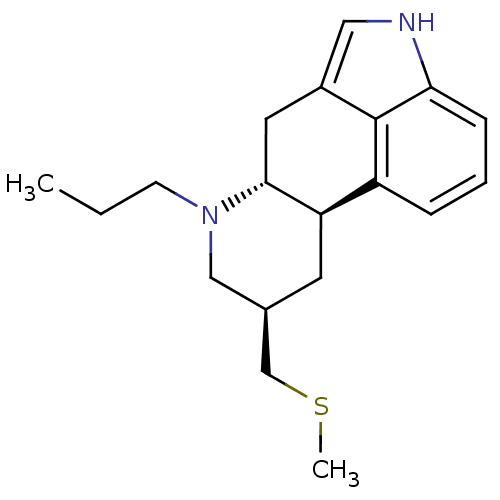
Affinity DataKi: 1nMAssay Description:Compound was tested for antagonistic activity against D2 receptor from rat striatum, used [3H]raclopride as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Binding affinity towards Dopamine receptor D2 by displacement of [3H]-U-86,170.More data for this Ligand-Target Pair
Affinity DataKi: 26nMAssay Description:Displacement of [125]iodosulpride from human recombinant dopamine D2L receptor expressed in CHO cells after 30 minsMore data for this Ligand-Target Pair
Affinity DataKi: 50nMAssay Description:Affinity towards Dopamine receptor D2More data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Inhibitory activity against Dopamine receptor D2 in rat corpus striatum using [3H]-Apomorphine as radioligandMore data for this Ligand-Target Pair
Affinity DataIC50: 28nMAssay Description:Displacement of [3H]nemonapride from Rattus norvegicus (rat) chimeric dopamine D2 trunk/D2 tail receptor transfected in african green monkey COS7 cel...More data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Displacement of [3H]nemonapride from Rattus norvegicus (rat) wild type dopamine D2 receptor transfected in african green monkey COS7 cells after 1 hr...More data for this Ligand-Target Pair
Affinity DataIC50: 48nMAssay Description:Inhibitory activity against Dopamine receptor D2 in calf corpus striatum using [3H]-Spiperone as radioligandMore data for this Ligand-Target Pair
Affinity DataKd: 45nMAssay Description:In vitro affinity at mutant D2 receptor (S194A) in C6 (glioma) cell membranes.More data for this Ligand-Target Pair
Affinity DataKd: 4nMAssay Description:In vitro affinity at mutant D2 receptor (S194A) in C6 (glioma) cell membranes.More data for this Ligand-Target Pair
Affinity DataEC50: 7.90nMAssay Description:Agonist activity at Rattus norvegicus (rat) dopamine D2 receptor transfected in african green monkey COS7 cells assessed as inhibition of forskolin-s...More data for this Ligand-Target Pair
Affinity DataKd: 40nMAssay Description:In vitro affinity at mutant D2 receptor (S197A) in C6 (glioma) cell membranes.More data for this Ligand-Target Pair
Affinity DataKd: 30nMAssay Description:In vitro affinity at wild type Dopamine receptor D2 on C6 (glioma) cell membranes.More data for this Ligand-Target Pair