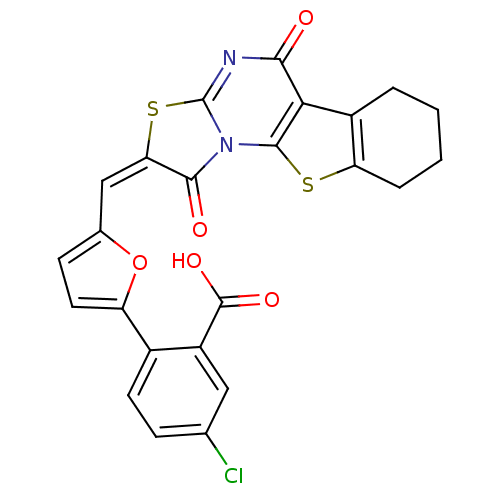
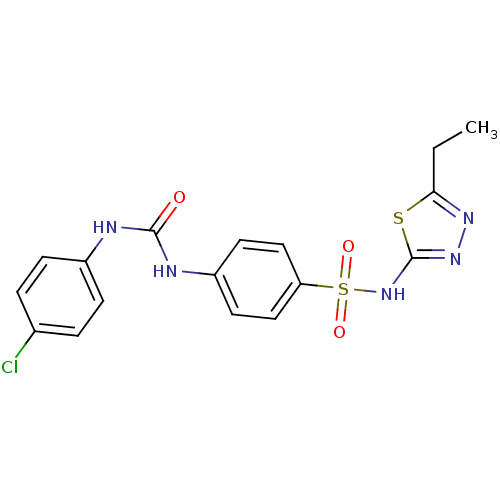
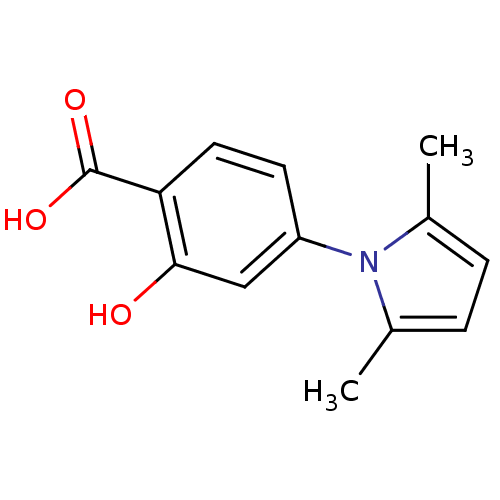
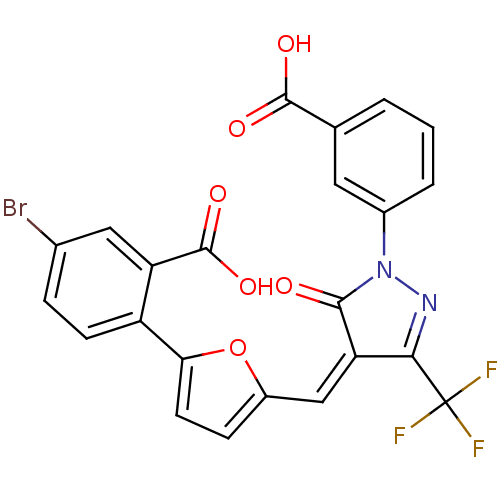
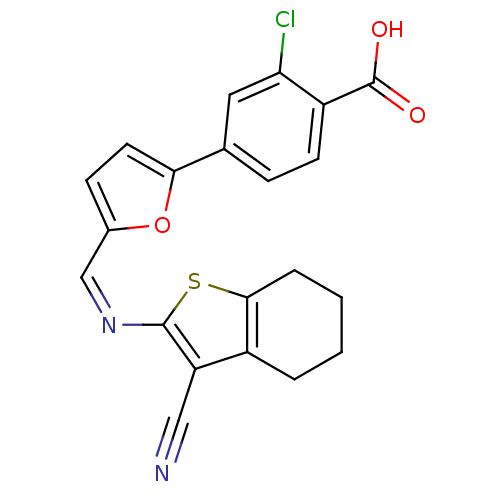
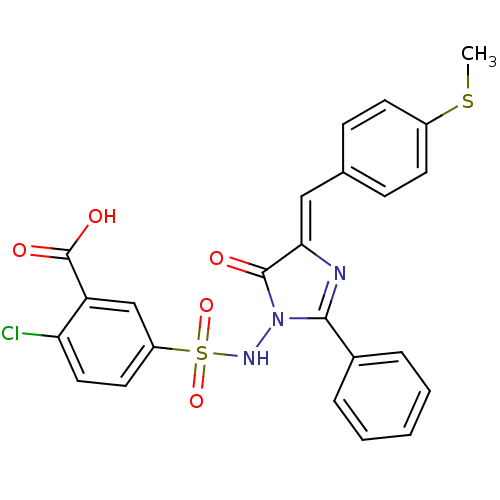
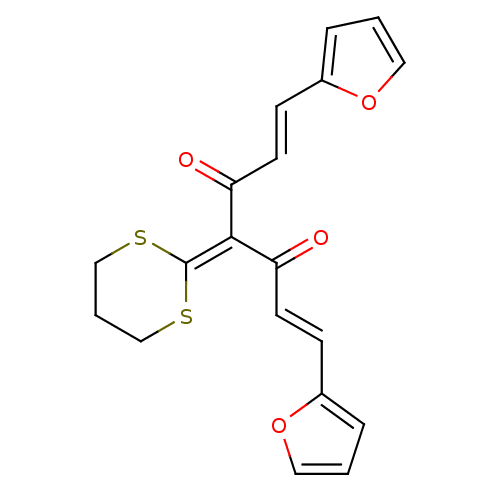
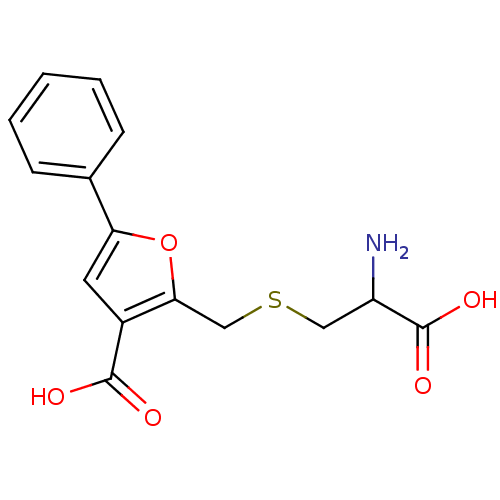
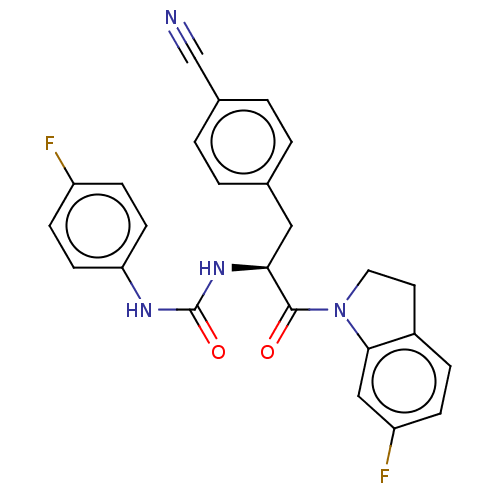
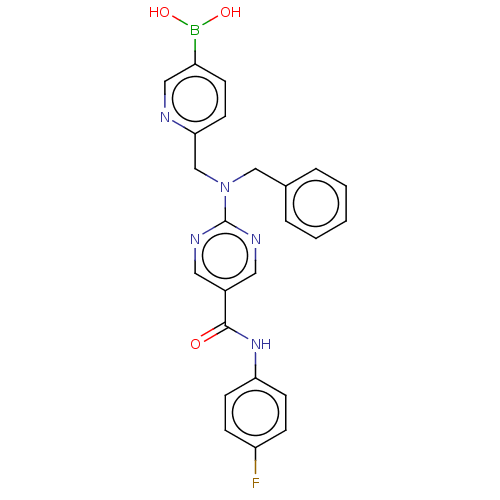
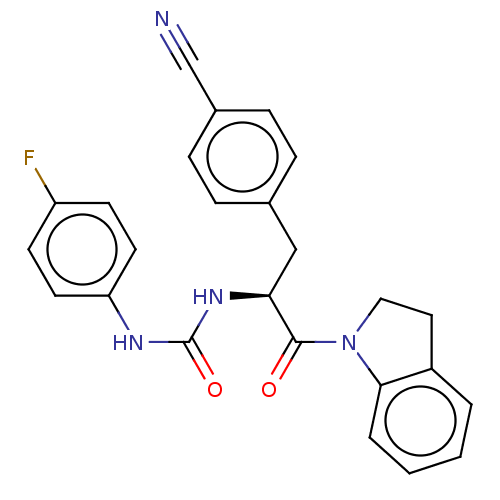
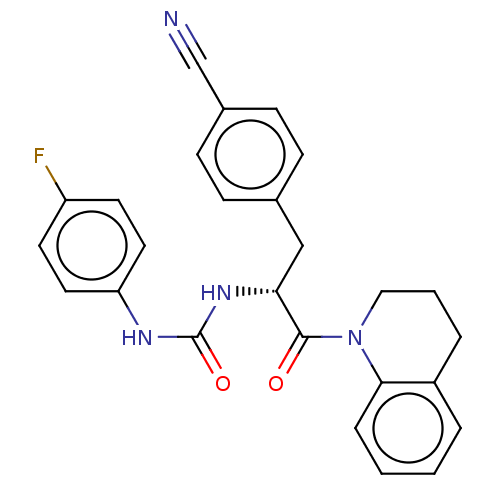
Affinity DataKi: 800nM ΔG°: -35.0kJ/mole IC50: 1.10E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 900nM ΔG°: -34.7kJ/mole IC50: 4.80E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 900nM ΔG°: -34.7kJ/mole IC50: 7.60E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, the concentrations of inhibitor that caused 50% inhibition of enzymati...More data for this Ligand-Target Pair
Affinity DataKi: 900nM ΔG°: -34.7kJ/mole IC50: 8.20E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nM ΔG°: -34.2kJ/mole IC50: 3.70E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+3nM ΔG°: -33.5kJ/mole IC50: 3.80E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 1.60E+3nM ΔG°: -33.3kJ/mole IC50: 1.80E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 1.60E+3nM ΔG°: -33.3kJ/mole IC50: 2.90E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, the concentrations of inhibitor that caused 50% inhibition of enzymati...More data for this Ligand-Target Pair
Affinity DataKi: 1.80E+3nM ΔG°: -33.0kJ/mole IC50: 8.70E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 2.10E+3nM ΔG°: -32.6kJ/mole IC50: 1.00E+4nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 2.40E+3nM ΔG°: -32.3kJ/mole IC50: 2.90E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 2.40E+3nM ΔG°: -32.3kJ/mole IC50: 5.20E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 2.50E+3nM ΔG°: -32.2kJ/mole IC50: 3.30E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 2.70E+3nM ΔG°: -32.0kJ/mole IC50: 3.20E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 2.90E+3nM ΔG°: -31.8kJ/mole IC50: 4.10E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 3.10E+3nM ΔG°: -31.7kJ/mole IC50: 4.20E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 