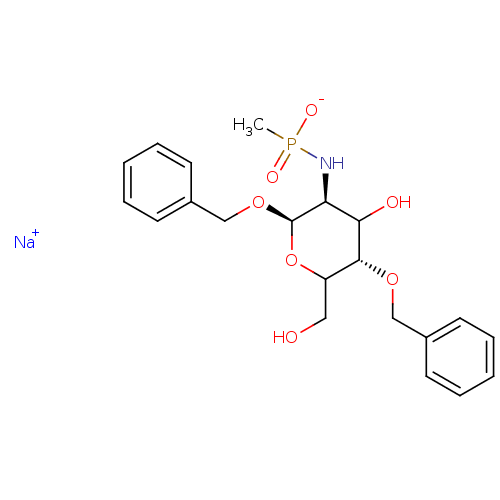
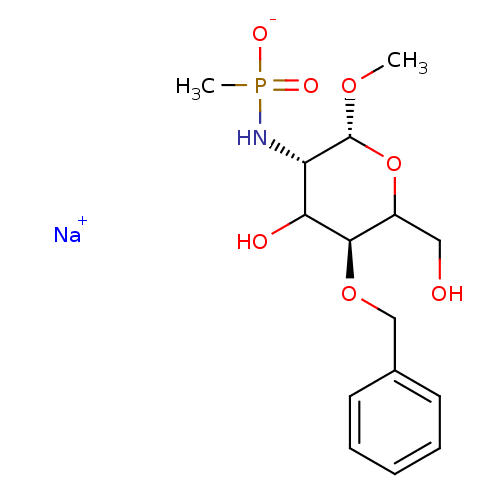
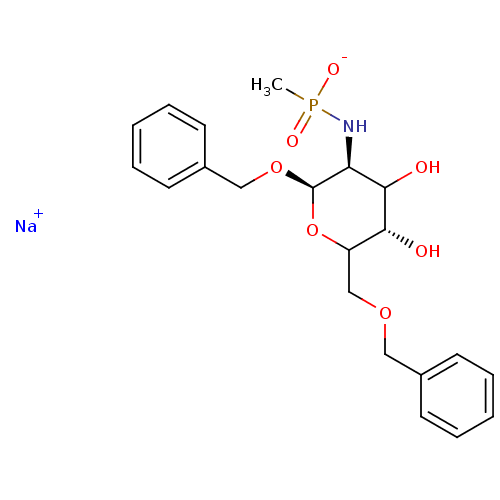
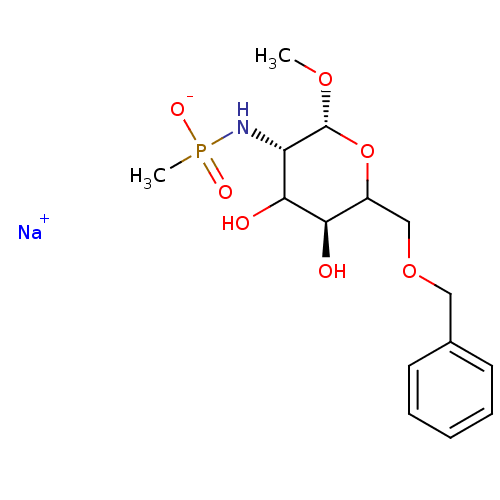
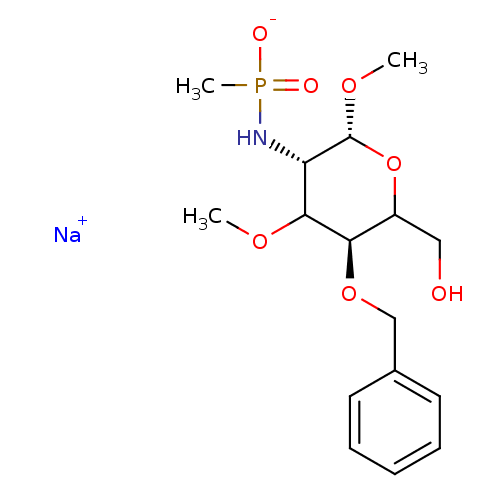
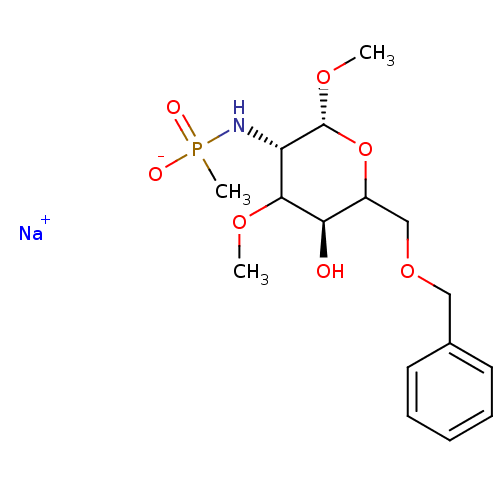
TargetPeptidoglycan-N-acetylglucosamine deacetylase(Streptococcus pneumoniae (Firmicutes))
University Of Toronto
University Of Toronto
Affinity DataKi: 8.00E+4nM ΔG°: -23.4kJ/molepH: 7.0 T: 2°CAssay Description:In a standard optimized reaction, SpPgdA was added to a microcentrifuge tube and incubated at 30 �C for 5 min. Chitotriose was added as a substrate t...More data for this Ligand-Target Pair
TargetPeptidoglycan-N-acetylglucosamine deacetylase(Streptococcus pneumoniae (Firmicutes))
University Of Toronto
University Of Toronto
Affinity DataKi: 1.70E+5nM ΔG°: -21.5kJ/molepH: 7.0 T: 2°CAssay Description:In a standard optimized reaction, SpPgdA was added to a microcentrifuge tube and incubated at 30 �C for 5 min. Chitotriose was added as a substrate t...More data for this Ligand-Target Pair
TargetPeptidoglycan-N-acetylglucosamine deacetylase(Streptococcus pneumoniae (Firmicutes))
University Of Toronto
University Of Toronto
Affinity DataKi: 2.20E+5nM ΔG°: -20.9kJ/molepH: 7.0 T: 2°CAssay Description:In a standard optimized reaction, SpPgdA was added to a microcentrifuge tube and incubated at 30 �C for 5 min. Chitotriose was added as a substrate t...More data for this Ligand-Target Pair
TargetPeptidoglycan-N-acetylglucosamine deacetylase(Streptococcus pneumoniae (Firmicutes))
University Of Toronto
University Of Toronto
Affinity DataKi: 5.80E+5nM ΔG°: -18.5kJ/molepH: 7.0 T: 2°CAssay Description:In a standard optimized reaction, SpPgdA was added to a microcentrifuge tube and incubated at 30 �C for 5 min. Chitotriose was added as a substrate t...More data for this Ligand-Target Pair
TargetPeptidoglycan-N-acetylglucosamine deacetylase(Streptococcus pneumoniae (Firmicutes))
University Of Toronto
University Of Toronto
