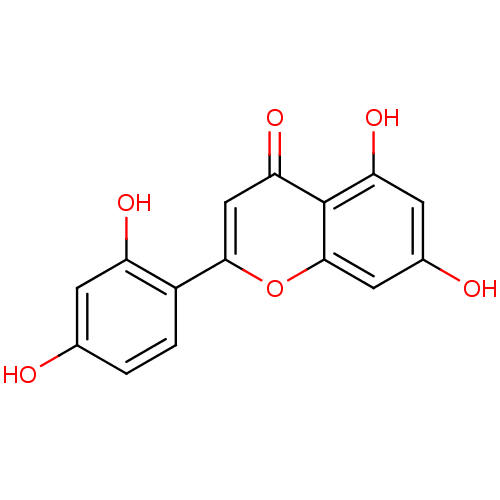
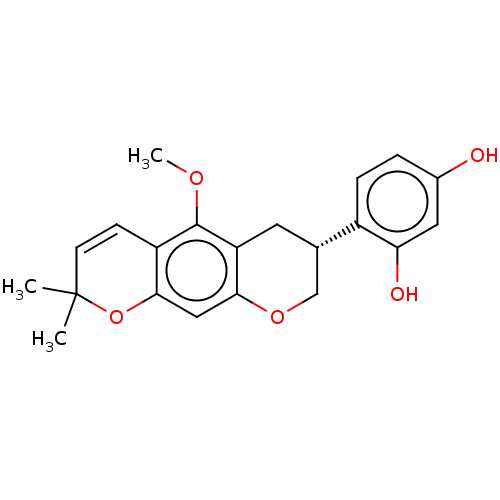
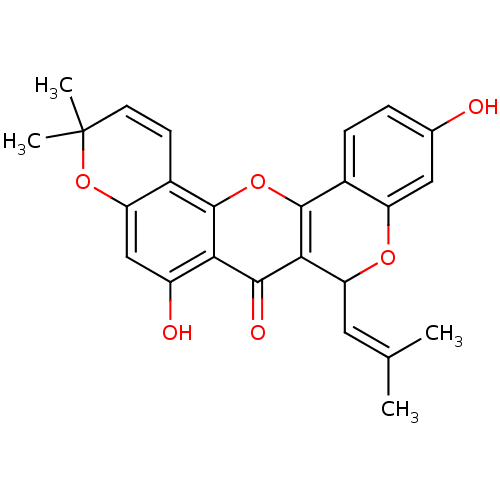
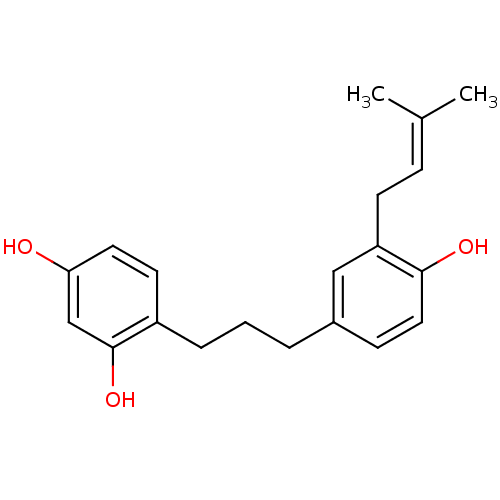
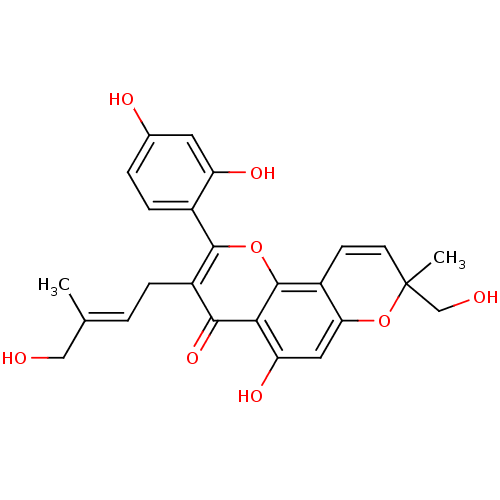
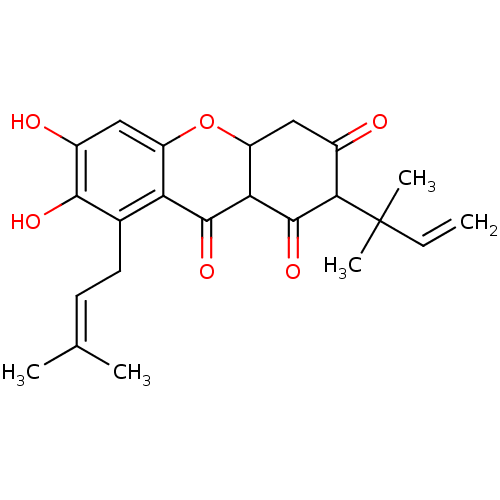
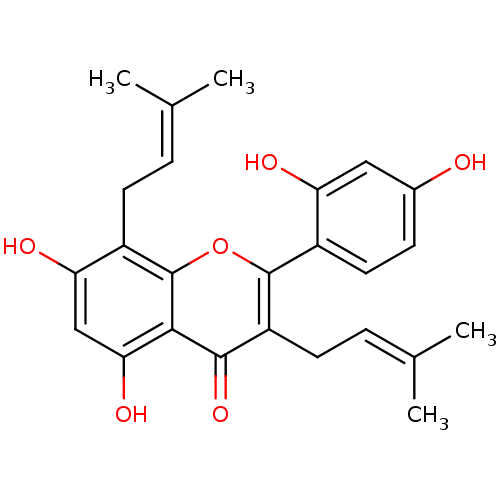
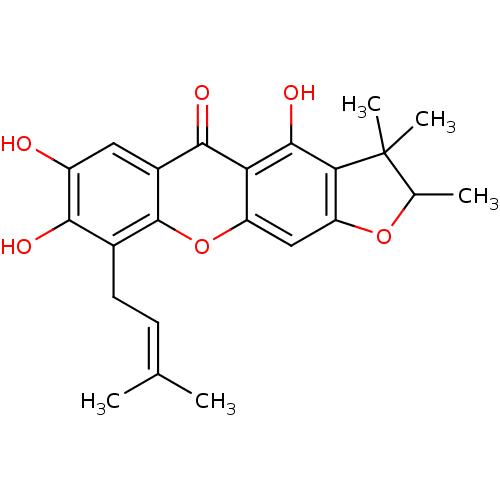
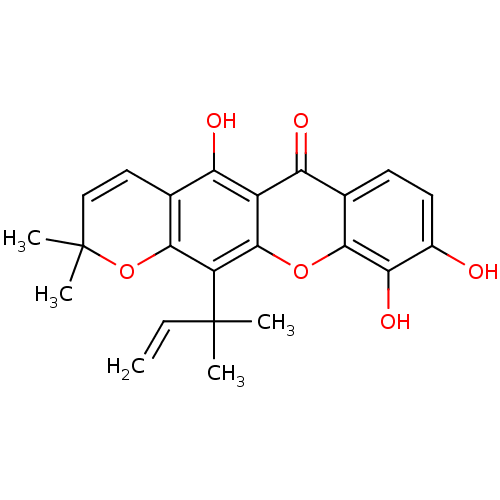
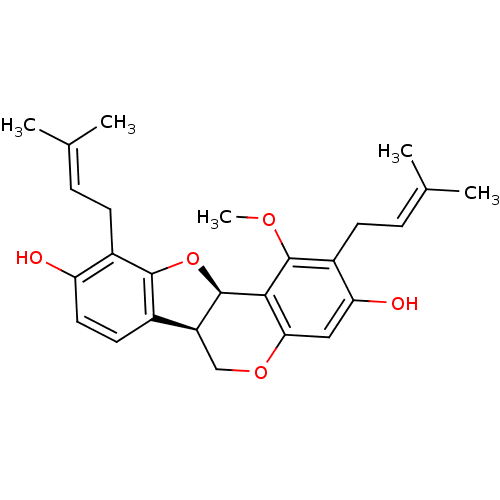
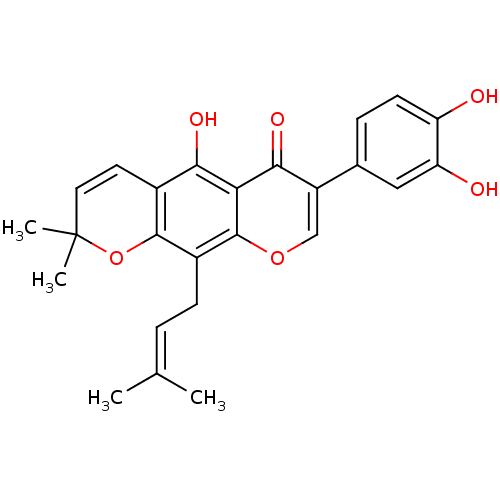
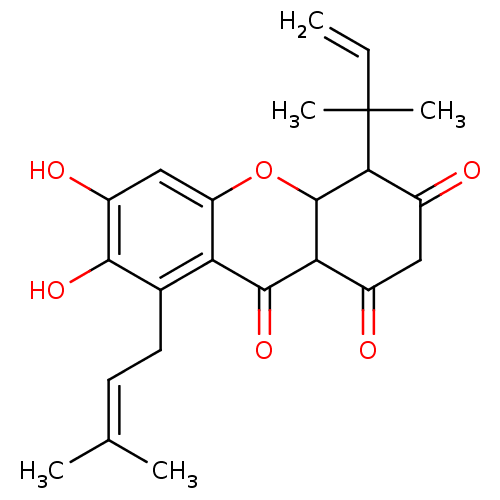
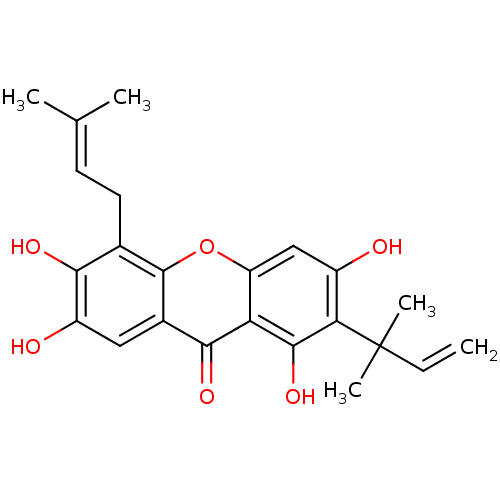
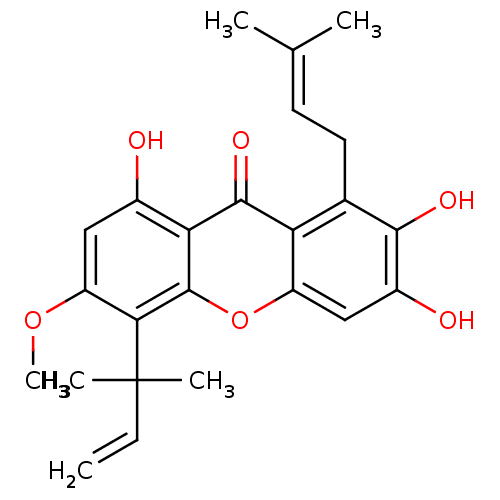
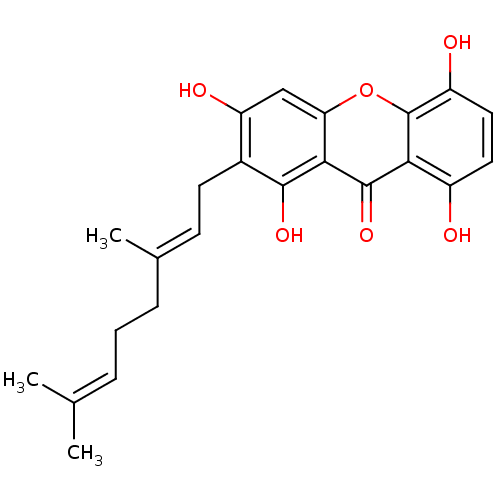
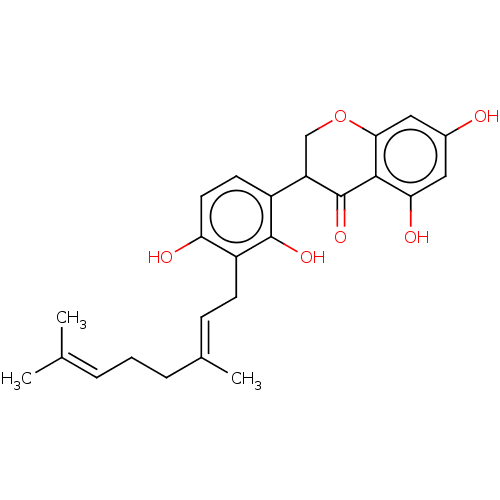
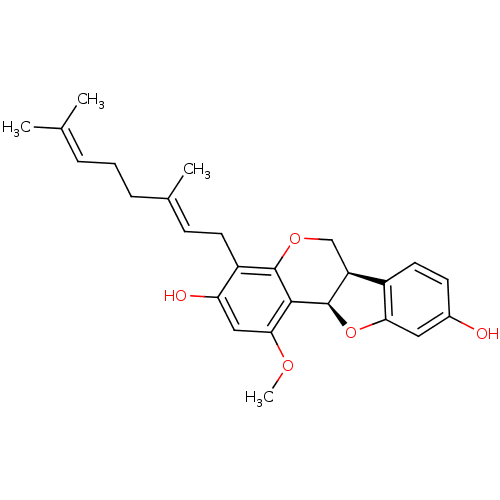
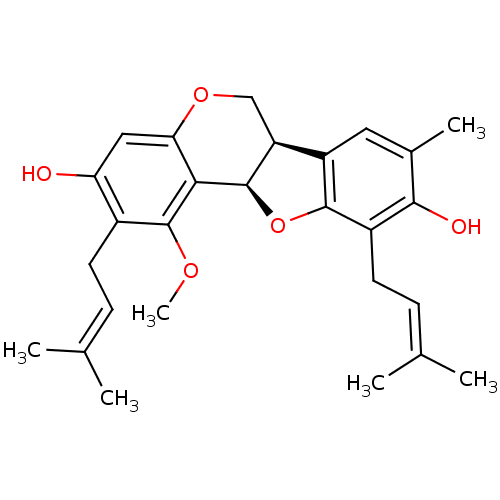
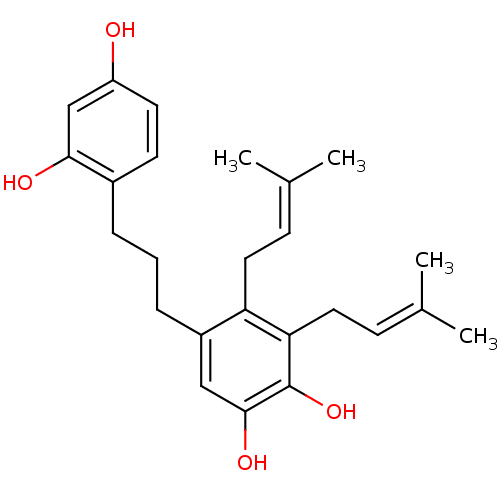
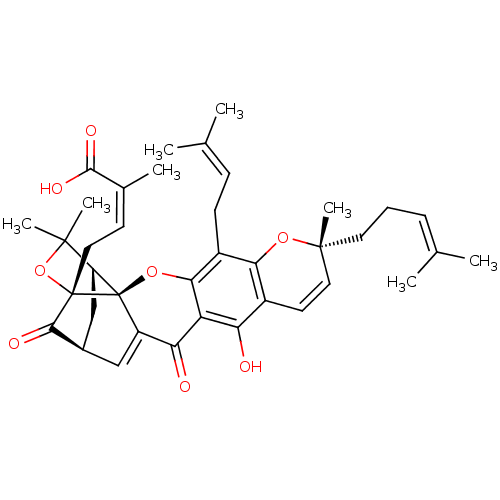
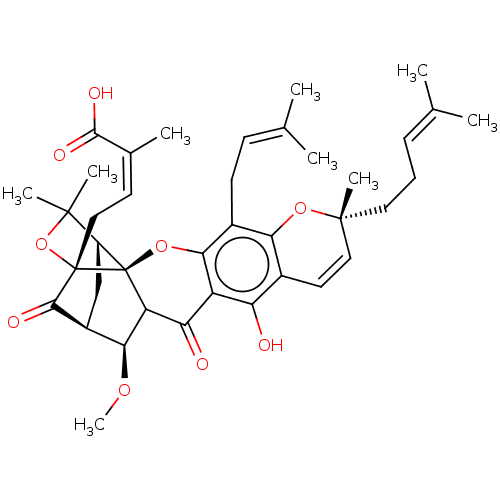
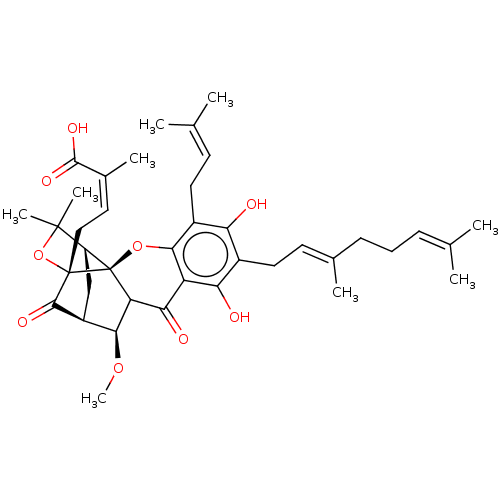
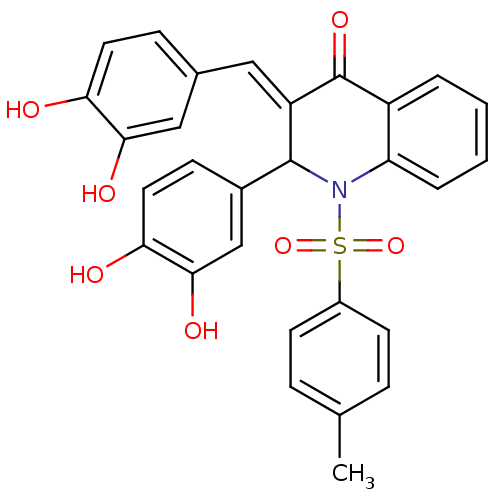
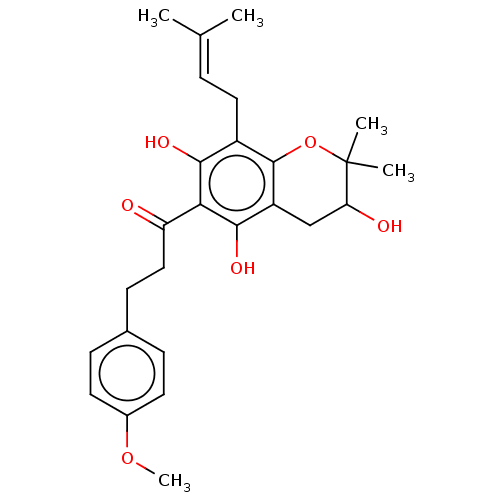
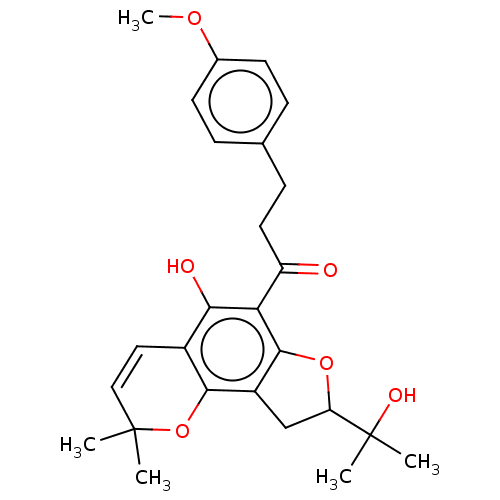
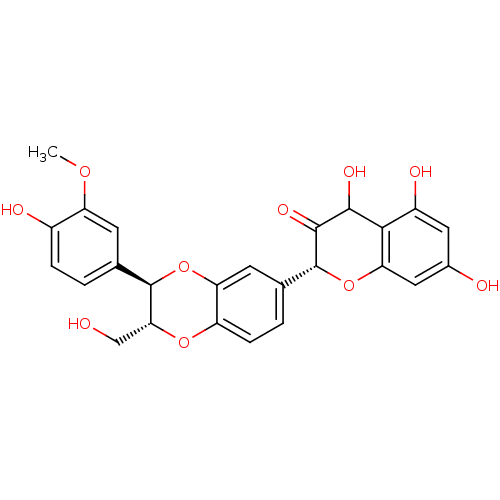
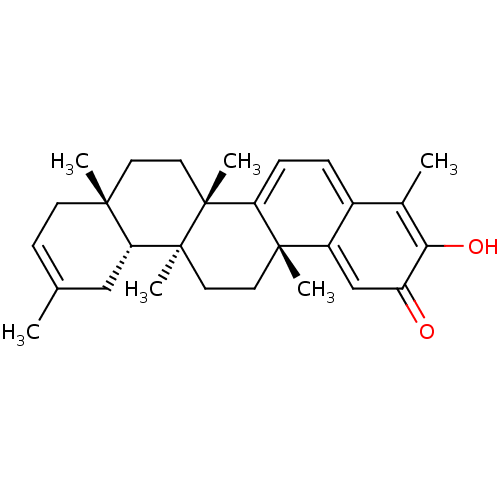
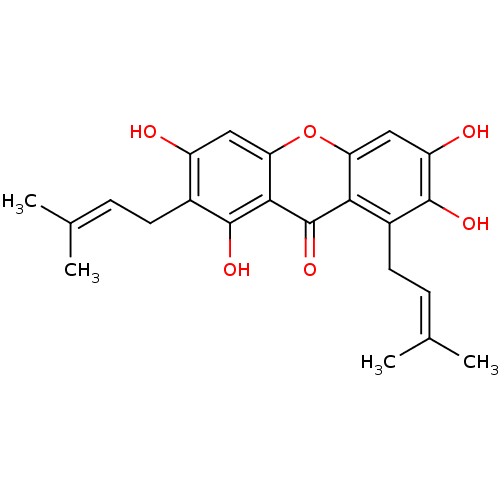
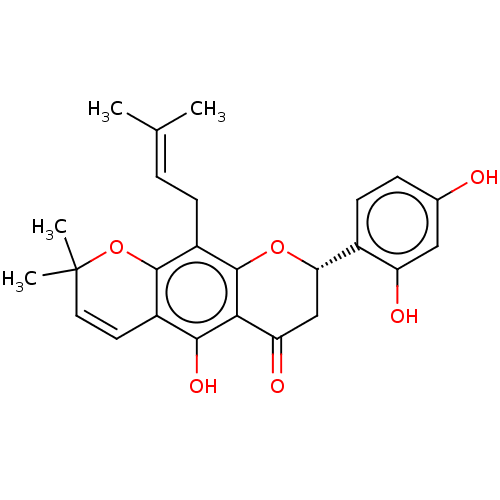
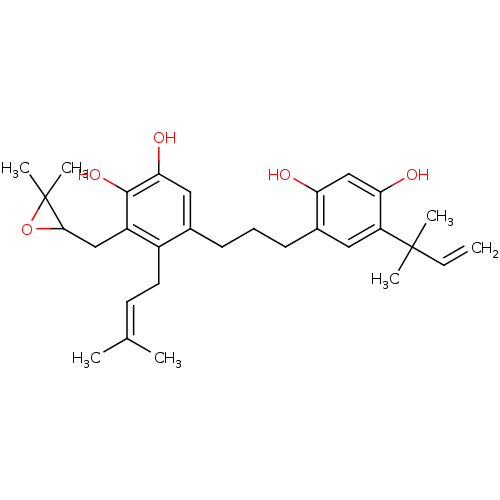
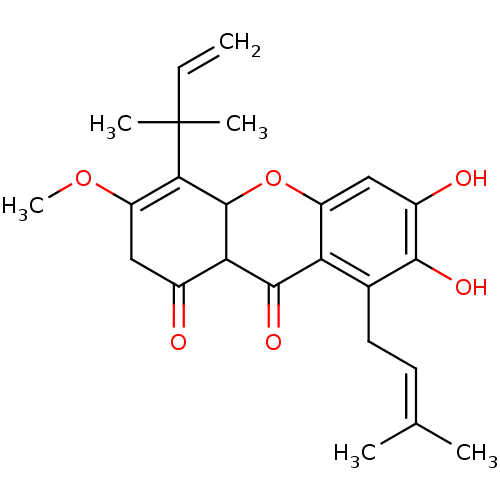
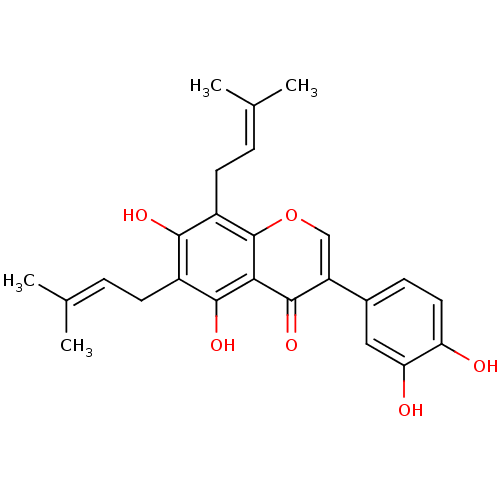
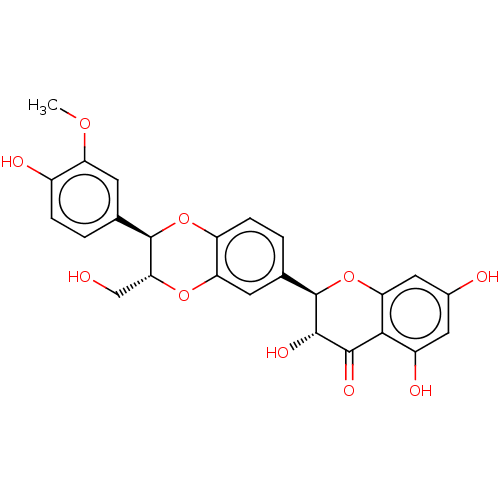
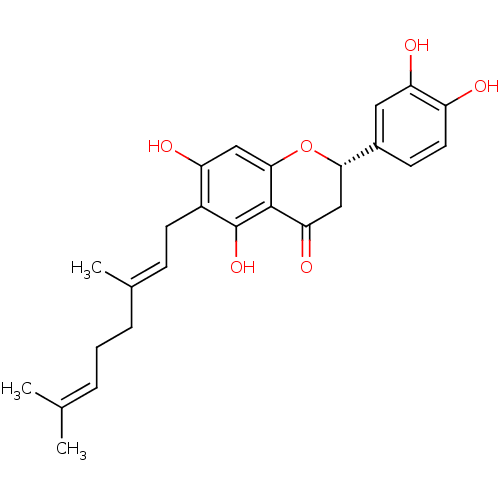
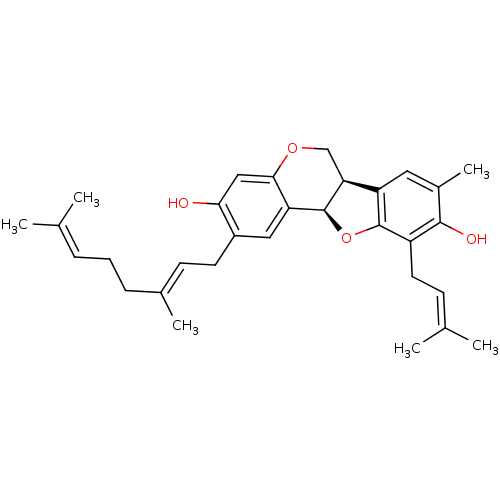
