Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Histone-lysine N-methyltransferase EHMT1
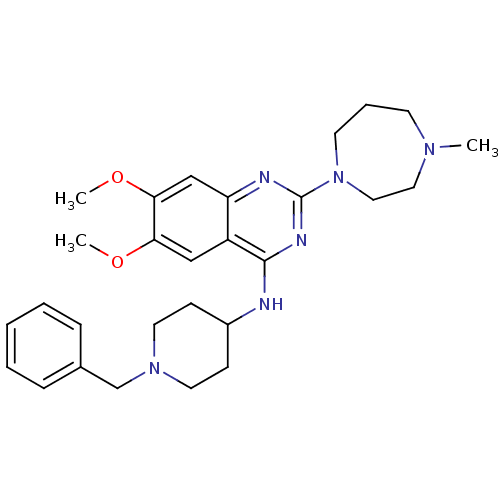
Ligand
BDBM50300028
Substrate
n/a
Meas. Tech.
ChEMBL_647514 (CHEMBL1217454)
IC50
27±n/a nM
Citation
 Liu, F; Chen, X; Allali-Hassani, A; Quinn, AM; Wigle, TJ; Wasney, GA; Dong, A; Senisterra, G; Chau, I; Siarheyeva, A; Norris, JL; Kireev, DB; Jadhav, A; Herold, JM; Janzen, WP; Arrowsmith, CH; Frye, SV; Brown, PJ; Simeonov, A; Vedadi, M; Jin, J Protein lysine methyltransferase G9a inhibitors: design, synthesis, and structure activity relationships of 2,4-diamino-7-aminoalkoxy-quinazolines. J Med Chem 53:5844-57 (2010) [PubMed] Article
Liu, F; Chen, X; Allali-Hassani, A; Quinn, AM; Wigle, TJ; Wasney, GA; Dong, A; Senisterra, G; Chau, I; Siarheyeva, A; Norris, JL; Kireev, DB; Jadhav, A; Herold, JM; Janzen, WP; Arrowsmith, CH; Frye, SV; Brown, PJ; Simeonov, A; Vedadi, M; Jin, J Protein lysine methyltransferase G9a inhibitors: design, synthesis, and structure activity relationships of 2,4-diamino-7-aminoalkoxy-quinazolines. J Med Chem 53:5844-57 (2010) [PubMed] Article More Info.:
Target
Name:
Histone-lysine N-methyltransferase EHMT1
Synonyms:
EHMT1 | EHMT1_HUMAN | EUHMTASE1 | Eu-HMTase1 | Euchromatic histone-lysine N-methyltransferase 1 | G9a-like protein 1 | GLP | GLP1 | H3-K9-HMTase 5 | Histone H3-K9 methyltransferase 5 | Histone-lysine N-methyltransferase EHMT1/EHMT2 | KIAA1876 | KMT1D | Lysine N-methyltransferase 1D
Type:
PROTEIN
Mol. Mass.:
141443.67
Organism:
Homo sapiens (Human)
Description:
ChEMBL_1450367
Residue:
1298
Sequence:
MAAADAEAVPARGEPQQDCCVKTELLGEETPMAADEGSAEKQAGEAHMAADGETNGSCENSDASSHANAAKHTQDSARVNPQDGTNTLTRIAENGVSERDSEAAKQNHVTADDFVQTSVIGSNGYILNKPALQAQPLRTTSTLASSLPGHAAKTLPGGAGKGRTPSAFPQTPAAPPATLGEGSADTEDRKLPAPGADVKVHRARKTMPKSVVGLHAASKDPREVREARDHKEPKEEINKNISDFGRQQLLPPFPSLHQSLPQNQCYMATTKSQTACLPFVLAAAVSRKKKRRMGTYSLVPKKKTKVLKQRTVIEMFKSITHSTVGSKGEKDLGASSLHVNGESLEMDSDEDDSEELEEDDGHGAEQAAAFPTEDSRTSKESMSEADRAQKMDGESEEEQESVDTGEEEEGGDESDLSSESSIKKKFLKRKGKTDSPWIKPARKRRRRSRKKPSGALGSESYKSSAGSAEQTAPGDSTGYMEVSLDSLDLRVKGILSSQAEGLANGPDVLETDGLQEVPLCSCRMETPKSREITTLANNQCMATESVDHELGRCTNSVVKYELMRPSNKAPLLVLCEDHRGRMVKHQCCPGCGYFCTAGNFMECQPESSISHRFHKDCASRVNNASYCPHCGEESSKAKEVTIAKADTTSTVTPVPGQEKGSALEGRADTTTGSAAGPPLSEDDKLQGAASHVPEGFDPTGPAGLGRPTPGLSQGPGKETLESALIALDSEKPKKLRFHPKQLYFSARQGELQKVLLMLVDGIDPNFKMEHQNKRSPLHAAAEAGHVDICHMLVQAGANIDTCSEDQRTPLMEAAENNHLEAVKYLIKAGALVDPKDAEGSTCLHLAAKKGHYEVVQYLLSNGQMDVNCQDDGGWTPMIWATEYKHVDLVKLLLSKGSDINIRDNEENICLHWAAFSGCVDIAEILLAAKCDLHAVNIHGDSPLHIAARENRYDCVVLFLSRDSDVTLKNKEGETPLQCASLNSQVWSALQMSKALQDSAPDRPSPVERIVSRDIARGYERIPIPCVNAVDSEPCPSNYKYVSQNCVTSPMNIDRNITHLQYCVCIDDCSSSNCMCGQLSMRCWYDKDGRLLPEFNMAEPPLIFECNHACSCWRNCRNRVVQNGLRARLQLYRTRDMGWGVRSLQDIPPGTFVCEYVGELISDSEADVREEDSYLFDLDNKDGEVYCIDARFYGNVSRFINHHCEPNLVPVRVFMAHQDLRFPRIAFFSTRLIEAGEQLGFDYGERFWDIKGKLFSCRCGSPKCRHSSAALAQRQASAAQEAQEDGLPDTSSAAAADPL