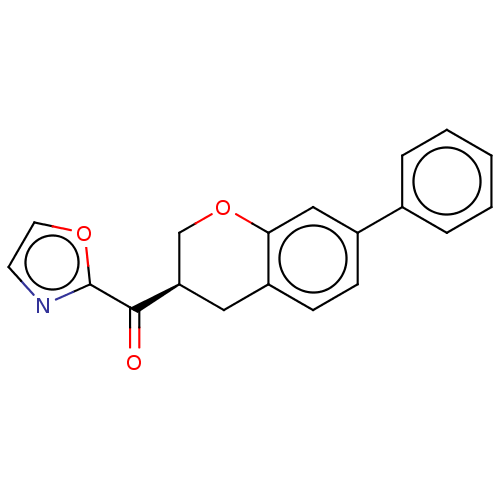
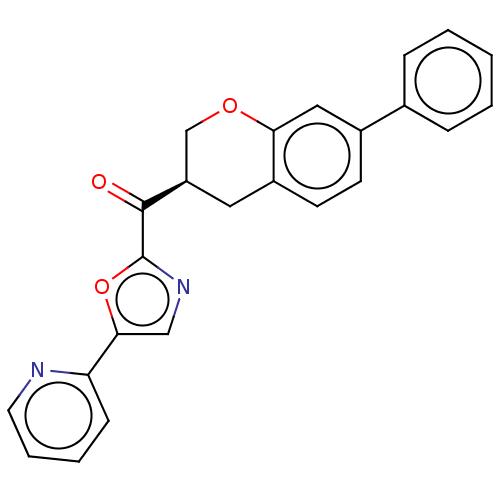
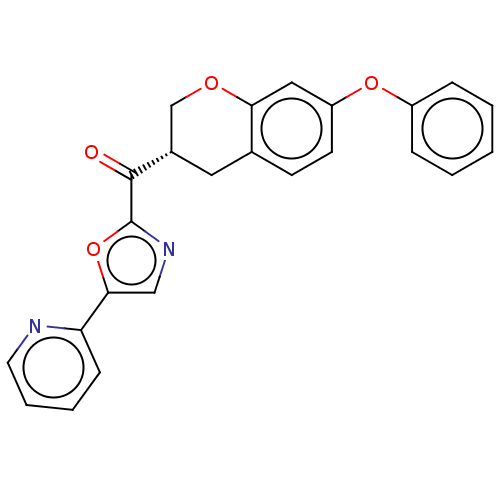
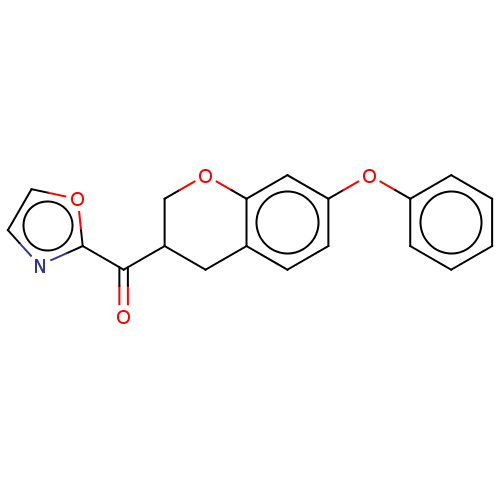
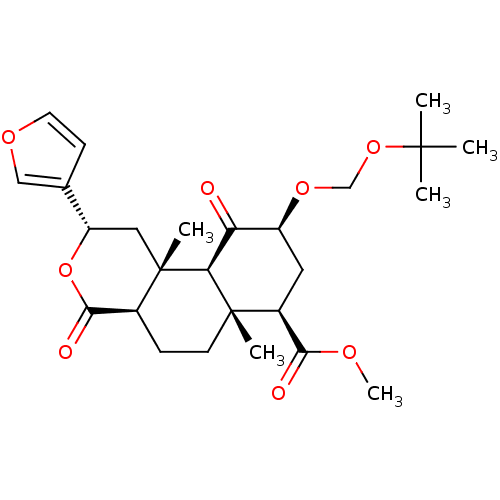
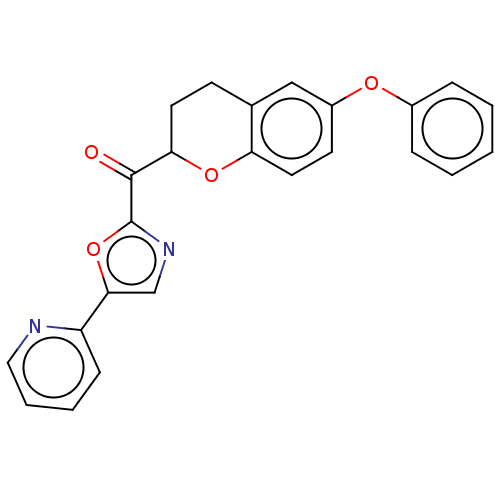
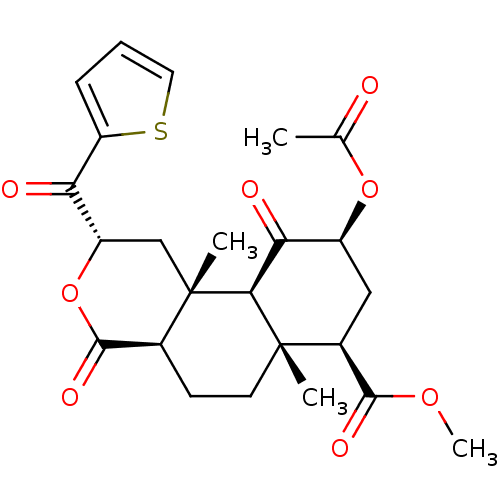
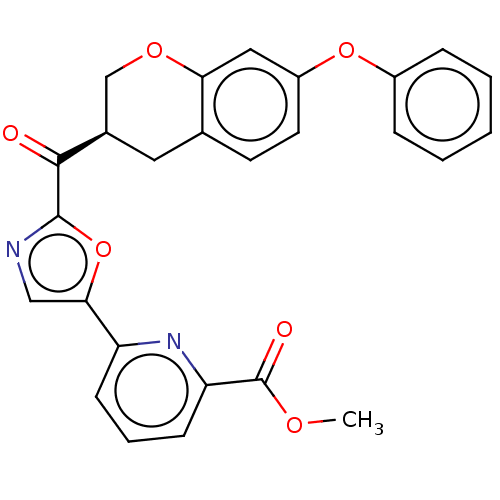
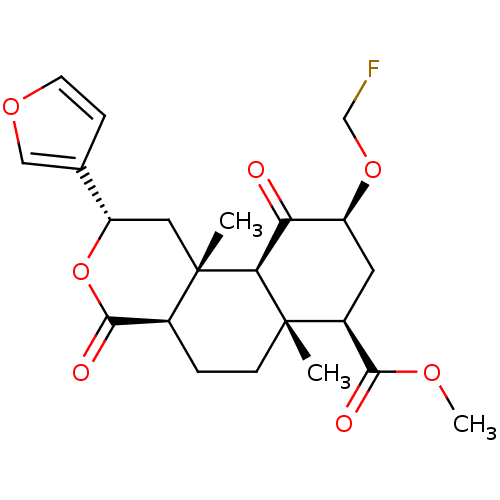
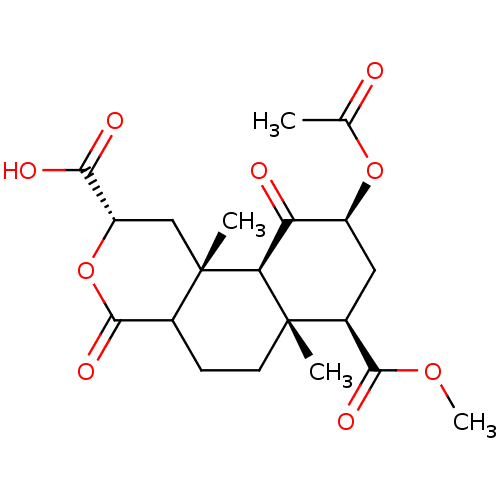
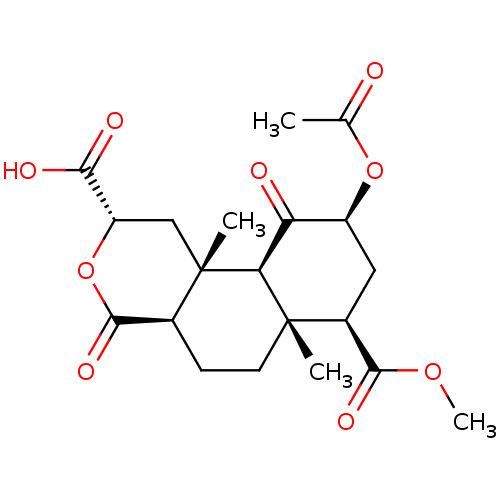
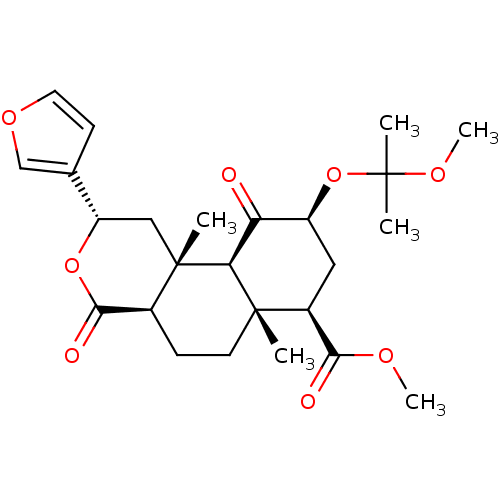
Affinity DataKi: 0.320nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
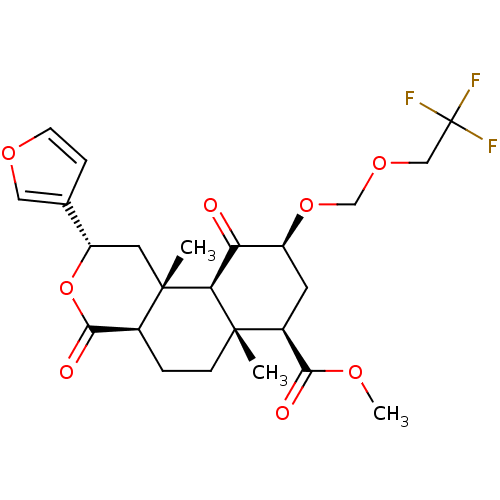
Affinity DataKi: 0.600nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
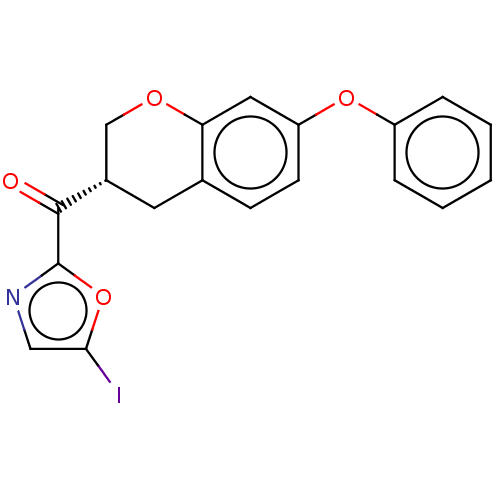
Affinity DataKi: 1.60nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
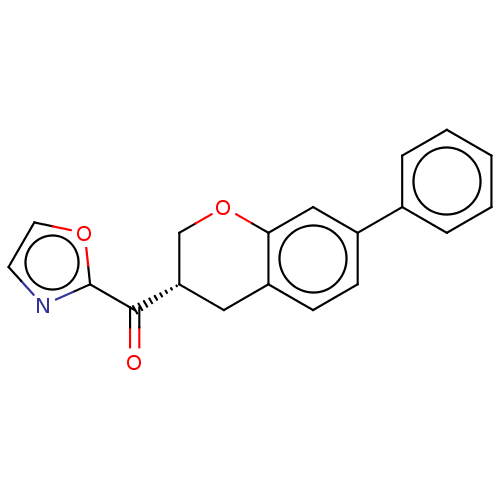
Affinity DataKi: 1.60nMAssay Description:Displacement of [3H]diprenorphine from human KOPR expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 1.90nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Displacement of [3H]diprenorphine from human KOPR expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.90nMAssay Description:Displacement of [3H]diprenorphine from human KOPR expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 4.40nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 4.70nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1(Mus musculus (mouse))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 4.70nMAssay Description:Inhibition of mouse FAAHMore data for this Ligand-Target Pair
