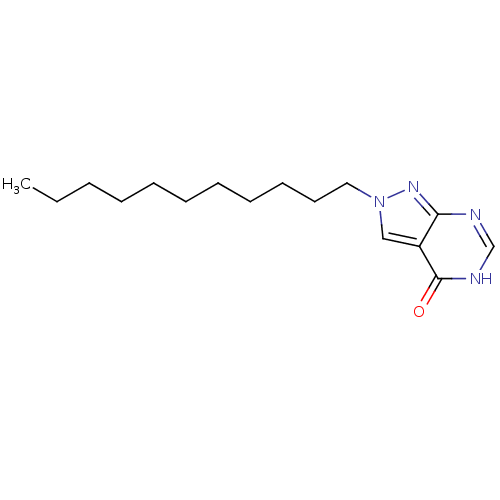
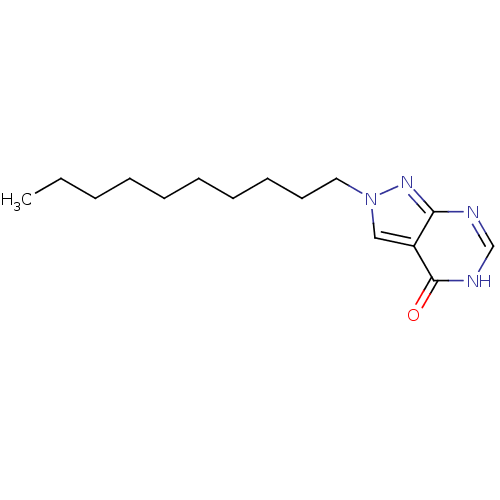
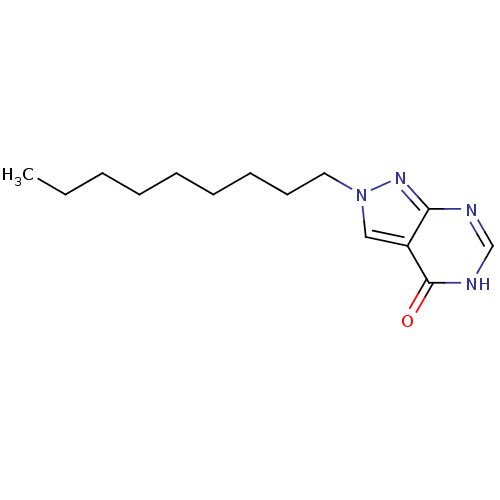
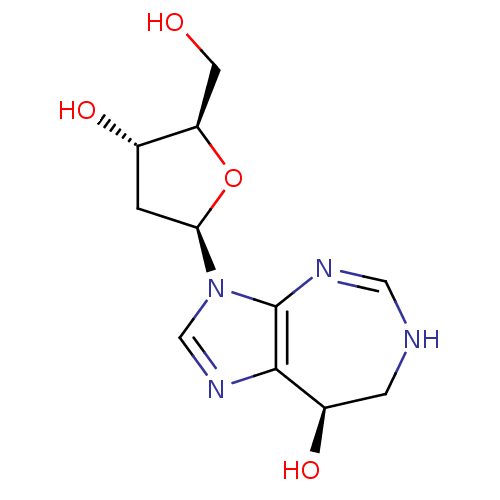
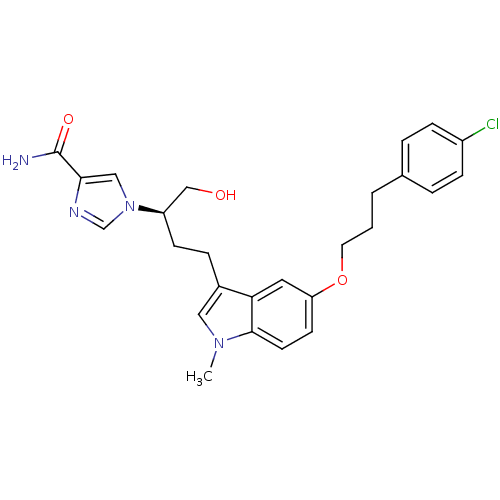
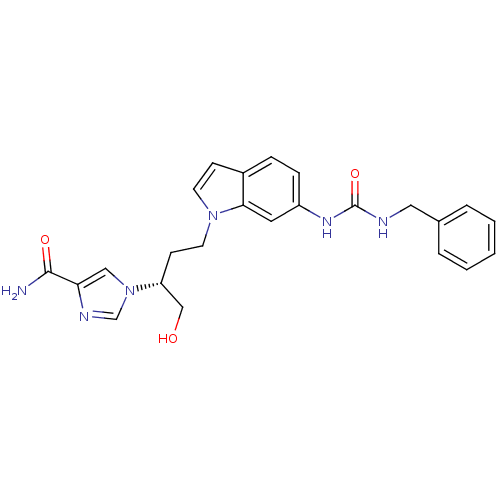
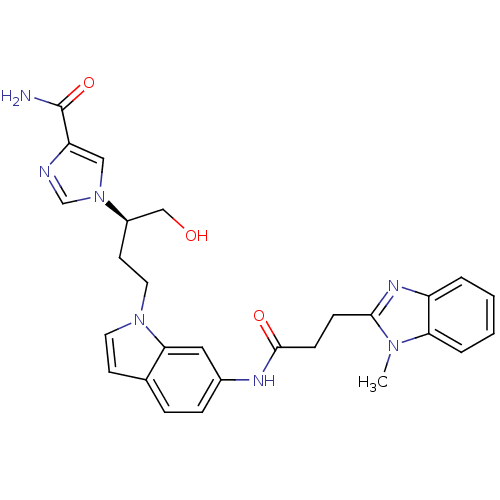
Affinity DataKi: 4.00E+3nMpH: 7.0Assay Description:Inhibitory constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined before heatingMore data for this Ligand-Target Pair
Affinity DataKi: 5.00E+3nMpH: 7.0Assay Description:Inhibitor constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0, was determined after heating to 100 degrees celsiusMore data for this Ligand-Target Pair
Affinity DataKi: 8.00E+3nMpH: 7.0Assay Description:Inhibitor constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined after heating to 100 degrees celsiusMore data for this Ligand-Target Pair
Affinity DataKi: 1.06E+4nM ΔG°: -28.4kJ/molepH: 7.0 T: 2°CAssay Description:The target compounds were screened against calf intestine ADA in vitro by monitoring the rate of hydrolyzing adenosine into inosine spectrophotometri...More data for this Ligand-Target Pair
Affinity DataKi: 1.21E+4nM ΔG°: -28.1kJ/molepH: 7.0 T: 2°CAssay Description:The target compounds were screened against calf intestine ADA in vitro by monitoring the rate of hydrolyzing adenosine into inosine spectrophotometri...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nMpH: 7.0Assay Description:Inhibitory constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0, was determined before heatingMore data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nMpH: 7.0Assay Description:Inhibitor constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined after heating to 100 degrees celsiusMore data for this Ligand-Target Pair
