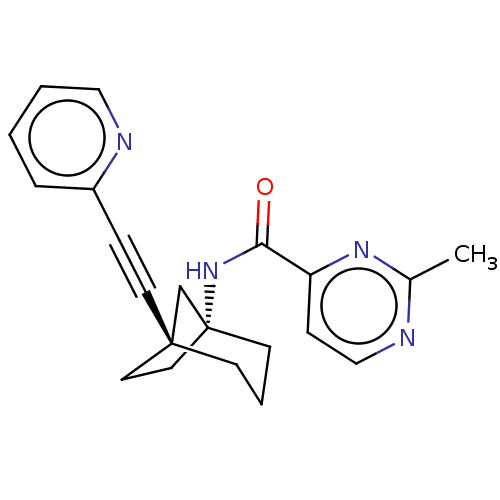
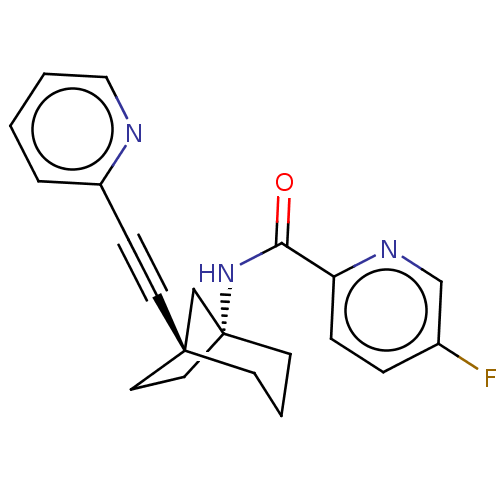
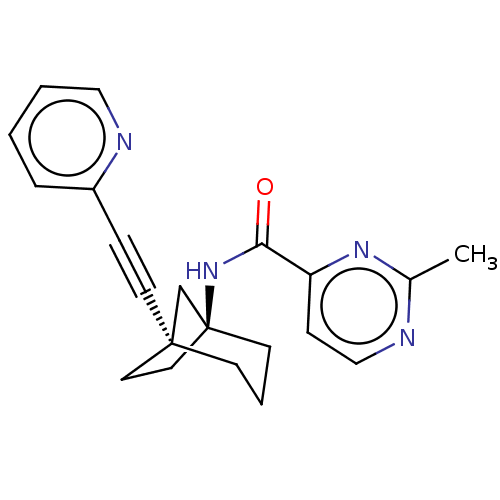
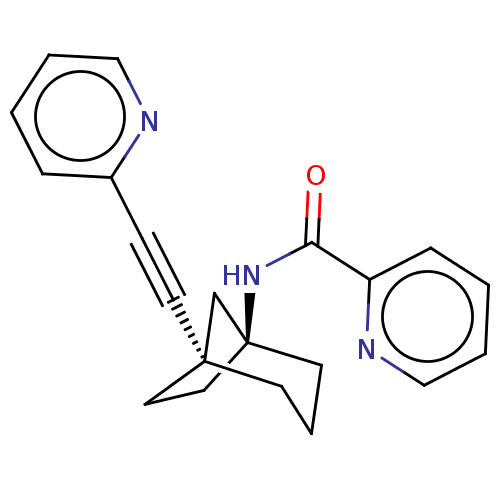
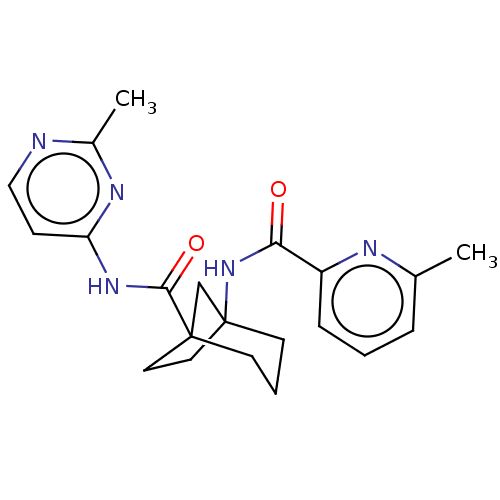
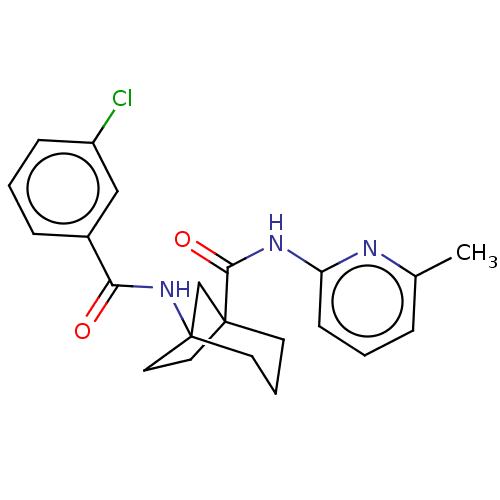
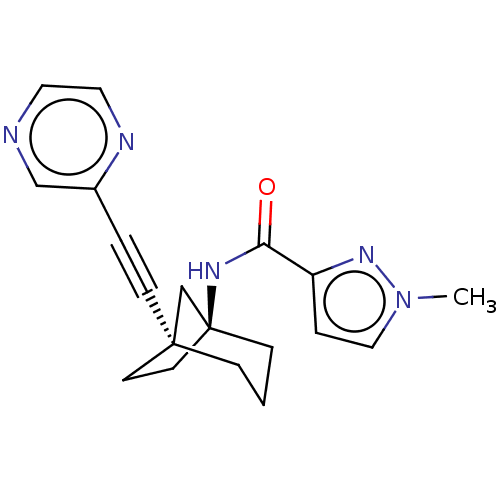
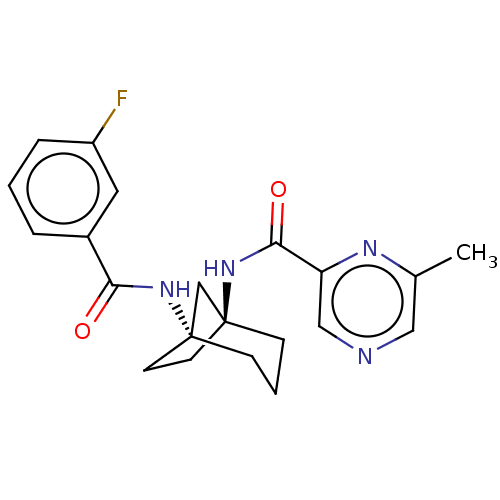
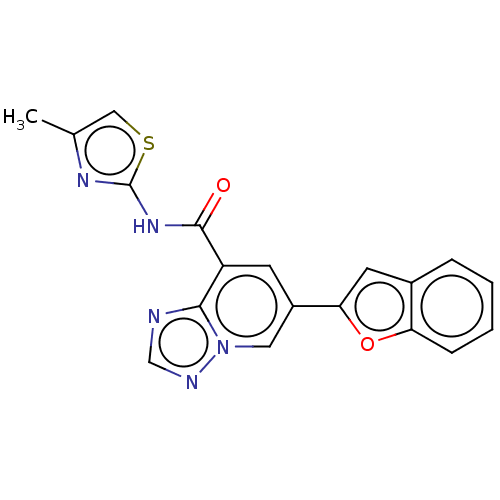
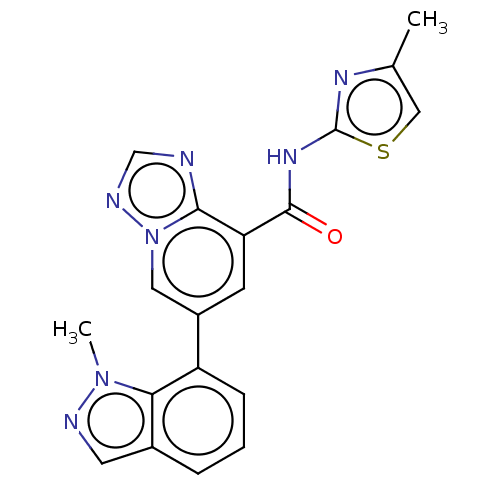
Affinity DataKi: 3nM ΔG°: -48.6kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 4.5nM ΔG°: -47.6kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 4.70nM ΔG°: -47.5kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 6.20nM ΔG°: -46.8kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -45.7kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 11nM ΔG°: -45.4kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 17nM ΔG°: -44.3kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 22nM ΔG°: -43.7kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 27nM ΔG°: -43.2kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol. Pharmacol., 2003, 64, 731-740] with slight modifications, including that a r...More data for this Ligand-Target Pair
Affinity DataKi: 34nM ΔG°: -42.6kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 35nM ΔG°: -42.6kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol. Pharmacol., 2003, 64, 731-740] with slight modifications, including that a r...More data for this Ligand-Target Pair
Affinity DataKi: 35nM ΔG°: -42.6kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 38nM ΔG°: -42.4kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 39nM ΔG°: -42.3kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol. Pharmacol., 2003, 64, 731-740] with slight modifications, including that a r...More data for this Ligand-Target Pair
Affinity DataKi: 65nM ΔG°: -41.0kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol. Pharmacol., 2003, 64, 731-740] with slight modifications, including that a r...More data for this Ligand-Target Pair
Affinity DataKi: 78nM ΔG°: -40.6kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol. Pharmacol., 2003, 64, 731-740] with slight modifications, including that a r...More data for this Ligand-Target Pair
Affinity DataKi: 91nM ΔG°: -40.2kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 120nM ΔG°: -39.5kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 280nM ΔG°: -37.4kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol. Pharmacol., 2003, 64, 731-740] with slight modifications, including that a r...More data for this Ligand-Target Pair
