Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Beta-secretase 1
Ligand
BDBM50128041
Substrate
n/a
Meas. Tech.
ChEMBL_39223 (CHEMBL655069)
Ki
10491±n/a nM
Citation
 Tounge, BA; Reynolds, CH Calculation of the binding affinity of beta-secretase inhibitors using the linear interaction energy method. J Med Chem 46:2074-82 (2003) [PubMed] Article
Tounge, BA; Reynolds, CH Calculation of the binding affinity of beta-secretase inhibitors using the linear interaction energy method. J Med Chem 46:2074-82 (2003) [PubMed] Article More Info.:
Target
Name:
Beta-secretase 1
Synonyms:
ASP2 | Asp 2 | Aspartyl protease 2 | BACE | BACE1 | BACE1_HUMAN | Beta-secretase (BACE) | Beta-secretase 1 | Beta-secretase 1 (BACE 1) | Beta-secretase 1 (BACE-1) | Beta-secretase 1 (BACE1) | Beta-site APP cleaving enzyme 1 | Beta-site amyloid precursor protein cleaving enzyme 1 | KIAA1149 | Memapsin-2 | Membrane-associated aspartic protease 2 | beta-Secretase (BACE-1) | beta-Secretase (BACE1)
Type:
Protein
Mol. Mass.:
55755.10
Organism:
Homo sapiens (Human)
Description:
P56817
Residue:
501
Sequence:
MAQALPWLLLWMGAGVLPAHGTQHGIRLPLRSGLGGAPLGLRLPRETDEEPEEPGRRGSFVEMVDNLRGKSGQGYYVEMTVGSPPQTLNILVDTGSSNFAVGAAPHPFLHRYYQRQLSSTYRDLRKGVYVPYTQGKWEGELGTDLVSIPHGPNVTVRANIAAITESDKFFINGSNWEGILGLAYAEIARPDDSLEPFFDSLVKQTHVPNLFSLQLCGAGFPLNQSEVLASVGGSMIIGGIDHSLYTGSLWYTPIRREWYYEVIIVRVEINGQDLKMDCKEYNYDKSIVDSGTTNLRLPKKVFEAAVKSIKAASSTEKFPDGFWLGEQLVCWQAGTTPWNIFPVISLYLMGEVTNQSFRITILPQQYLRPVEDVATSQDDCYKFAISQSSTGTVMGAVIMEGFYVVFDRARKRIGFAVSACHVHDEFRTAAVEGPFVTLDMEDCGYNIPQTDESTLMTIAYVMAAICALFMLPLCLMVCQWRCLRCLRQQHDDFADDISLLK
Inhibitor
Name:
BDBM50128041
Synonyms:
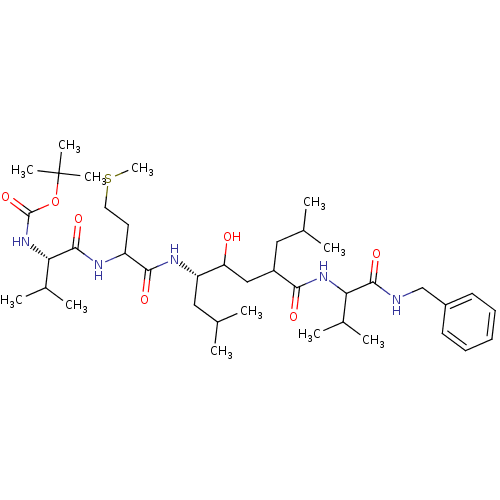
(1-{1-[4-(1-Benzylcarbamoyl-2-methyl-propylcarbamoyl)-2-hydroxy-1-isobutyl-6-methyl-heptylcarbamoyl]-3-methylsulfanyl-propylcarbamoyl}-2-methyl-propyl)-carbamic acid tert-butyl ester | CHEMBL302651
Type:
Small organic molecule
Emp. Form.:
C40H69N5O7S
Mol. Mass.:
764.07
SMILES:
CSCCC(NC(=O)[C@@H](NC(=O)OC(C)(C)C)C(C)C)C(=O)N[C@@H](CC(C)C)C(O)CC(CC(C)C)C(=O)NC(C(C)C)C(=O)NCc1ccccc1