Affinity DataKi: 2.10E+4nMpH: 7.0Assay Description:Inhibitor constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined after heating to 100 degrees celsiusMore data for this Ligand-Target Pair
Affinity DataKi: 2.10E+4nMpH: 7.0Assay Description:Inhibitor constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined after heating to 100 degrees celsiusMore data for this Ligand-Target Pair
Affinity DataKi: 2.65E+4nM ΔG°: -26.1kJ/molepH: 7.0 T: 2°CAssay Description:The target compounds were screened against calf intestine ADA in vitro by monitoring the rate of hydrolyzing adenosine into inosine spectrophotometri...More data for this Ligand-Target Pair
Affinity DataKi: 2.80E+4nMpH: 7.0Assay Description:Inhibitory constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined before heatingMore data for this Ligand-Target Pair
Affinity DataKi: 2.80E+4nMpH: 7.0Assay Description:Inhibitory constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined before heatingMore data for this Ligand-Target Pair
Affinity DataKi: 3.00E+4nMpH: 7.0Assay Description:Inhibitor constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined after heating to 100 degrees celsiusMore data for this Ligand-Target Pair
Affinity DataKi: 3.00E+4nMpH: 7.0Assay Description:Inhibitory constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined before heatingMore data for this Ligand-Target Pair
Affinity DataKi: 3.80E+4nMpH: 7.0Assay Description:Inhibitor constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined after heating to 100 degrees celsiusMore data for this Ligand-Target Pair