Affinity DataKi: 340nM ΔG°: -36.9kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol. Pharmacol., 2003, 64, 731-740] with slight modifications, including that a r...More data for this Ligand-Target Pair
Affinity DataKi: 370nM ΔG°: -36.7kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol. Pharmacol., 2003, 64, 731-740] with slight modifications, including that a r...More data for this Ligand-Target Pair
Affinity DataKi: 500nM ΔG°: -36.0kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol. Pharmacol., 2003, 64, 731-740] with slight modifications, including that a r...More data for this Ligand-Target Pair
Affinity DataKi: 590nM ΔG°: -35.6kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nM ΔG°: -33.8kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol. Pharmacol., 2003, 64, 731-740] with slight modifications, including that a r...More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+3nM ΔG°: -33.6kJ/molepH: 7.4 T: 2°CAssay Description:Binding assays were performed as described in [J. A. O'Brien et al. Mol Pharmacol., 2003, 64, 731-740] with slight modifications, including that ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.320nMpH: 7.4 T: 2°CAssay Description:The cDNA for rat metabotropic glutamate receptor 5 (rmGluR5) and the cDNA for human metabotropic glutamate receptor 5 (rmGluR5) were generous gifts f...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMpH: 7.4Assay Description:The rmGluR5 or hmGluR5 was stably expressed in a HEK 293 cell line and gown in Dulbecco's Modified Eagle Medium (DMEM) (Invitrogen, Carlsbad, Cal...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMpH: 7.4Assay Description:The rmGluR5 or hmGluR5 was stably expressed in a HEK 293 cell line and gown in Dulbecco's Modified Eagle Medium (DMEM) (Invitrogen, Carlsbad, Cal...More data for this Ligand-Target Pair
Affinity DataIC50: 3.70nMpH: 7.4 T: 2°CAssay Description:The cDNA for rat metabotropic glutamate receptor 5 (rmGluR5) and the cDNA for human metabotropic glutamate receptor 5 (rmGluR5) were generous gifts f...More data for this Ligand-Target Pair
Affinity DataIC50: 3.80nMpH: 7.4Assay Description:The rmGluR5 or hmGluR5 was stably expressed in a HEK 293 cell line and gown in Dulbecco's Modified Eagle Medium (DMEM) (Invitrogen, Carlsbad, Cal...More data for this Ligand-Target Pair
