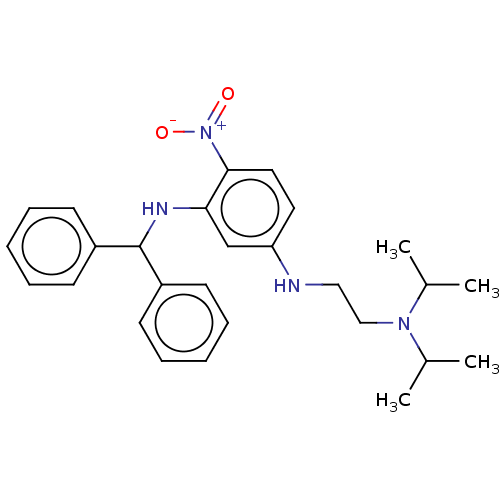
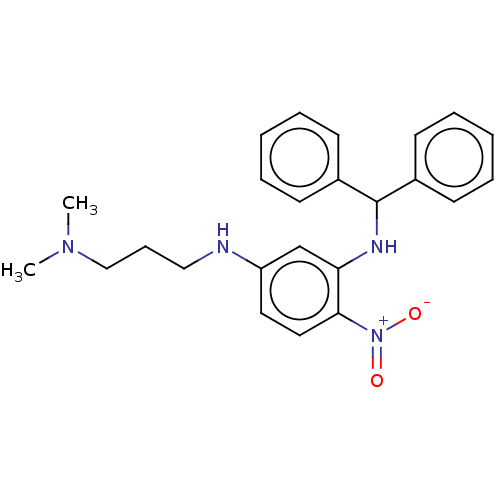
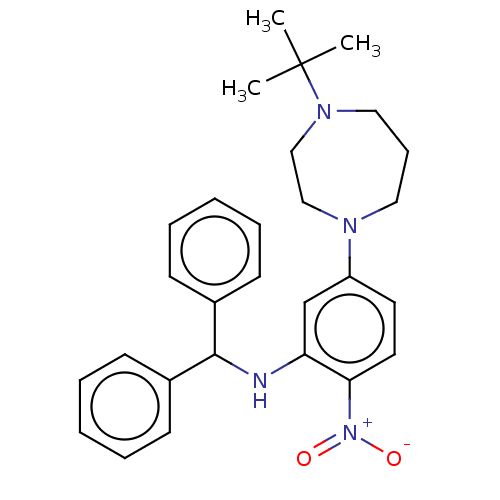
Affinity DataKi: 19nM ΔG°: -44.8kJ/molepH: 7.5 T: 2°CAssay Description:Reaction velocities were determined at each dGTP concentration and used to create double reciprocal plots of velocity versus dGTP concentration. The ...More data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:Competitive inhibition of Bacillus anthracis recombinant lethal factor expressed in Escherichia coli by linear double-reciprocal plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 63nM ΔG°: -41.8kJ/molepH: 7.5 T: 2°CAssay Description:Reaction velocities were determined at each dGTP concentration and used to create double reciprocal plots of velocity versus dGTP concentration. The ...More data for this Ligand-Target Pair
Affinity DataKi: 300nMAssay Description:Inhibition of Clostridium botulinum toxin BoNT/A light chainMore data for this Ligand-Target Pair
Affinity DataKi: 330nMAssay Description:Inhibition of Clostridium botulinum botulinum neurotoxin type A light chainMore data for this Ligand-Target Pair
Affinity DataKi: 478nMAssay Description:Non-competitive inhibition of Bacillus anthracis lethal factor using 3.12 uM MCA-KKVYPYPME[dnp]K amide as substrate after 30 mins by Eadie-Hofstee pl...More data for this Ligand-Target Pair
Affinity DataKi: 500nMAssay Description:Competitive inhibition of Bacillus anthracis lethal factorMore data for this Ligand-Target Pair
Affinity DataKi: 550nMAssay Description:Activity determined in mouse liver S-adenosyl-L-homocysteine hydrolase and expressed as KI values.More data for this Ligand-Target Pair
Affinity DataKi: 570nMAssay Description:Inhibition of Bacillus anthracis lethal factor by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 760nMAssay Description:Inhibition of Clostridium botulinum toxin BoNT/A light chainMore data for this Ligand-Target Pair
Affinity DataKi: 1.04E+3nMAssay Description:Activity determined in mouse liver S-adenosyl-L-homocysteine hydrolase and expressed as KI values.More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nMAssay Description:Inhibition of Bacillus anthracis lethal factorMore data for this Ligand-Target Pair
Affinity DataKi: 1.14E+3nMAssay Description:Non-competitive inhibition of Bacillus anthracis lethal factor assessed as MCA-KKVYPYPME[dnp]K amide cleavage after 30 mins by Eadie-Hofstee plot ana...More data for this Ligand-Target Pair
Affinity DataKi: 1.26E+3nMAssay Description:Non-competitive inhibition of Bacillus anthracis lethal factor using 3.12 uM MCA-KKVYPYPME[dnp]K amide as substrate after 30 mins by Eadie-Hofstee pl...More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+3nMAssay Description:Activity determined in rat liver S-adenosyl-L-homocysteine hydrolase and expressed as Kinactivator values; NA= not applicableMore data for this Ligand-Target Pair
Affinity DataKi: 1.60E+3nMAssay Description:Activity determined in rat liver S-adenosyl-L-homocysteine hydrolase and expressed as Kinactivator values; NA= not applicableMore data for this Ligand-Target Pair
Affinity DataKi: 2.40E+3nMAssay Description:Activity determined in rat liver S-adenosyl-L-homocysteine hydrolase and expressed as KI values.More data for this Ligand-Target Pair
Affinity DataKi: 2.88E+3nMAssay Description:Non-competitive inhibition of Bacillus anthracis lethal factor assessed as MCA-KKVYPYPME[dnp]K amide cleavage after 30 mins by Eadie-Hofstee plot ana...More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+3nMAssay Description:Competitive inhibitory activity against rat liver S-Adenosyl-homocysteine hydrolaseMore data for this Ligand-Target Pair