3.30E+3nM ΔG°: -31.5kJ/mole IC50: 4.70E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 4.20E+3nM ΔG°: -30.9kJ/mole IC50: 9.20E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 5.40E+3nM ΔG°: -30.3kJ/mole IC50: 1.08E+4nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 5.60E+3nM ΔG°: -30.2kJ/mole IC50: 6.80E+3nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, the concentrations of inhibitor that caused 50% inhibition of enzymati...More data for this Ligand-Target Pair
Affinity DataKi: 7.20E+3nM ΔG°: -29.5kJ/mole IC50: 1.39E+4nMpH: 7.2 T: 2°CAssay Description:For selected lead compounds from fluorescence-based high-throughput screening, Km and Vmax were calculated using the double-reciprocal Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30nMAssay Description:Antagonist activity at CXCR1 (unknown origin) transfected in RBL cells assessed as inhibition of IL-8-mediated intracellular calcium release preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Antagonist activity at human FPR2 transfected in human HL-60 cells assessed as inhibition of WKYMVM-induced intracellular calcium mobilization preinc...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Antagonist activity at CXCR2 (unknown origin) transfected in RBL cells assessed as inhibition of IL-8-mediated intracellular calcium release preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Antagonist activity at CXCR1 (unknown origin) transfected in RBL cells assessed as inhibition of IL-8-mediated intracellular calcium release preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Antagonist activity at human FPR2 transfected in human HL-60 cells assessed as inhibition of WKYMVM-induced intracellular calcium mobilization preinc...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Antagonist activity at human FPR2 transfected in human HL-60 cells assessed as inhibition of WKYMVM-induced intracellular calcium mobilization preinc...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Antagonist activity at human FPR2 transfected in human HL-60 cells assessed as inhibition of WKYMVM-induced intracellular calcium mobilization preinc...