Affinity DataKi: 5.30nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 6.60nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 7.10nMAssay Description:Displacement of [3H]diprenorphine from human KOPR expressed in CHO cellsMore data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 11nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 16nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 17nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 17nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 17nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 17nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 18nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 18nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 20nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:In vitro binding assay: The affinities of compounds for opioid receptors were determined by competitive inhibition of [3H]diprenorphine binding to ka...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Displacement of [3H]diprenorphine from human KOPR expressed in CHO cellsMore data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 20nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 20nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 23nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 27nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 31nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 32nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 33nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
Affinity DataKi: 35nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 36nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 36nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
Affinity DataKi: 38nMAssay Description:In vitro binding assay: The affinities of compounds for opioid receptors were determined by competitive inhibition of [3H]diprenorphine binding to ka...More data for this Ligand-Target Pair
Affinity DataKi: 38nMAssay Description:Displacement of [3H]diprenorphine from human KOPR expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Displacement of [3H]diprenorphine from human KOPR expressed in CHO cellsMore data for this Ligand-Target Pair
TargetFatty-acid amide hydrolase 1 [30-579](Rattus norvegicus (rat))
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 41nMAssay Description:Inhibition of rat recombinant FAAH expressed in Escherichia coli using [14C]oleamide as substrate assessed as oleic acid formation by Dixon plot anal...More data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:In vitro binding assay: The affinities of compounds for opioid receptors were determined by competitive inhibition of [3H]diprenorphine binding to ka...More data for this Ligand-Target Pair
Affinity DataKi: 50nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 55nMAssay Description:In vitro binding assay: The affinities of compounds for opioid receptors were determined by competitive inhibition of [3H]diprenorphine binding to ka...More data for this Ligand-Target Pair
Affinity DataKi: 55nMAssay Description:Displacement of [3H]diprenorphine from human KOPR expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 72nMAssay Description:Displacement of [3H]diprenorphine from human kappa opioid receptor expressed in CHO cellsMore data for this Ligand-Target Pair



 3D Structure (crystal)
3D Structure (crystal)