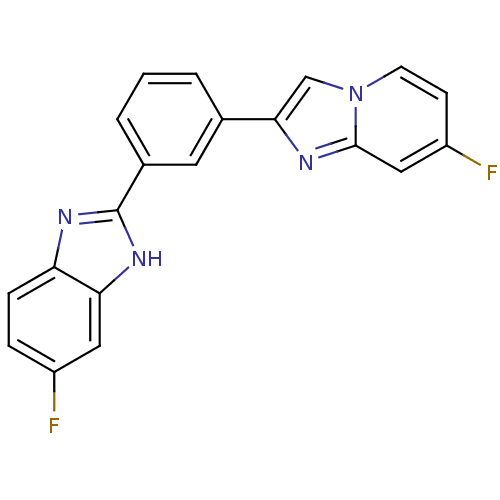
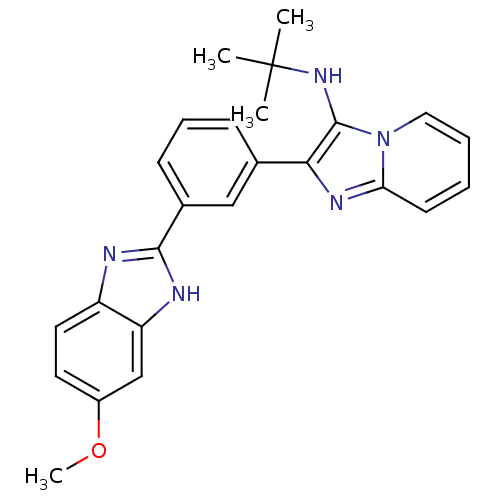
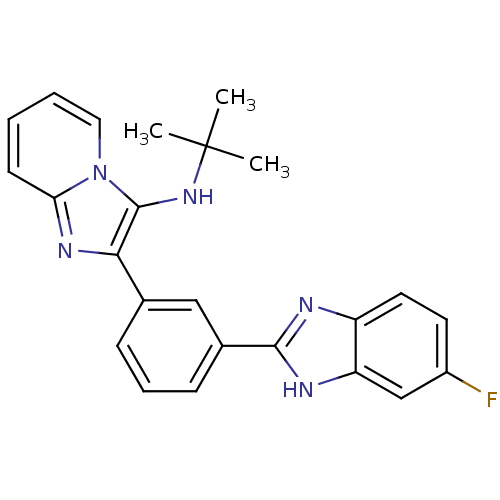
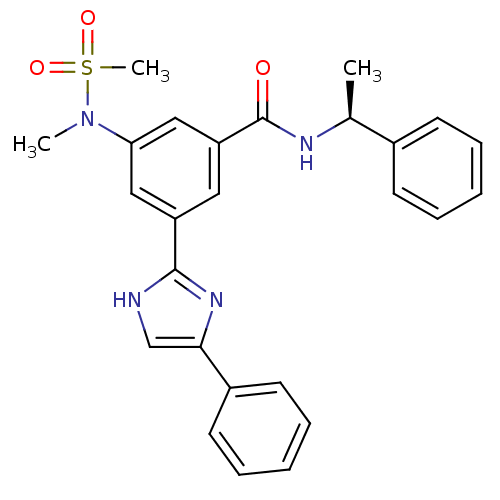
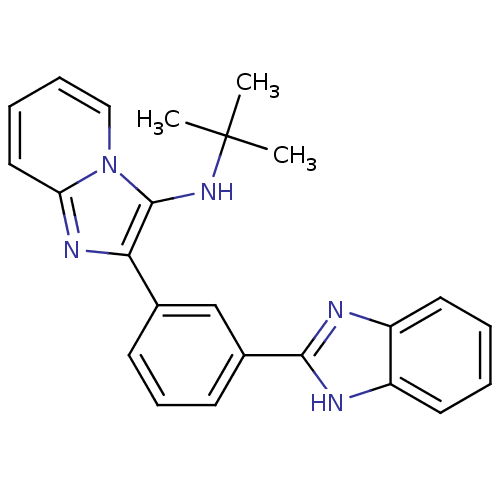
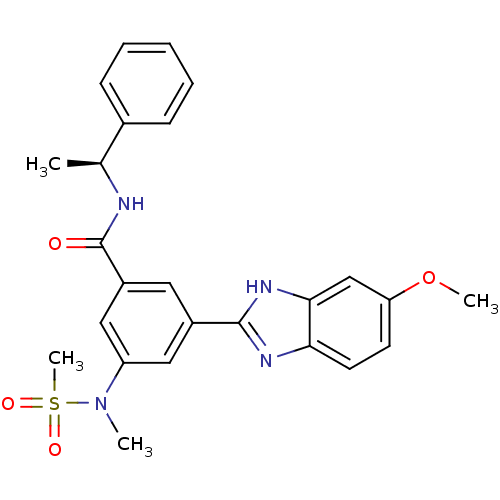
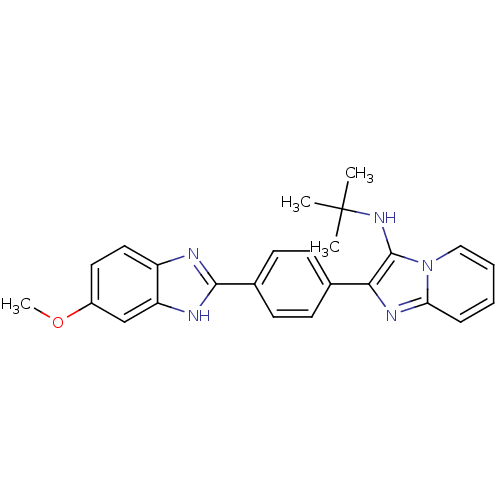
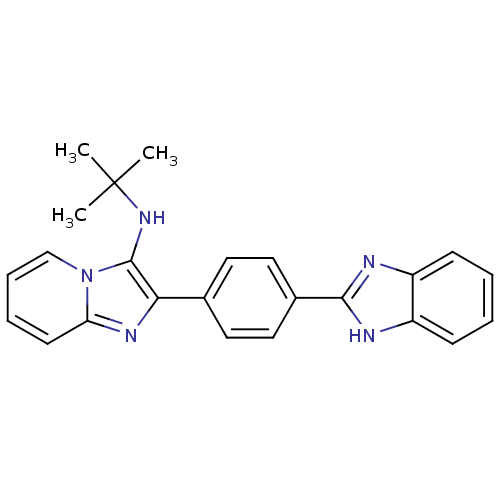
Affinity DataKi: 17nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
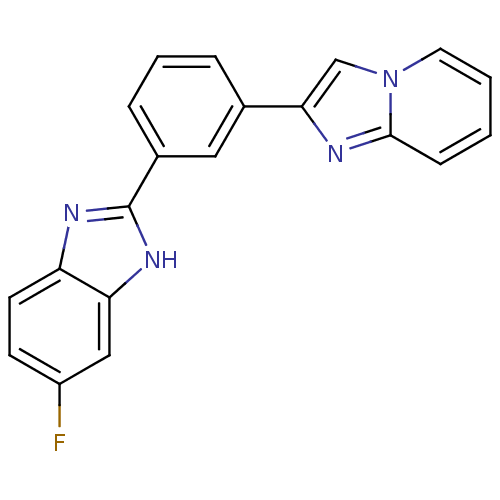
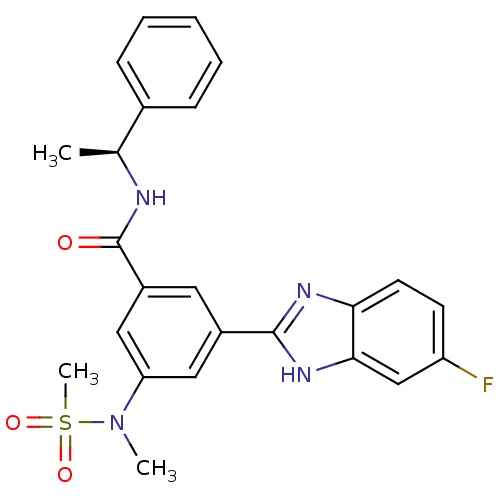
Affinity DataKi: 25nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
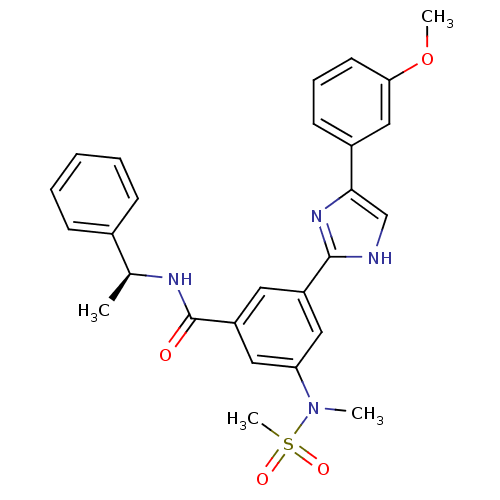
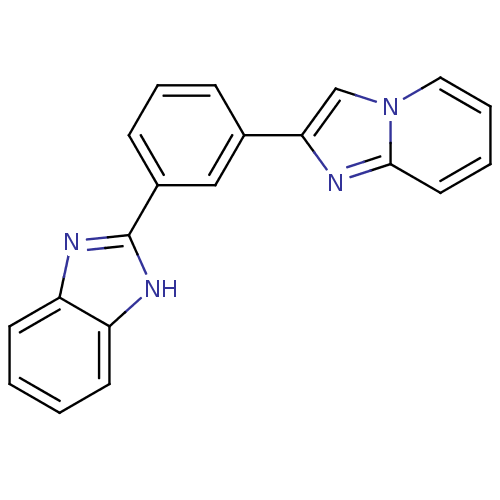
Affinity DataKi: 34nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
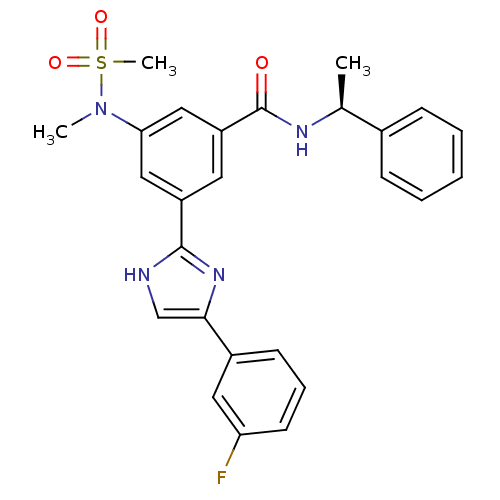
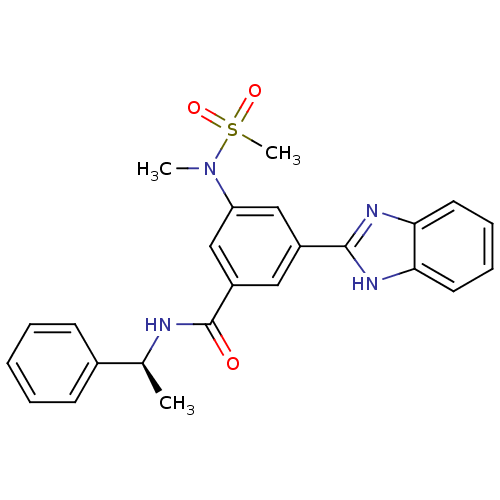
Affinity DataKi: 62nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 180nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 253nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 301nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 304nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 495nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 525nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 758nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 926nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 1.04E+3nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 1.73E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 1.82E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 2.07E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 2.07E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 2.16E+3nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 2.28E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 2.35E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 2.39E+3nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 2.81E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 3.08E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 3.22E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 3.57E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 3.62E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 3.67E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 3.78E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 4.00E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 4.11E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 5.31E+3nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 5.51E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by Michaelis menton equationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 35nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 77nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 178nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 187nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 312nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 315nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 514nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 545nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 786nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 961nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 1.08E+3nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 1.79E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 1.87E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 2.13E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 2.13E+3nMAssay Description:Inhibition of BACE2 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 2.24E+3nMAssay Description:Inhibition of BACE1 expressing APP Swedish mutant using rhodamine-labeled EVNLDAEFK-quencher as substrate after 60 mins by FRET analysisChecked by AuthorMore data for this Ligand-Target Pair