Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Glutamate receptor ionotropic, kainate 2
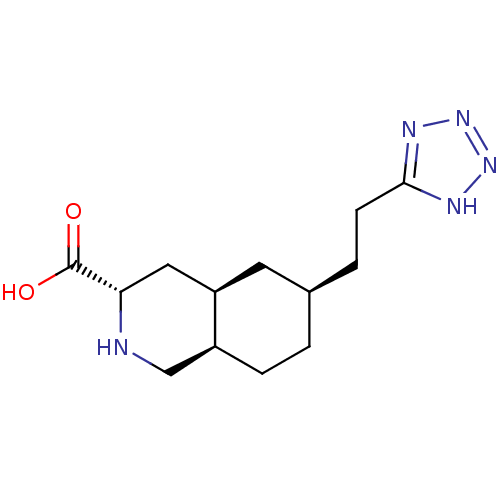
Ligand
BDBM86751
Substrate
n/a
Meas. Tech.
ChEMBL_90593 (CHEMBL701338)
Ki
>100000±n/a nM
Citation
 Filla, SA; Winter, MA; Johnson, KW; Bleakman, D; Bell, MG; Bleisch, TJ; Castaño, AM; Clemens-Smith, A; del Prado, M; Dieckman, DK; Dominguez, E; Escribano, A; Ho, KH; Hudziak, KJ; Katofiasc, MA; Martinez-Perez, JA; Mateo, A; Mathes, BM; Mattiuz, EL; Ogden, AM; Phebus, LA; Stack, DR; Stratford, RE; Ornstein, PL Ethyl (3S,4aR,6S,8aR)-6-(4-ethoxycar- bonylimidazol-1-ylmethyl)decahydroiso-quinoline-3-carboxylic ester: a prodrug of a GluR5 kainate receptor antagonist active in two animal models of acute migraine. J Med Chem 45:4383-6 (2002) [PubMed] Article
Filla, SA; Winter, MA; Johnson, KW; Bleakman, D; Bell, MG; Bleisch, TJ; Castaño, AM; Clemens-Smith, A; del Prado, M; Dieckman, DK; Dominguez, E; Escribano, A; Ho, KH; Hudziak, KJ; Katofiasc, MA; Martinez-Perez, JA; Mateo, A; Mathes, BM; Mattiuz, EL; Ogden, AM; Phebus, LA; Stack, DR; Stratford, RE; Ornstein, PL Ethyl (3S,4aR,6S,8aR)-6-(4-ethoxycar- bonylimidazol-1-ylmethyl)decahydroiso-quinoline-3-carboxylic ester: a prodrug of a GluR5 kainate receptor antagonist active in two animal models of acute migraine. J Med Chem 45:4383-6 (2002) [PubMed] Article More Info.:
Target
Name:
Glutamate receptor ionotropic, kainate 2
Synonyms:
EAA4 | Excitatory amino acid receptor 4 | GLUR6 | GRIK2 | GRIK2_HUMAN | GluK2 | GluR-6 | Glutamate kainate | Glutamate receptor 6 | Glutamate receptor ionotropic kainate 2 | Glutamate receptor, ionotropic kainate 2 | Glutamate-Kainate | Glutamate-Kainate, GluR6
Type:
Enzyme Catalytic Domain
Mol. Mass.:
102592.78
Organism:
Homo sapiens (Human)
Description:
Q13002
Residue:
908
Sequence:
MKIIFPILSNPVFRRTVKLLLCLLWIGYSQGTTHVLRFGGIFEYVESGPMGAEELAFRFAVNTINRNRTLLPNTTLTYDTQKINLYDSFEASKKACDQLSLGVAAIFGPSHSSSANAVQSICNALGVPHIQTRWKHQVSDNKDSFYVSLYPDFSSLSRAILDLVQFFKWKTVTVVYDDSTGLIRLQELIKAPSRYNLRLKIRQLPADTKDAKPLLKEMKRGKEFHVIFDCSHEMAAGILKQALAMGMMTEYYHYIFTTLDLFALDVEPYRYSGVNMTGFRILNTENTQVSSIIEKWSMERLQAPPKPDSGLLDGFMTTDAALMYDAVHVVSVAVQQFPQMTVSSLQCNRHKPWRFGTRFMSLIKEAHWEGLTGRITFNKTNGLRTDFDLDVISLKEEGLEKIGTWDPASGLNMTESQKGKPANITDSLSNRSLIVTTILEEPYVLFKKSDKPLYGNDRFEGYCIDLLRELSTILGFTYEIRLVEDGKYGAQDDANGQWNGMVRELIDHKADLAVAPLAITYVREKVIDFSKPFMTLGISILYRKPNGTNPGVFSFLNPLSPDIWMYILLAYLGVSCVLFVIARFSPYEWYNPHPCNPDSDVVENNFTLLNSFWFGVGALMQQGSELMPKALSTRIVGGIWWFFTLIIISSYTANLAAFLTVERMESPIDSADDLAKQTKIEYGAVEDGATMTFFKKSKISTYDKMWAFMSSRRQSVLVKSNEEGIQRVLTSDYAFLMESTTIEFVTQRNCNLTQIGGLIDSKGYGVGTPMGSPYRDKITIAILQLQEEGKLHMMKEKWWRGNGCPEEESKEASALGVQNIGGIFIVLAAGLVLSVFVAVGEFLYKSKKNAQLEKRSFCSAMVEELRMSLKCQRRLKHKPQAPVIVKTEEVINMHTFNDRRLPGKETMA