Affinity DataKi: 3.20E+3nMAssay Description:Competitive inhibitory activity against rat liver S-Adenosyl-homocysteine hydrolaseMore data for this Ligand-Target Pair
Affinity DataKi: 6.50E+3nMAssay Description:Activity determined in rat liver S-adenosyl-L-homocysteine hydrolase and expressed as KI values.More data for this Ligand-Target Pair
Affinity DataKi: 9.70E+3nMAssay Description:Activity determined in rat liver S-adenosyl-L-homocysteine hydrolase and expressed as Kinactivator values.More data for this Ligand-Target Pair
Affinity DataKi: 9.70E+3nMAssay Description:Competitive inhibitory activity against rat liver S-adenosyl-L-homocysteine hydrolaseMore data for this Ligand-Target Pair
Affinity DataKi: 1.00E+4nMAssay Description:Activity determined in mouse liver S-adenosyl-L-homocysteine hydrolase and expressed as KI values.More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+4nMAssay Description:Competitive inhibitory activity against rat liver S-Adenosyl-homocysteine hydrolaseMore data for this Ligand-Target Pair
Affinity DataKi: 3.00E+4nMAssay Description:Activity determined in mouse liver S-adenosyl-L-homocysteine hydrolase and expressed as KI values.More data for this Ligand-Target Pair
Affinity DataKi: 1.17E+5nM ΔG°: -22.8kJ/molepH: 7.5 T: 2°CAssay Description:Reaction velocities were determined at each dGTP concentration and used to create double reciprocal plots of velocity versus dGTP concentration. The ...More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+5nMAssay Description:Activity determined in rat liver S-adenosyl-L-homocysteine hydrolase and expressed as Kinactivator values; NA= not applicableMore data for this Ligand-Target Pair
Affinity DataKi: >4.82E+5nM ΔG°: >-19.7kJ/molepH: 7.5 T: 2°CAssay Description:Reaction velocities were determined at each dGTP concentration and used to create double reciprocal plots of velocity versus dGTP concentration. The ...More data for this Ligand-Target Pair
Affinity DataKi: >4.82E+5nM ΔG°: >-19.2kJ/molepH: 7.5 T: 2°CAssay Description:Reaction velocities were determined at each dGTP concentration and used to create double reciprocal plots of velocity versus dGTP concentration. The ...More data for this Ligand-Target Pair
Affinity DataKi: >4.82E+5nM ΔG°: >-19.2kJ/molepH: 7.5 T: 2°CAssay Description:Reaction velocities were determined at each dGTP concentration and used to create double reciprocal plots of velocity versus dGTP concentration. The ...More data for this Ligand-Target Pair
Affinity DataKi: 6.84E+5nM ΔG°: -18.8kJ/molepH: 7.5 T: 2°CAssay Description:Reaction velocities were determined at each dGTP concentration and used to create double reciprocal plots of velocity versus dGTP concentration. The ...More data for this Ligand-Target Pair
Affinity DataKi: >1.01E+6nM ΔG°: >-17.8kJ/molepH: 7.5 T: 2°CAssay Description:Reaction velocities were determined at each dGTP concentration and used to create double reciprocal plots of velocity versus dGTP concentration. The ...More data for this Ligand-Target Pair
Affinity DataKi: >1.01E+6nM ΔG°: >-17.4kJ/molepH: 7.5 T: 2°CAssay Description:Reaction velocities were determined at each dGTP concentration and used to create double reciprocal plots of velocity versus dGTP concentration. The ...More data for this Ligand-Target Pair
