Affinity DataKi: 0.230nMAssay Description:Displacement of [125I]-SI-Ang-2 from AT1 receptor in Rattus norvegicus Sprague-Dawley (rat) liver membranes after 2 hrMore data for this Ligand-Target Pair
Affinity DataKi: 8nM ΔG°: -10.3kcal/molepH: 8.2 T: 2°CAssay Description:Binding assays were performed in 96-well plate format, using a classical filtration assay with a human full length PPARγ construct (GST-PPAR LBD...More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Binding affinity to GST-tagged PPARG LBD (unknown origin) by Fluormone Pan-PPAR Green tracer based TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 27nMAssay Description:Agonist activity at PPARgamma ligand binding domain (unknown origin) using fluormone Pan-PPAR green tracer by TR-FRET assay based competitive ligand ...More data for this Ligand-Target Pair
TargetCDGSH iron-sulfur domain-containing protein 1(Homo sapiens (Human))
West Virginia University
Curated by ChEMBL
West Virginia University
Curated by ChEMBL
Affinity DataKi: 31nMAssay Description:Displacement of [3H]rosiglitazone from recombinant human C-terminal His-tagged MitoNEET cytosolic domain (32 to 108 residues) expressed in Escherichi...More data for this Ligand-Target Pair
Affinity DataKi: 47nMAssay Description:Binding affinity against Peroxisome proliferator activated receptor gamma (PPAR gamma)More data for this Ligand-Target Pair
Affinity DataKi: 59nMAssay Description:Displacement of fluormone PPAR green tracer ligand from recombinant GST-tagged PPARgamma ligand binding domain (unknown origin) incubated for 4 hrs b...More data for this Ligand-Target Pair

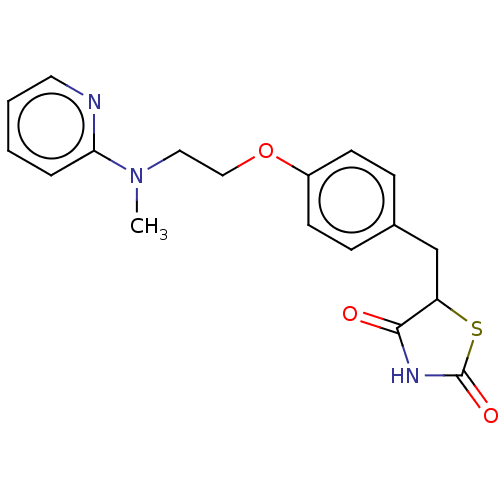
 3D Structure (crystal)
3D Structure (crystal)