Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Glycogen phosphorylase, liver form
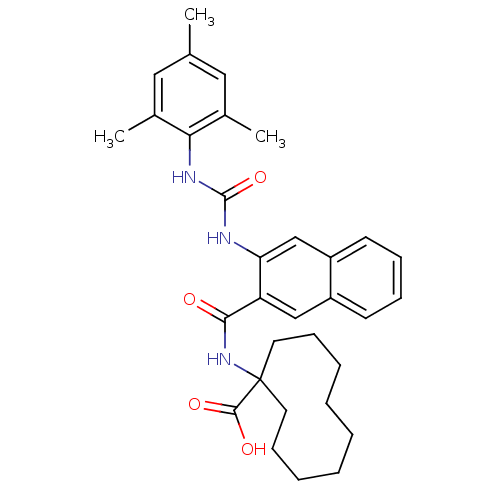
Ligand
BDBM50256330
Substrate
n/a
Meas. Tech.
ChEMBL_515050 (CHEMBL1034792)
IC50
14±n/a nM
Citation
 Sparks, SM; Banker, P; Bickett, DM; Carter, HL; Clancy, DC; Dickerson, SH; Dwornik, KA; Garrido, DM; Golden, PL; Nolte, RT; Peat, AJ; Sheckler, LR; Tavares, FX; Thomson, SA; Wang, L; Weiel, JE Anthranilimide-based glycogen phosphorylase inhibitors for the treatment of type 2 diabetes: 1. Identification of 1-amino-1-cycloalkyl carboxylic acid headgroups. Bioorg Med Chem Lett 19:976-80 (2009) [PubMed] Article
Sparks, SM; Banker, P; Bickett, DM; Carter, HL; Clancy, DC; Dickerson, SH; Dwornik, KA; Garrido, DM; Golden, PL; Nolte, RT; Peat, AJ; Sheckler, LR; Tavares, FX; Thomson, SA; Wang, L; Weiel, JE Anthranilimide-based glycogen phosphorylase inhibitors for the treatment of type 2 diabetes: 1. Identification of 1-amino-1-cycloalkyl carboxylic acid headgroups. Bioorg Med Chem Lett 19:976-80 (2009) [PubMed] Article More Info.:
Target
Name:
Glycogen phosphorylase, liver form
Synonyms:
Glycogen Phosphorylase (PYGL) | Glycogen Phosphorylase, liver form | Liver glycogen phosphorylase | PYGL | PYGL_HUMAN
Type:
Homodimer
Mol. Mass.:
97153.98
Organism:
Homo sapiens (Human)
Description:
Dimers associate into a tetramer to form the enzymatically active phosphorylase A.
Residue:
847
Sequence:
MAKPLTDQEKRRQISIRGIVGVENVAELKKSFNRHLHFTLVKDRNVATTRDYYFALAHTVRDHLVGRWIRTQQHYYDKCPKRVYYLSLEFYMGRTLQNTMINLGLQNACDEAIYQLGLDIEELEEIEEDAGLGNGGLGRLAACFLDSMATLGLAAYGYGIRYEYGIFNQKIRDGWQVEEADDWLRYGNPWEKSRPEFMLPVHFYGKVEHTNTGTKWIDTQVVLALPYDTPVPGYMNNTVNTMRLWSARAPNDFNLRDFNVGDYIQAVLDRNLAENISRVLYPNDNFFEGKELRLKQEYFVVAATLQDIIRRFKASKFGSTRGAGTVFDAFPDQVAIQLNDTHPALAIPELMRIFVDIEKLPWSKAWELTQKTFAYTNHTVLPEALERWPVDLVEKLLPRHLEIIYEINQKHLDRIVALFPKDVDRLRRMSLIEEEGSKRINMAHLCIVGSHAVNGVAKIHSDIVKTKVFKDFSELEPDKFQNKTNGITPRRWLLLCNPGLAELIAEKIGEDYVKDLSQLTKLHSFLGDDVFLRELAKVKQENKLKFSQFLETEYKVKINPSSMFDVQVKRIHEYKRQLLNCLHVITMYNRIKKDPKKLFVPRTVIIGGKAAPGYHMAKMIIKLITSVADVVNNDPMVGSKLKVIFLENYRVSLAEKVIPATDLSEQISTAGTEASGTGNMKFMLNGALTIGTMDGANVEMAEEAGEENLFIFGMRIDDVAALDKKGYEAKEYYEALPELKLVIDQIDNGFFSPKQPDLFKDIINMLFYHDRFKVFADYEAYVKCQDKVSQLYMNPKAWNTMVLKNIAASGKFSSDRTIKEYAQNIWNVEPSDLKISLSNESNKVNGN