Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Voltage-dependent N-type calcium channel subunit alpha-1B
Ligand
BDBM34648
Substrate
n/a
Meas. Tech.
Homogeneous Time-Resolved Fluorescence Resonance Energy Transfer (HTRF) Assay
IC50
10600±n/a nM
Citation
 PubChem, PC Homogeneous Time-Resolved Fluorescence Resonance Energy Transfer (HTRF) Assay PubChem Bioassay (2009)[AID]
PubChem, PC Homogeneous Time-Resolved Fluorescence Resonance Energy Transfer (HTRF) Assay PubChem Bioassay (2009)[AID] More Info.:
Target
Name:
Voltage-dependent N-type calcium channel subunit alpha-1B
Synonyms:
BIII | Brain calcium channel III | CAC1B_HUMAN | CACH5 | CACNA1B | CACNL1A5 | Calcium channel (Type N) | Calcium channel, L type, alpha-1 polypeptide isoform 5 | Voltage-dependent N-type calcium channel subunit alpha-1B | Voltage-dependent N-type calcium channel subunit alpha-1B/Voltage-dependent calcium channel subunit alpha-2/delta-1/Voltage-dependent L-type calcium channel subunit beta-3 | Voltage-gated N-type calcium channel alpha-1B subunit | Voltage-gated N-type calcium channel alpha-1B subunit/Amyloid beta A4 precursor protein-binding family A member 1 | Voltage-gated calcium channel | Voltage-gated calcium channel subunit alpha Cav2.2 | Voltage-gated calcium channel subunit alpha Cav2.2 ((alpha 1B, beta 1b, alpha 2 delta-1) | calcium channel, voltage-dependent, N type, alpha 1B subunit
Type:
Enzyme
Mol. Mass.:
262548.16
Organism:
Homo sapiens (Human)
Description:
Q00975
Residue:
2339
Sequence:
MVRFGDELGGRYGGPGGGERARGGGAGGAGGPGPGGLQPGQRVLYKQSIAQRARTMALYNPIPVKQNCFTVNRSLFVFSEDNVVRKYAKRITEWPPFEYMILATIIANCIVLALEQHLPDGDKTPMSERLDDTEPYFIGIFCFEAGIKIIALGFVFHKGSYLRNGWNVMDFVVVLTGILATAGTDFDLRTLRAVRVLRPLKLVSGIPSLQVVLKSIMKAMVPLLQIGLLLFFAILMFAIIGLEFYMGKFHKACFPNSTDAEPVGDFPCGKEAPARLCEGDTECREYWPGPNFGITNFDNILFAILTVFQCITMEGWTDILYNTNDAAGNTWNWLYFIPLIIIGSFFMLNLVLGVLSGEFAKERERVENRRAFLKLRRQQQIERELNGYLEWIFKAEEVMLAEEDRNAEEKSPLDVLKRAATKKSRNDLIHAEEGEDRFADLCAVGSPFARASLKSGKTESSSYFRRKEKMFRFFIRRMVKAQSFYWVVLCVVALNTLCVAMVHYNQPRRLTTTLYFAEFVFLGLFLTEMSLKMYGLGPRSYFRSSFNCFDFGVIVGSVFEVVWAAIKPGSSFGISVLRALRLLRIFKVTKYWSSLRNLVVSLLNSMKSIISLLFLLFLFIVVFALLGMQLFGGQFNFQDETPTTNFDTFPAAILTVFQILTGEDWNAVMYHGIESQGGVSKGMFSSFYFIVLTLFGNYTLLNVFLAIAVDNLANAQELTKDEEEMEEAANQKLALQKAKEVAEVSPMSAANISIAARQQNSAKARSVWEQRASQLRLQNLRASCEALYSEMDPEERLRFATTRHLRPDMKTHLDRPLVVELGRDGARGPVGGKARPEAAEAPEGVDPPRRHHRHRDKDKTPAAGDQDRAEAPKAESGEPGAREERPRPHRSHSKEAAGPPEARSERGRGPGPEGGRRHHRRGSPEEAAEREPRRHRAHRHQDPSKECAGAKGERRARHRGGPRAGPREAESGEEPARRHRARHKAQPAHEAVEKETTEKEATEKEAEIVEADKEKELRNHQPREPHCDLETSGTVTVGPMHTLPSTCLQKVEEQPEDADNQRNVTRMGSQPPDPNTIVHIPVMLTGPLGEATVVPSGNVDLESQAEGKKEVEADDVMRSGPRPIVPYSSMFCLSPTNLLRRFCHYIVTMRYFEVVILVVIALSSIALAAEDPVRTDSPRNNALKYLDYIFTGVFTFEMVIKMIDLGLLLHPGAYFRDLWNILDFIVVSGALVAFAFSGSKGKDINTIKSLRVLRVLRPLKTIKRLPKLKAVFDCVVNSLKNVLNILIVYMLFMFIFAVIAVQLFKGKFFYCTDESKELERDCRGQYLDYEKEEVEAQPRQWKKYDFHYDNVLWALLTLFTVSTGEGWPMVLKHSVDATYEEQGPSPGYRMELSIFYVVYFVVFPFFFVNIFVALIIITFQEQGDKVMSECSLEKNERACIDFAISAKPLTRYMPQNRQSFQYKTWTFVVSPPFEYFIMAMIALNTVVLMMKFYDAPYEYELMLKCLNIVFTSMFSMECVLKIIAFGVLNYFRDAWNVFDFVTVLGSITDILVTEIAETNNFINLSFLRLFRAARLIKLLRQGYTIRILLWTFVQSFKALPYVCLLIAMLFFIYAIIGMQVFGNIALDDDTSINRHNNFRTFLQALMLLFRSATGEAWHEIMLSCLSNQACDEQANATECGSDFAYFYFVSFIFLCSFLMLNLFVAVIMDNFEYLTRDSSILGPHHLDEFIRVWAEYDPAACGRISYNDMFEMLKHMSPPLGLGKKCPARVAYKRLVRMNMPISNEDMTVHFTSTLMALIRTALEIKLAPAGTKQHQCDAELRKEISVVWANLPQKTLDLLVPPHKPDEMTVGKVYAALMIFDFYKQNKTTRDQMQQAPGGLSQMGPVSLFHPLKATLEQTQPAVLRGARVFLRQKSSTSLSNGGAIQNQESGIKESVSWGTQRTQDAPHEARPPLERGHSTEIPVGRSGALAVDVQMQSITRRGPDGEPQPGLESQGRAASMPRLAAETQPVTDASPMKRSISTLAQRPRGTHLCSTTPDRPPPSQASSHHHHHRCHRRRDRKQRSLEKGPSLSADMDGAPSSAVGPGLPPGEGPTGCRRERERRQERGRSQERRQPSSSSSEKQRFYSCDRFGGREPPKPKPSLSSHPTSPTAGQEPGPHPQGSGSVNGSPLLSTSGASTPGRGGRRQLPQTPLTPRPSITYKTANSSPIHFAGAQTSLPAFSPGRLSRGLSEHNALLQRDPLSQPLAPGSRIGSDPYLGQRLDSEASVHALPEDTLTFEEAVATNSGRSSRTSYVSSLTSQSHPLRRVPNGYHCTLGLSSGGRARHSYHHPDQDHWC
Inhibitor
Name:
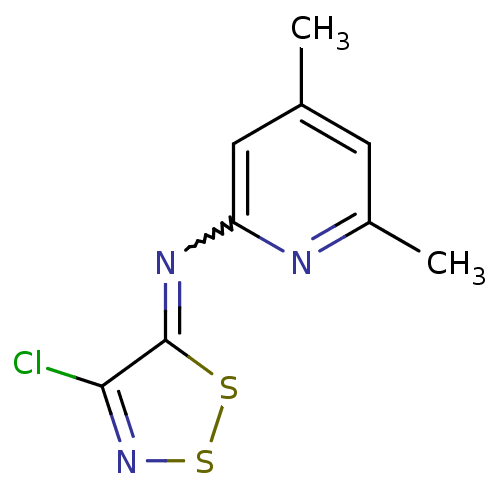
BDBM34648
Synonyms:
(4-chlorodithiazol-5-ylidene)-(4,6-dimethyl-2-pyridyl)amine | 4-chloranyl-N-(4,6-dimethylpyridin-2-yl)-1,2,3-dithiazol-5-imine | 4-chloro-N-(4,6-dimethyl-2-pyridinyl)-5-dithiazolimine | 4-chloro-N-(4,6-dimethylpyridin-2-yl)dithiazol-5-imine | MLS000104333 | SMR000054268 | cid_757789
Type:
Small organic molecule
Emp. Form.:
C9H8ClN3S2
Mol. Mass.:
257.763
SMILES:
Cc1cc(C)nc(c1)N=c1ssnc1Cl |w:8.8|