Affinity DataKi: 0.610nM ΔG°: -53.5kJ/moleT: 2°CAssay Description:Mushroom tyrosinase using either L-DOPA or L-tyrosine as substrate. In spectrophotometric experiments, enzyme activity was monitored by dopachrome f...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Time dependent inhibition of monophenolase activity of mushroom tyrosinase using L-tyrosine as substrate measured every 30 secs by Morrison and Walsh...More data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Competitive inhibition of monophenolase activity of mushroom tyrosinase using L-tyrosine as substrate preincubated for 5 to 60 mins by Lineweaver-Bur...More data for this Ligand-Target Pair
Affinity DataKi: 46.4nM ΔG°: -42.6kJ/moleT: 2°CAssay Description:Mushroom tyrosinase using either L-DOPA or L-tyrosine as substrate. In spectrophotometric experiments, enzyme activity was monitored by dopachrome f...More data for this Ligand-Target Pair
Affinity DataKi: 48.5nMAssay Description:Inhibition of mushroom tyrosinase by kinetic based assayMore data for this Ligand-Target Pair
Affinity DataKi: 49.2nM ΔG°: -42.4kJ/moleT: 2°CAssay Description:Mushroom tyrosinase using either L-DOPA or L-tyrosine as substrate. In spectrophotometric experiments, enzyme activity was monitored by dopachrome f...More data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 58nMAssay Description:Competitive inhibition of Clostridium perfringens neuraminidase by Lineweaver-Burke plot and Dixon plotMore data for this Ligand-Target Pair
