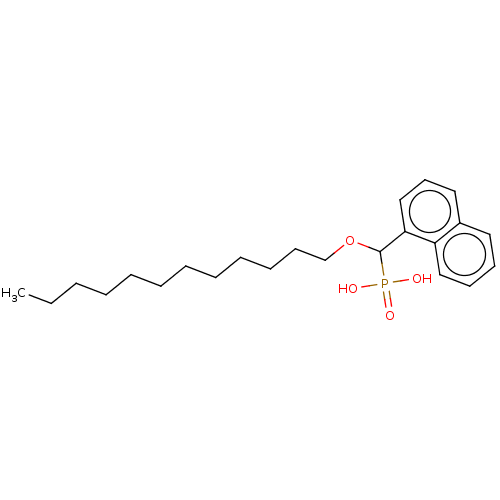
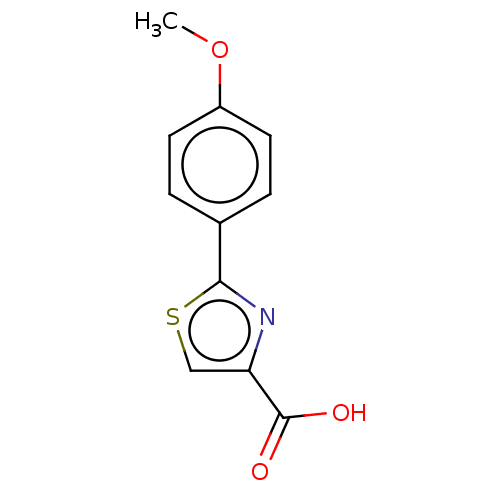
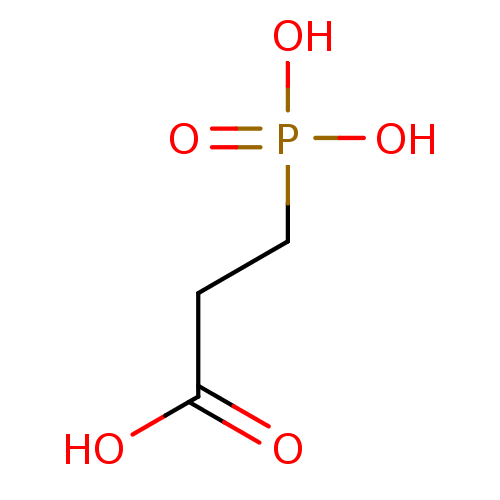
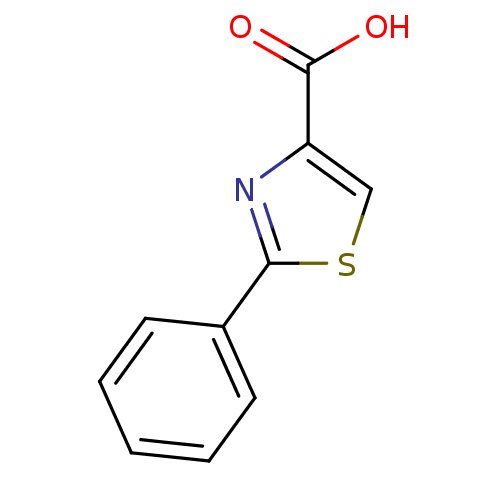
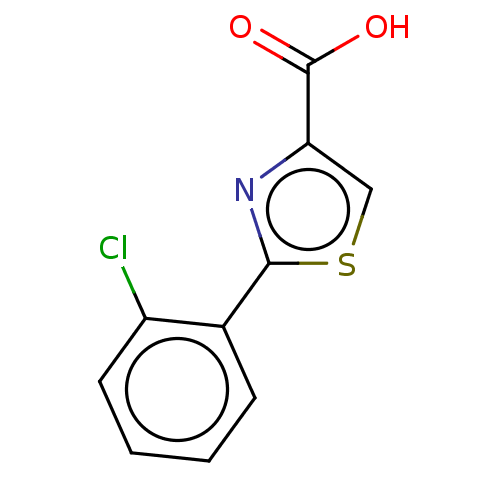