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:Antagonist activity at CXCR2 (unknown origin) transfected in RBL cells assessed as inhibition of IL-8-mediated intracellular calcium release preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:Inhibition of CXCR2 (unknown origin) transfected with RBL cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:Antagonist activity at CXCR2 (unknown origin) transfected in RBL cells assessed as inhibition of IL-8-mediated intracellular calcium release preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Antagonist activity at CXCR2 in human PMNs assessed as inhibition of CXCL1-induced intracellular Ca2+ release by fluorescence based calcium flux assa...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Antagonist activity at human FPR2 transfected in human HL-60 cells assessed as inhibition of WKYMVM-induced intracellular calcium mobilization preinc...More data for this Ligand-Target Pair
Affinity DataIC50: 31nMAssay Description:Inhibition of CXCR1 (unknown origin) transfected with RBL cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 31nMAssay Description:Antagonist activity at CXCR1 (unknown origin) transfected in RBL cells assessed as inhibition of IL-8-mediated intracellular calcium release preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 38nMAssay Description:Antagonist activity at CXCR2 in human PMNs assessed as inhibition of CXCL1-induced intracellular Ca2+ release by fluorescence based calcium flux assa...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Antagonist activity at human CXCR2 expressed in HEK293 cells assessed as inhibition of CXCL8-induced intracellular Ca2+ release by fluorescence based...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of recombinant Cdc25A (unknown origin) using OMFP as substrate measured every 30 sec for 10 mins by fluorometric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:Antagonist activity at human CXCR2 expressed in HEK293 cells assessed as inhibition of CVCL8-induced [35S]GTPgammaS binding after 60 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 78nMAssay Description:Antagonist activity at CXCR2 in human PMNs assessed as inhibition of CXCL1-induced intracellular Ca2+ release by fluorescence based calcium flux assa...More data for this Ligand-Target Pair