Affinity DataKi: 80.5nM ΔG°: -41.2kJ/moleT: 2°CAssay Description:Mushroom tyrosinase using either L-DOPA or L-tyrosine as substrate. In spectrophotometric experiments, enzyme activity was monitored by dopachrome f...More data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 98nMAssay Description:Competitive inhibition of Clostridium perfringens neuraminidase by Lineweaver-Burke plot and Dixon plotMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 103nMAssay Description:Competitive inhibition of Clostridium perfringens neuraminidase by Lineweaver-Burke plot and Dixon plotMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 110nMAssay Description:Non-competitive inhibition of Clostridium perfringens neuraminidase using 4-methylumbelliferyl-alpha-D-N-acetylneuraminic acid as substrate by Linewe...More data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 127nMAssay Description:Apparent binding affinity at Clostridium perfringens neuraminidase by fluorimetryMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 130nMAssay Description:Non-competitive inhibition of Clostridium perfringens neuraminidase using 4-methylumbelliferyl-alpha-D-N-acetylneuraminic acid sodium salt hydrate as...More data for this Ligand-Target Pair
Affinity DataKi: 130nMAssay Description:Competitive inhibition of diphenolase activity of mushroom tyrosinase using L-DOPA as substrate preincubated for 5 to 60 mins by Lineweaver-Burk and ...More data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 136nMAssay Description:Competitive inhibition of Clostridium perfringens neuraminidase by Lineweaver-Burke plot and Dixon plotMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 138nMAssay Description:Competitive inhibition of Clostridium perfringens neuraminidase by Lineweaver-Burke plot and