Affinity DataKi: 7.20E+5nM ΔG°: -17.9kJ/molepH: 7.0 T: 2°CAssay Description:In a standard optimized reaction, SpPgdA was added to a microcentrifuge tube and incubated at 30 �C for 5 min. Chitotriose was added as a substrate t...More data for this Ligand-Target Pair
TargetPoly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase(Escherichia coli (Enterobacteria))
University Of Toronto
University Of Toronto
Affinity DataKi: >1.00E+6nMpH: 7.5Assay Description:Reactions were carried out in solutions (50 �L) containing PgaB (10 �M) in HEPES (100 mM, pH 7.5), and varying concentrations of ACC (0.05-10 mM) at ...More data for this Ligand-Target Pair
TargetPoly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase(Escherichia coli (Enterobacteria))
University Of Toronto
University Of Toronto
Affinity DataKi: >1.00E+6nMpH: 7.5Assay Description:Reactions were carried out in solutions (50 �L) containing PgaB (10 �M) in HEPES (100 mM, pH 7.5), and varying concentrations of ACC (0.05-10 mM) at ...More data for this Ligand-Target Pair
TargetPoly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase(Escherichia coli (Enterobacteria))
University Of Toronto
University Of Toronto
Affinity DataKi: >1.00E+6nMpH: 7.5Assay Description:Reactions were carried out in solutions (50 �L) containing PgaB (10 �M) in HEPES (100 mM, pH 7.5), and varying concentrations of ACC (0.05-10 mM) at ...More data for this Ligand-Target Pair
TargetPoly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase(Escherichia coli (Enterobacteria))
University Of Toronto
University Of Toronto
Affinity DataKi: >1.00E+6nMpH: 7.5Assay Description:Reactions were carried out in solutions (50 �L) containing PgaB (10 �M) in HEPES (100 mM, pH 7.5), and varying concentrations of ACC (0.05-10 mM) at ...More data for this Ligand-Target Pair
TargetPoly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase(Escherichia coli (Enterobacteria))
University Of Toronto
University Of Toronto
Affinity DataKi: >1.00E+6nMpH: 7.5Assay Description:Reactions were carried out in solutions (50 �L) containing PgaB (10 �M) in HEPES (100 mM, pH 7.5), and varying concentrations of ACC (0.05-10 mM) at ...More data for this Ligand-Target Pair