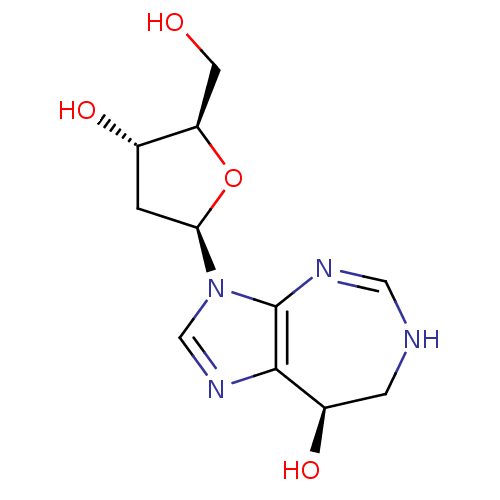
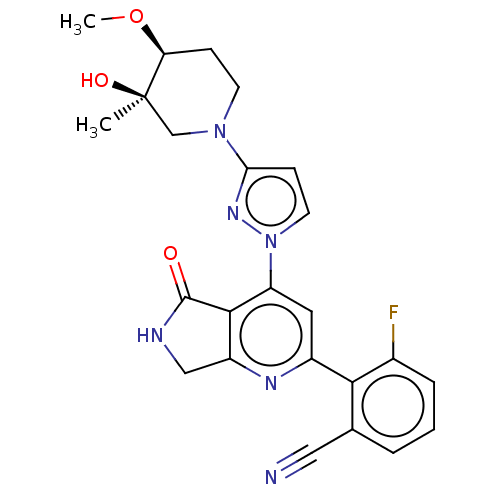
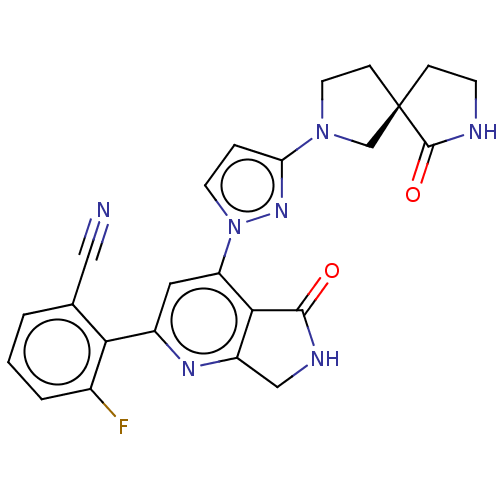
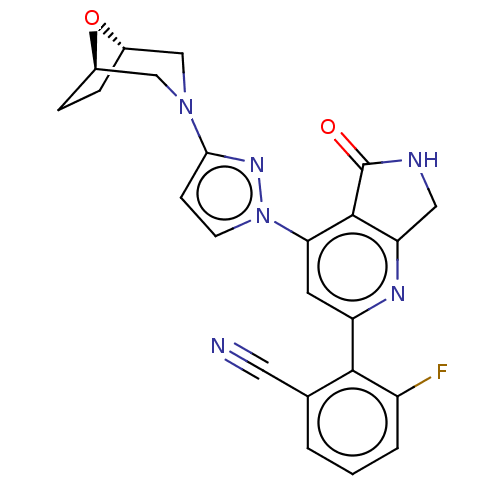
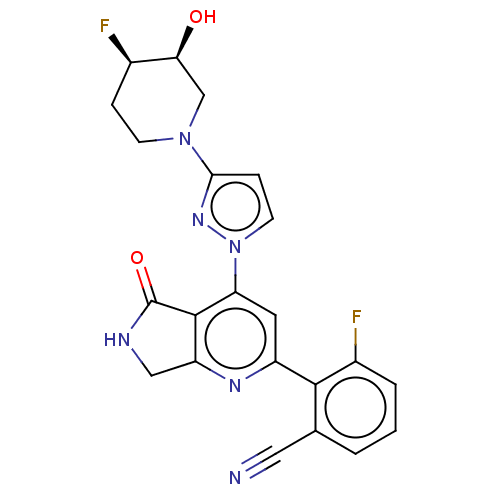
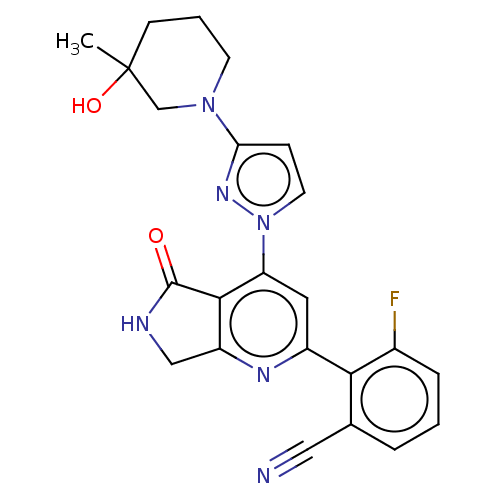
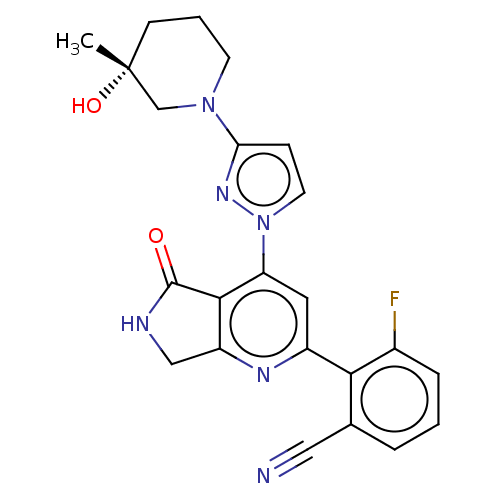
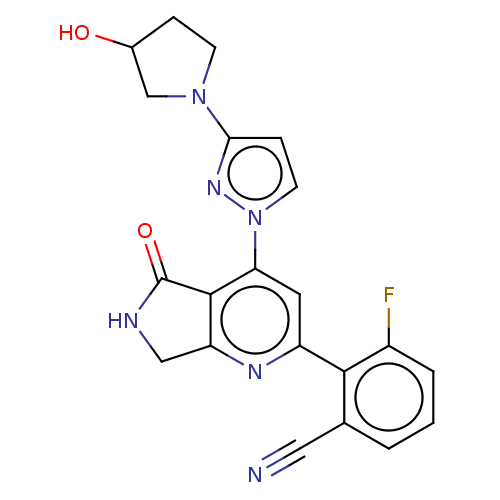
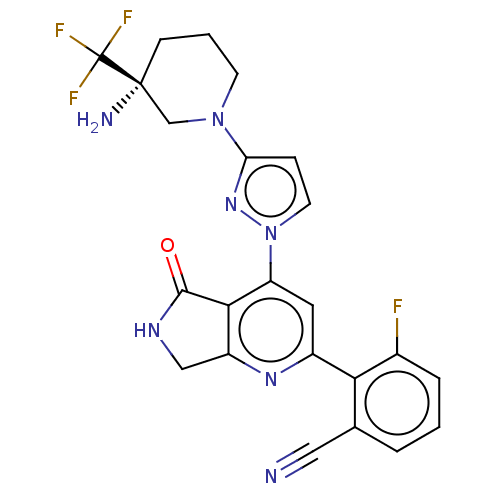
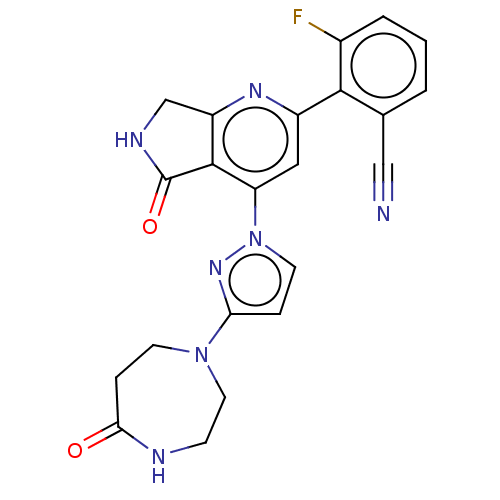
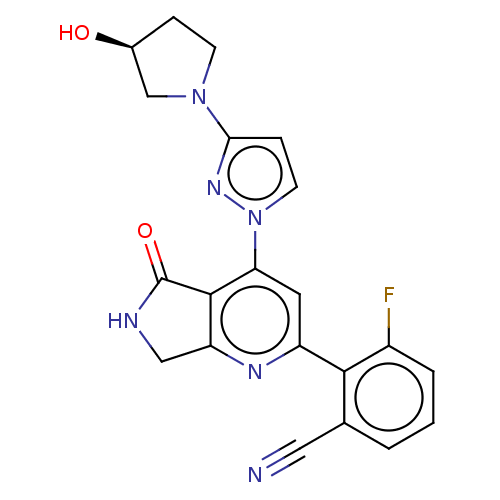
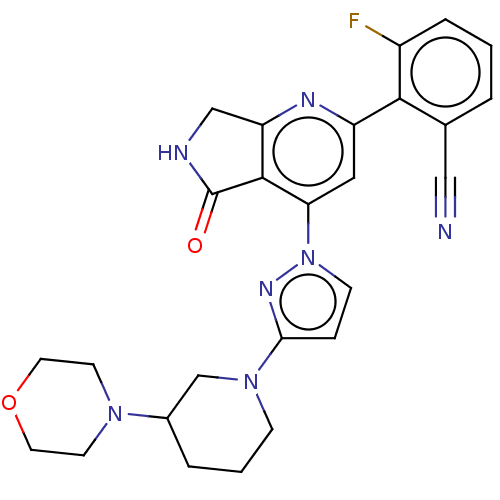
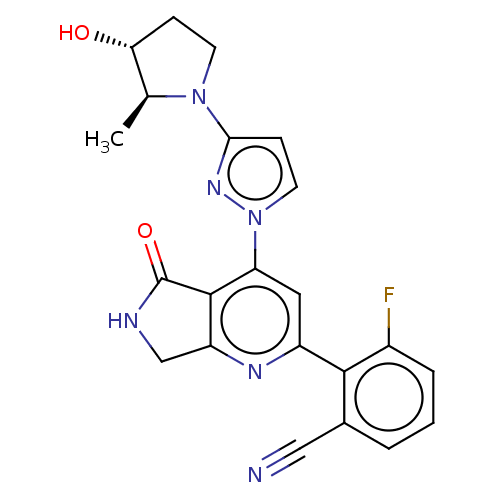
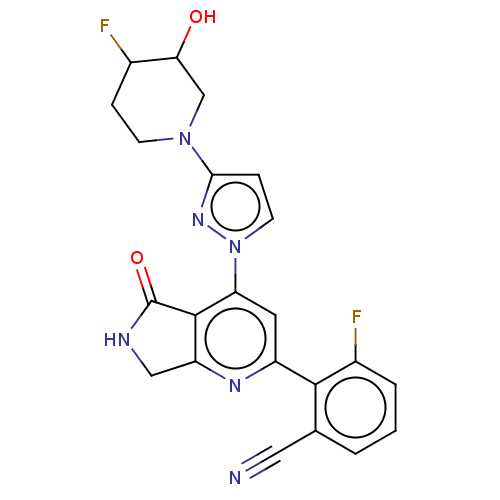
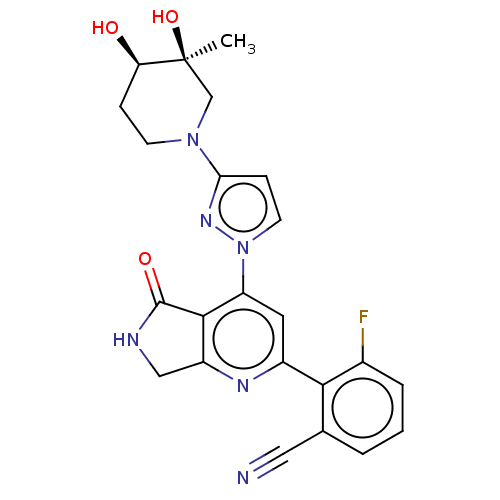
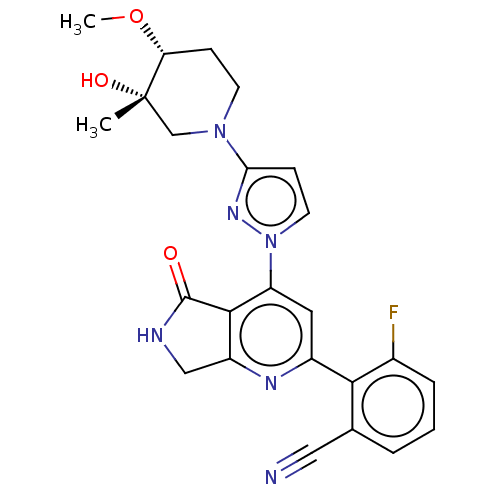
Affinity DataKi: 0.00250nMAssay Description:Binding affinity (Ki) at calf intestinal adenosine deaminase.Checked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The