Dixon plotMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 143nMAssay Description:Competitive inhibition of Clostridium perfringens neuraminidase by Lineweaver-Burke plot and Dixon plotMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 150nMAssay Description:Competitive inhibition of Clostridium perfringens neuraminidase by fluorometryMore data for this Ligand-Target Pair
Affinity DataKi: 151nMAssay Description:Time dependent inhibition of monophenolase activity of mushroom tyrosinase using L-tyrosine as substrate measured every 30 secs by Morrison and Walsh...More data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 160nMAssay Description:Non-competitive inhibition of Clostridium perfringens neuraminidase using 4-methylumbelliferyl-alpha-D-N-acetylneuraminic acid as substrate by Linewe...More data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 160nMAssay Description:Non-competitive inhibition of Clostridium perfringens neuraminidase using 4-methylumbelliferyl-alpha-D-N-acetylneuraminic acid as substrate by Linewe...More data for this Ligand-Target Pair
Affinity DataKi: 181nMAssay Description:Inhibition of mushroom tyrosinase by kinetic based assayMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 1(Homo sapiens (Human))
Gyeongsang National University
Curated by ChEMBL
Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 190nMAssay Description:Inhibition of recombinant human PTP1B (1 to 322 residues) expressed in Escherichia coli using pNPP as substrate by Dixon plot analysisMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 214nMAssay Description:Apparent binding affinity at Clostridium perfringens neuraminidase by fluorimetryMore data for this Ligand-Target Pair