Affinity DataIC50: 3.90nMpH: 7.4 T: 2°CAssay Description:The cDNA for rat metabotropic glutamate receptor 5 (rmGluR5) and the cDNA for human metabotropic glutamate receptor 5 (rmGluR5) were generous gifts f...More data for this Ligand-Target Pair
Affinity DataIC50: 4.70nMpH: 7.4 T: 2°CAssay Description:The cDNA for rat metabotropic glutamate receptor 5 (rmGluR5) and the cDNA for human metabotropic glutamate receptor 5 (rmGluR5) were generous gifts f...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30nMpH: 7.4Assay Description:The rmGluR5 or hmGluR5 was stably expressed in a HEK 293 cell line and gown in Dulbecco's Modified Eagle Medium (DMEM) (Invitrogen, Carlsbad, Cal...More data for this Ligand-Target Pair
Affinity DataIC50: 5.90nMpH: 7.4Assay Description:The rmGluR5 or hmGluR5 was stably expressed in a HEK 293 cell line and gown in Dulbecco's Modified Eagle Medium (DMEM) (Invitrogen, Carlsbad, Cal...More data for this Ligand-Target Pair
Affinity DataIC50: 6.60nMpH: 7.4Assay Description:The rmGluR5 or hmGluR5 was stably expressed in a HEK 293 cell line and gown in Dulbecco's Modified Eagle Medium (DMEM) (Invitrogen, Carlsbad, Cal...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMpH: 7.4 T: 2°CAssay Description:The cDNA for rat metabotropic glutamate receptor 5 (rmGluR5) and the cDNA for human metabotropic glutamate receptor 5 (rmGluR5) were generous gifts f...More data for this Ligand-Target Pair
Affinity DataIC50: 9.30nMpH: 7.4Assay Description:The rmGluR5 or hmGluR5 was stably expressed in a HEK 293 cell line and gown in Dulbecco's Modified Eagle Medium (DMEM) (Invitrogen, Carlsbad, Cal...More data for this Ligand-Target Pair
Affinity DataIC50: >10nMpH: 7.4 T: 2°CAssay Description:The following examples of the in vitro effects of the disclosed compounds are prophetic. An example of an in vitro assay method for assessing the rec...More data for this Ligand-Target Pair
Affinity DataIC50: >10nMpH: 7.4 T: 2°CAssay Description:The following examples of the in vitro effects of the disclosed compounds are prophetic. An example of an in vitro assay method for assessing the rec...More data for this Ligand-Target Pair
Affinity DataIC50: >10nMpH: 7.4 T: 2°CAssay Description:The following examples of the in vitro effects of the disclosed compounds are prophetic. An example of an in vitro assay method for assessing the rec...More data for this Ligand-Target Pair