Affinity DataKi: 4.60E+4nMpH: 7.0Assay Description:Inhibitory constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined before heatingMore data for this Ligand-Target Pair
Affinity DataKi: 4.60E+4nMpH: 7.0Assay Description:Inhibitor constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined after heating to 100 degrees celsiusMore data for this Ligand-Target Pair
Affinity DataKi: 5.14E+4nM ΔG°: -24.5kJ/molepH: 7.0 T: 2°CAssay Description:The target compounds were screened against calf intestine ADA in vitro by monitoring the rate of hydrolyzing adenosine into inosine spectrophotometri...More data for this Ligand-Target Pair
Affinity DataKi: 5.20E+4nM ΔG°: -24.5kJ/molepH: 7.0 T: 2°CAssay Description:The target compounds were screened against calf intestine ADA in vitro by monitoring the rate of hydrolyzing adenosine into inosine spectrophotometri...More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+5nMpH: 7.0Assay Description:Inhibitory constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined before heatingMore data for this Ligand-Target Pair
Affinity DataKi: 1.50E+5nMpH: 7.0Assay Description:Inhibitory constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined before heatingMore data for this Ligand-Target Pair
Affinity DataKi: 3.10E+5nMpH: 7.0Assay Description:Inhibitory constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0, was determined before heatingMore data for this Ligand-Target Pair
Affinity DataKi: 3.70E+5nMpH: 7.0Assay Description:Inhibitory constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined before heatingMore data for this Ligand-Target Pair