Affinity DataKi: 230nMAssay Description:Inhibition of monophenolase activity of mushroom tyrosinase using as L-tyrosine substrate by Lineweaver-Burk plot based kinetic assayMore data for this Ligand-Target Pair
Affinity DataKi: 290nMAssay Description:Inhibition of diphenolase activity of mushroom tyrosinase using as L-DOPA substrate by Lineweaver-Burk plot based kinetic assayMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 1(Homo sapiens (Human))
Gyeongsang National University
Curated by ChEMBL
Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 300nMAssay Description:Inhibition of recombinant human PTP1B (1 to 322 residues) expressed in Escherichia coli using pNPP as substrate by Dixon plot analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 1(Homo sapiens (Human))
Gyeongsang National University
Curated by ChEMBL
Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 390nMAssay Description:Inhibition of recombinant human PTP1B (1 to 322 residues) expressed in Escherichia coli using pNPP as substrate by Dixon plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 400nM ΔG°: -37.7kJ/mole IC50: 600nMpH: 7.6 T: 2°CAssay Description:Inhibition assay using trans-sialidase, a membrane-associated protein.More data for this Ligand-Target Pair
Affinity DataKi: 410nMAssay Description:Inhibition of monophenolase activity of mushroom tyrosinase using as L-tyrosine substrate by Lineweaver-Burk plot based kinetic assayMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 436nMAssay Description:Apparent binding affinity at Clostridium perfringens neuraminidase by fluorimetryMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 477nMAssay Description:Apparent binding affinity at Clostridium perfringens neuraminidase by fluorimetryMore data for this Ligand-Target Pair
Affinity DataKi: 480nMAssay Description:Competitive inhibition of mushroom tyrosinase using L-tyrosine as substrate preincubated for 10 mins with substrate followed by enzyme addition measu...More data for this Ligand-Target Pair
TargetTyrosine-protein phosphatase non-receptor type 1(Homo sapiens (Human))
Gyeongsang National University
Curated by ChEMBL
Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 499nMAssay Description:Inhibition of recombinant human PTP1B (1 to 322 residues) expressed in Escherichia coli using pNPP as substrate incubated for 10 mins measured for 30...More data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 510nMAssay Description:Apparent binding affinity at Clostridium perfringens neuraminidase by fluorimetryMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 568nMAssay Description:Apparent binding affinity at Clostridium perfringens neuraminidase by fluorimetryMore data for this Ligand-Target Pair
Affinity DataKi: 630nMAssay Description:Competitive inhibition of mushroom tyrosinase using L-tyrosine as substrate preincubated for 10 mins with substrate followed by enzyme addition measu...More data for this Ligand-Target Pair