Affinity DataIC50: 81nMAssay Description:Antagonist activity at CXCR2 (unknown origin) transfected in RBL cells assessed as inhibition of IL-8-mediated intracellular calcium release preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 85nMAssay Description:Antagonist activity at human FPR2 transfected in human HL-60 cells assessed as inhibition of WKYMVM-induced intracellular calcium mobilization preinc...More data for this Ligand-Target Pair
Affinity DataIC50: 105nMAssay Description:Antagonist activity at CXCR2 in human PMNs assessed as inhibition of CXCL1-induced intracellular Ca2+ release by fluorescence based calcium flux assa...More data for this Ligand-Target Pair
Affinity DataIC50: 121nMAssay Description:Antagonist activity at CXCR2 (unknown origin) transfected in RBL cells assessed as inhibition of IL-8-mediated intracellular calcium release preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Inhibition of CXCR2 (unknown origin) expressed in PathHunter celline by beta-arrestin assayMore data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:Antagonist activity at human FPR2 transfected in human HL-60 cells assessed as inhibition of WKYMVM-induced intracellular calcium mobilization preinc...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Antagonist activity at CXCR2 in human PMNs assessed as inhibition of CXCL1-induced intracellular Ca2+ release by fluorescence based calcium flux assa...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMAssay Description:Antagonist activity at human CXCR2 expressed in HEK293 cells assessed as inhibition of CXCL8-induced intracellular Ca2+ release by fluorescence based...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMAssay Description:Antagonist activity at human FPR2 transfected in human HL-60 cells assessed as inhibition of WKYMVM-induced intracellular calcium mobilization preinc...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Inhibition of recombinant Cdc25A (unknown origin) using OMFP as substrate measured every 30 sec for 10 mins by fluorometric assayMore data for this Ligand-Target Pair