Affinity DataKi: 5.20E+5nMpH: 7.0Assay Description:Inhibitor constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined after heating to 100 degrees celsiusMore data for this Ligand-Target Pair
Affinity DataKi: 7.80E+5nMpH: 7.0Assay Description:Inhibitory constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined before heatingMore data for this Ligand-Target Pair
Affinity DataKi: 8.00E+5nMpH: 7.0Assay Description:Inhibitory constant against adenosine deaminase in 0.05 M sodium phosphate buffer at pH 7.0 was determined before heatingMore data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -52.2kJ/molepH: 7.2 T: 2°CAssay Description:For the ADA activity assay, a reaction comprising 50 mM potassium phosphate buffer (pH 7.2), 0.1 mM Ara-A or adenosine, and 20 mg ADA was conducted a...More data for this Ligand-Target Pair
Affinity DataKi: 2.10nM ΔG°: -50.4kJ/molepH: 7.2 T: 2°CAssay Description:For the ADA activity assay, a reaction comprising 50 mM potassium phosphate buffer (pH 7.2), 0.1 mM Ara-A or adenosine, and 20 mg ADA was conducted a...More data for this Ligand-Target Pair
Affinity DataKi: 160nM ΔG°: -39.4kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 510nM ΔG°: -36.5kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 960nM ΔG°: -34.9kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 1.14E+3nM ΔG°: -34.5kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 1.15E+3nM ΔG°: -34.5kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 2.01E+3nM ΔG°: -33.1kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 2.42E+3nM ΔG°: -32.6kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 1.27E+4nM ΔG°: -28.4kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 1.56E+4nM ΔG°: -27.9kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 2.83E+4nM ΔG°: -26.4kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 4.79E+4nM ΔG°: -25.1kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 8.57E+4nM ΔG°: -23.6kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 1.08E+5nM ΔG°: -23.0kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 1.86E+5nM ΔG°: -21.6kJ/molepH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DatapH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DatapH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DatapH: 7.2 T: 2°CAssay Description:The activity of ADA was determined spectrophotometrically by monitoring the change in absorbance at 262 nm, due to the deamination of adenosine catal...More data for this Ligand-Target Pair
Affinity DataKi: 0.0330nM ΔG°: -59.2kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 4.90nM ΔG°: -47.0kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 7.5nM ΔG°: -45.9kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 7.70nM ΔG°: -45.8kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair
Affinity DataKi: 7.70nM ΔG°: -45.8kJ/molepH: 7.4 T: 2°CAssay Description:The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a...More data for this Ligand-Target Pair

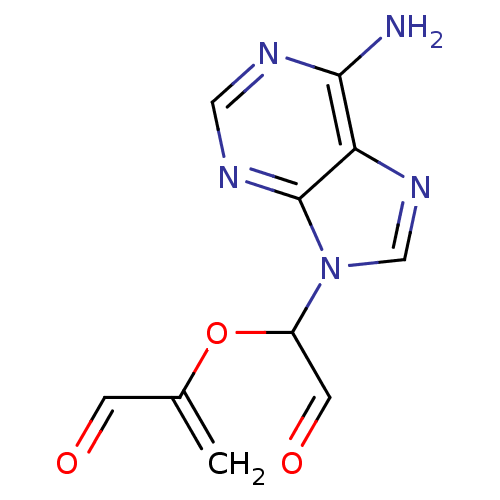
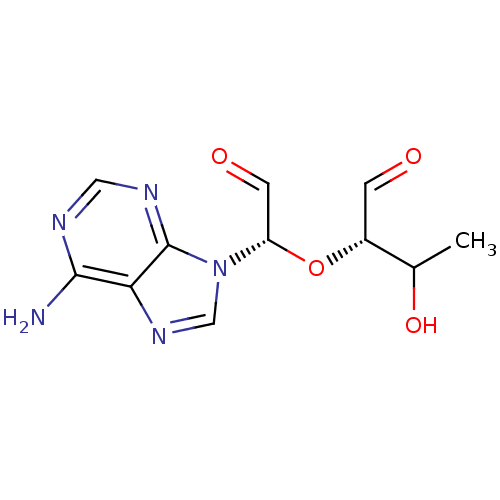
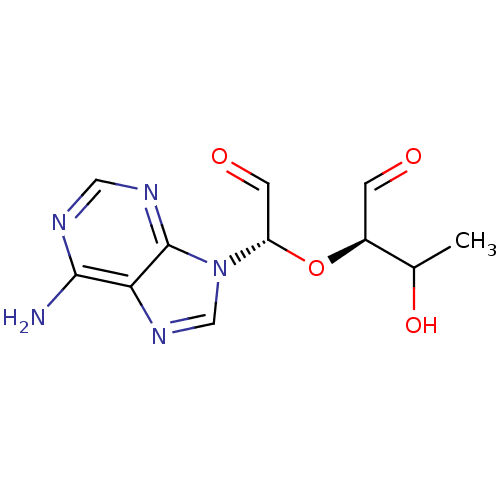
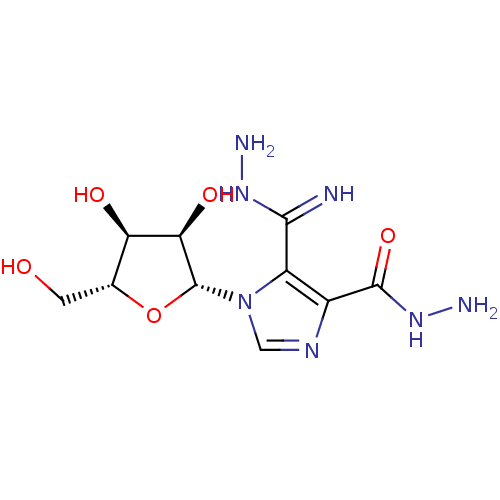
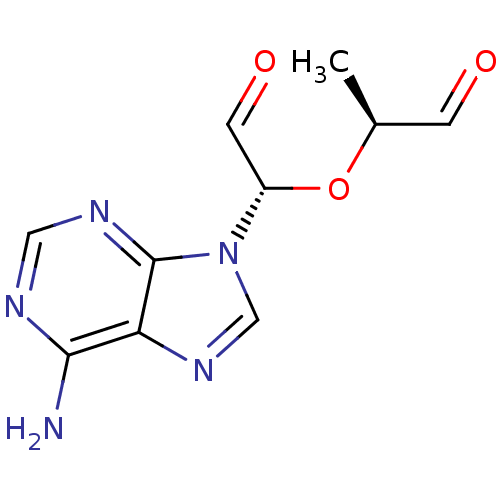
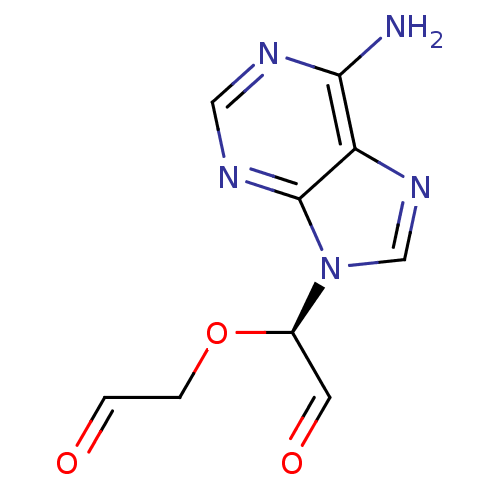
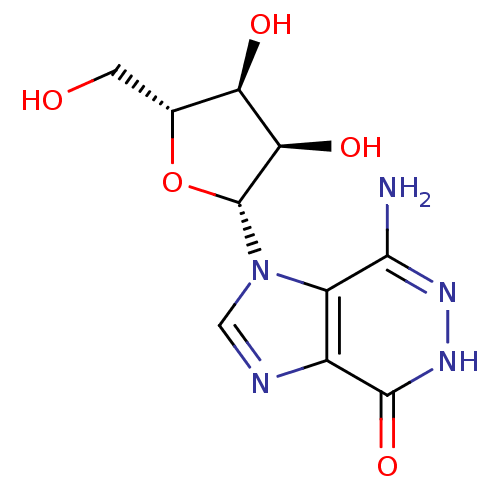
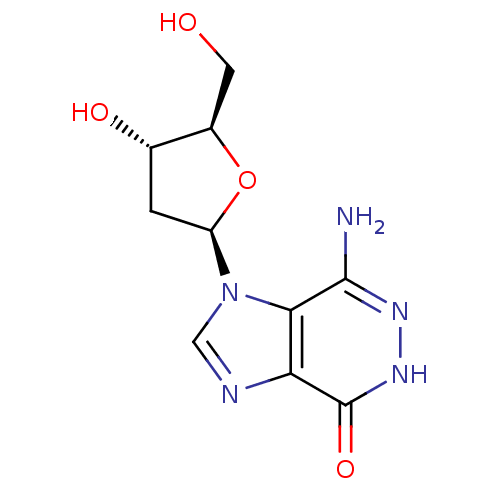
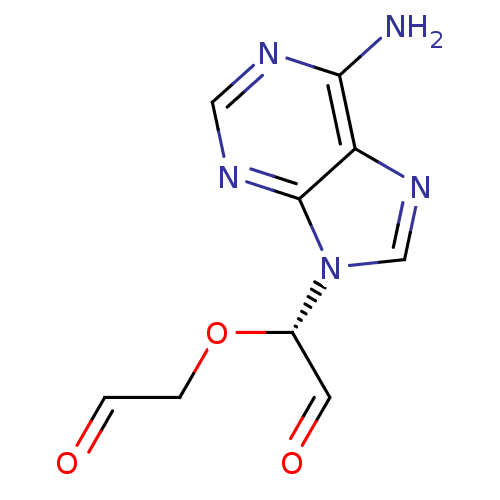

 3D Structure (crystal)
3D Structure (crystal)