Affinity DataIC50: >10nMpH: 7.4 T: 2°CAssay Description:The following examples of the in vitro effects of the disclosed compounds are prophetic. An example of an in vitro assay method for assessing the rec...More data for this Ligand-Target Pair
Affinity DataIC50: >10nMpH: 7.4 T: 2°CAssay Description:The following examples of the in vitro effects of the disclosed compounds are prophetic. An example of an in vitro assay method for assessing the rec...More data for this Ligand-Target Pair
Affinity DataIC50: >10nMpH: 7.4 T: 2°CAssay Description:The following examples of the in vitro effects of the disclosed compounds are prophetic. An example of an in vitro assay method for assessing the rec...More data for this Ligand-Target Pair
Affinity DataIC50: >10nMpH: 7.4 T: 2°CAssay Description:The following examples of the in vitro effects of the disclosed compounds are prophetic. An example of an in vitro assay method for assessing the rec...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMpH: 7.4 T: 2°CAssay Description:The following examples of the in vitro effects of the disclosed compounds are prophetic. An example of an in vitro assay method for assessing the rec...More data for this Ligand-Target Pair
Affinity DataIC50: >10nMpH: 7.4 T: 2°CAssay Description:The following examples of the in vitro effects of the disclosed compounds are prophetic. An example of an in vitro assay method for assessing the rec...More data for this Ligand-Target Pair
Affinity DataIC50: >10nMpH: 7.4 T: 2°CAssay Description:The following examples of the in vitro effects of the disclosed compounds are prophetic. An example of an in vitro assay method for assessing the rec...More data for this Ligand-Target Pair
Affinity DataIC50: >10nMpH: 7.4 T: 2°CAssay Description:The following examples of the in vitro effects of the disclosed compounds are prophetic. An example of an in vitro assay method for assessing the rec...More data for this Ligand-Target Pair
Affinity DataIC50: >10nMpH: 7.4 T: 2°CAssay Description:The following examples of the in vitro effects of the disclosed compounds are prophetic. An example of an in vitro assay method for assessing the rec...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMpH: 7.4 T: 2°CAssay Description:The following examples of the in vitro effects of the disclosed compounds are prophetic. An example of an in vitro assay method for assessing the rec...More data for this Ligand-Target Pair