Affinity DataKi: 700nMAssay Description:Mixed type inhibition of monophenolase activity of mushroom tyrosinase assessed as reduction in dopachrome formation using L-Tyrosine substrate by UV...More data for this Ligand-Target Pair
Affinity DataKi: 700nMAssay Description:Inhibition of monophenolase activity of mushroom tyrosinase assessed as inhibition constant for free enzyme-inhibitor complex by measuring reduction ...More data for this Ligand-Target Pair
Affinity DataKi: 770nMAssay Description:Inhibition of diphenolase activity of mushroom tyrosinase using as L-DOPA substrate by Lineweaver-Burk plot based kinetic assayMore data for this Ligand-Target Pair
TargetReplicase polyprotein 1ab(Human SARS coronavirus (SARS-CoV) (Severe acute re...)
Korea Research Institute Of Bioscience And Biotechnology
Curated by ChEMBL
Korea Research Institute Of Bioscience And Biotechnology
Curated by ChEMBL
Affinity DataKi: 800nMAssay Description:Inhibition of 3C-like protease of SARS coronavirus assessed as concentration of FRET peptide for 60 mins by dixon plotMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 800nMAssay Description:Competitive inhibition of Clostridium perfringens neuraminidase by fluorometryMore data for this Ligand-Target Pair
Affinity DataKi: 860nMAssay Description:Competitive inhibition of mushroom tyrosinase using L-tyrosine as substrate preincubated for 10 mins with substrate followed by enzyme addition measu...More data for this Ligand-Target Pair