Affinity DataKi: >1.01E+6nM ΔG°: >-17.8kJ/molepH: 7.5 T: 2°CAssay Description:Reaction velocities were determined at each dGTP concentration and used to create double reciprocal plots of velocity versus dGTP concentration. The ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.70nMAssay Description:Inhibition of SAHase from rabbit erythrocytes using S-adenosyl-L-homocysteine as substrate preincubated for 5 mins followed by substrate addition and...More data for this Ligand-Target Pair
Affinity DataIC50: <100nMAssay Description:Recombinant vesicular stomatitis viruses (VSVs) (serotype Indiana) expressing eGFP and EBOV, SUDV, or BDBV GP in place of VSV G, as well as those exp...More data for this Ligand-Target Pair
Affinity DataIC50: <100nMAssay Description:Recombinant vesicular stomatitis viruses (VSVs) (serotype Indiana) expressing eGFP and EBOV, SUDV, or BDBV GP in place of VSV G, as well as those exp...More data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:Recombinant vesicular stomatitis viruses (VSVs) (serotype Indiana) expressing eGFP and EBOV, SUDV, or BDBV GP in place of VSV G, as well as those exp...More data for this Ligand-Target Pair
Affinity DataIC50: 390nMAssay Description:Recombinant vesicular stomatitis viruses (VSVs) (serotype Indiana) expressing eGFP and EBOV, SUDV, or BDBV GP in place of VSV G, as well as those exp...More data for this Ligand-Target Pair
Affinity DataIC50: 410nMAssay Description:Inhibition of Clostridium botulinum toxin BoNT/A light chainMore data for this Ligand-Target Pair
Affinity DataIC50: 480nMAssay Description:The authentic filoviruses Ebola virus/H. sapiens-tc/COD/1995/Kikwit-9510621 (EBOV/Kik-9510621; �EBOV-Zaire 1995�) (Jahrling et al., J. Infect. Dis., ...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:Inhibition of Clostridium botulinum toxin BoNT/A light chainMore data for this Ligand-Target Pair
Affinity DataIC50: 530nMAssay Description:The authentic filoviruses Ebola virus/H. sapiens-tc/COD/1995/Kikwit-9510621 (EBOV/Kik-9510621; �EBOV-Zaire 1995�) (Jahrling et al., J. Infect. Dis., ...More data for this Ligand-Target Pair
Affinity DataIC50: 560nMAssay Description:Recombinant vesicular stomatitis viruses (VSVs) (serotype Indiana) expressing eGFP and EBOV, SUDV, or BDBV GP in place of VSV G, as well as those exp...More data for this Ligand-Target Pair
Affinity DataIC50: 560nMAssay Description:The authentic filoviruses Ebola virus/H. sapiens-tc/COD/1995/Kikwit-9510621 (EBOV/Kik-9510621; �EBOV-Zaire 1995�) (Jahrling et al., J. Infect. Dis., ...More data for this Ligand-Target Pair
Affinity DataIC50: 580nMAssay Description:Recombinant vesicular stomatitis viruses (VSVs) (serotype Indiana) expressing eGFP and EBOV, SUDV, or BDBV GP in place of VSV G, as well as those exp...More data for this Ligand-Target Pair
Affinity DataIC50: 600nMAssay Description:Recombinant vesicular stomatitis viruses (VSVs) (serotype Indiana) expressing eGFP and EBOV, SUDV, or BDBV GP in place of VSV G, as well as those exp...More data for this Ligand-Target Pair
Affinity DataIC50: 600nMAssay Description:The authentic filoviruses Ebola virus/H. sapiens-tc/COD/1995/Kikwit-9510621 (EBOV/Kik-9510621; �EBOV-Zaire 1995�) (Jahrling et al., J. Infect. Dis., ...More data for this Ligand-Target Pair
Affinity DataIC50: <600nMAssay Description:The authentic filoviruses Ebola virus/H. sapiens-tc/COD/1995/Kikwit-9510621 (EBOV/Kik-9510621; �EBOV-Zaire 1995�) (Jahrling et al., J. Infect. Dis., ...More data for this Ligand-Target Pair

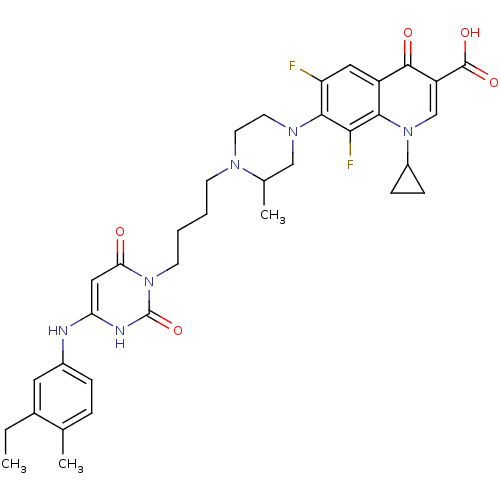
 3D Structure (crystal)
3D Structure (crystal)