TargetEnoyl-[acyl-carrier-protein] reductase [NADH] [T276S](Yersinia pestis (Enterobacteria))
University Of Wurzburg
University Of Wurzburg
Affinity DataIC50: 2.00E+3nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Yersinia pestis (Enterobacteria))
University Of Wurzburg
University Of Wurzburg
Affinity DataIC50: 5.50E+4nMAssay Description:IC50 values for the inhibition of wildtype and mutant forms of ypFabV were performed by adding varying concentrations of inhibitor dissolved in dimet...More data for this Ligand-Target Pair

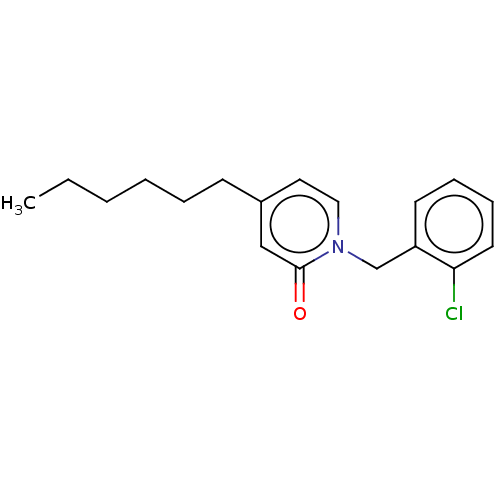
 3D Structure (crystal)
3D Structure (crystal)