Affinity DataKi: 870nMAssay Description:Inhibition of mushroom tyrosinase by kinetic based assayMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 950nMAssay Description:Competitive inhibition of Clostridium perfringens neuraminidase by Lineweaver-Burke plot and Dixon plotMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 1.02E+3nMAssay Description:Non-competitive inhibition of Clostridium perfringens neuraminidase using 4-methylumbelliferyl-alpha-D-N-acetylneuraminic acid sodium salt hydrate as...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nMAssay Description:Inhibition of monophenolase activity of mushroom tyrosinase assessed as inhibition constant for free enzyme-inhibitor complex by measuring reduction ...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nMAssay Description:Mixed type inhibition of monophenolase activity of mushroom tyrosinase assessed as reduction in dopachrome formation using L-Tyrosine substrate by UV...More data for this Ligand-Target Pair
TargetCarboxylic ester hydrolase(Equus caballus (Horse))
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 1.20E+3nMAssay Description:Mixed type inhibition of equine BChE using butyrylthiocholine iodide as substrate by Lineweaver-Burk double-reciprocal-plot and dixon plot analysisMore data for this Ligand-Target Pair
TargetSialidase(Clostridium perfringens)
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Graduate School Of Gyeongsang National University
Curated by ChEMBL
Affinity DataKi: 1.20E+3nMAssay Description:Non-competitive inhibition of Clostridium perfringens neuraminidase using 4-methylumbelliferyl-alpha-D-N-acetylneuraminic acid as substrate by Linewe...More data for this Ligand-Target Pair