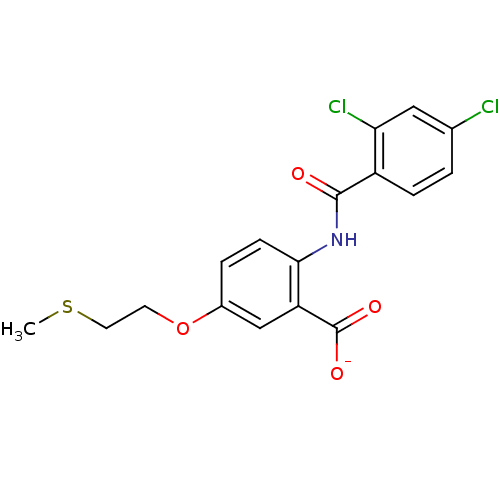
BDBM50121417 CHEMBL119646::Lithium; 2-(2,4-dichloro-benzoylamino)-5-(2-methylsulfanyl-ethoxy)-benzoate
SMILES CSCCOc1ccc(NC(=O)c2ccc(Cl)cc2Cl)c(c1)C([O-])=O
InChI Key InChIKey=NUFOUZIOJRXDFK-UHFFFAOYSA-M
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 4 hits for monomerid = 50121417
Found 4 hits for monomerid = 50121417
TargetPeroxisome proliferator-activated receptor gamma(Homo sapiens (Human))
Biovitrum
Curated by ChEMBL
Biovitrum
Curated by ChEMBL
Affinity DataKi: 1.60E+3nMAssay Description:Ligand binding affinity was determined by displacement of a tritiated tracer by the unlabeled compound to a GST-PPAR fusion protein for Peroxisome pr...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor alpha(Homo sapiens (Human))
Biovitrum
Curated by ChEMBL
Biovitrum
Curated by ChEMBL
Affinity DataKi: 1.90E+4nMAssay Description:Ligand binding affinity was determined by displacement of a tritiated tracer by the unlabeled compound to a GST-PPAR fusion protein for Peroxisome pr...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Homo sapiens (Human))
Biovitrum
Curated by ChEMBL
Biovitrum
Curated by ChEMBL
Affinity DataEC50: 600nMAssay Description:Transactivation potency was measured by luciferase activity in Caco- 2/TC7 cells transiently co-transfected for the fusion-protein Gal4-PPAR gammaMore data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor alpha(Homo sapiens (Human))
Biovitrum
Curated by ChEMBL
Biovitrum
Curated by ChEMBL
Affinity DataEC50: 2.50E+3nMAssay Description:Ligand binding affinity was determined by displacement of a tritiated tracer by the unlabeled compound to a GST-PPAR fusion protein for Peroxisome pr...More data for this Ligand-Target Pair