caliper machine employs an off chip mobility shift assay to detect phosphorylated peptide substrates from kinase assays, using microfluidics tech...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Peptide substrate, [KKSRGDYMTMQIG], (20 μM) is prepared in reaction buffer (20 mM Hepes pH 7.5, 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/mL...More data for this Ligand-Target Pair
Affinity DataKi: 0.0510nMAssay Description:Inhibition of TYK2 JH1 domain (unknown origin) measured in presence of 10 uM ATPMore data for this Ligand-Target Pair
Affinity DataKi: 0.0810nMAssay Description:Inhibition of TYK2 JH1 domain (unknown origin) measured in presence of 10 uM ATPMore data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:Inhibition of TYK2 JH1 domain (unknown origin) measured in presence of 10 uM ATPMore data for this Ligand-Target Pair
TargetC-C chemokine receptor type 8(Homo sapiens (Human))
Millennium Pharmaceuticals
Curated by ChEMBL
Millennium Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.170nMAssay Description:Binding affinity to human CCR8 expressed in L1.2 cells by FMAT assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.190nMAssay Description:Inhibition of TYK2 JH1 domain (unknown origin) measured in presence of 10 uM ATPMore data for this Ligand-Target Pair
Affinity DataKi: 0.210nMAssay Description:Inhibition of TYK2 JH1 domain (unknown origin) measured in presence of 10 uM ATPMore data for this Ligand-Target Pair
Affinity DataKi: 0.230nMAssay Description:Inhibition of TYK2 JH1 domain (unknown origin) measured in presence of 10 uM ATPMore data for this Ligand-Target Pair