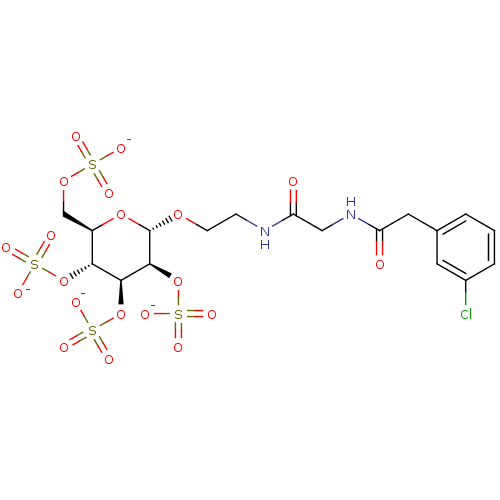
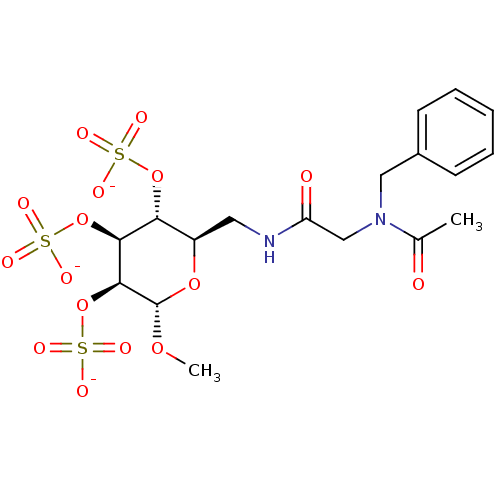
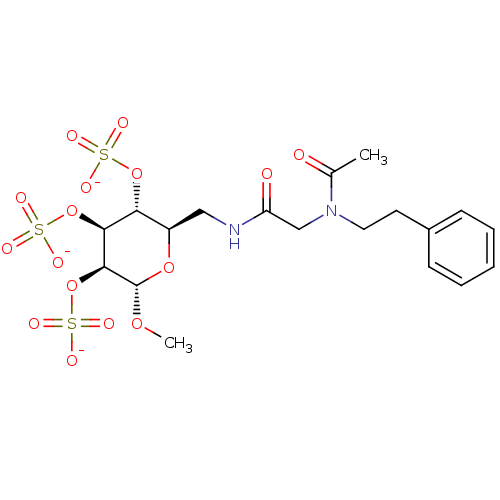
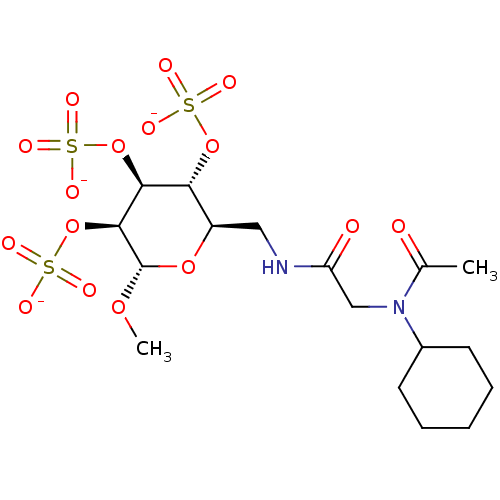
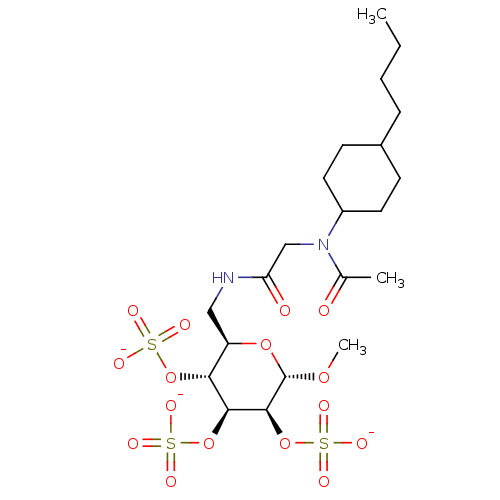
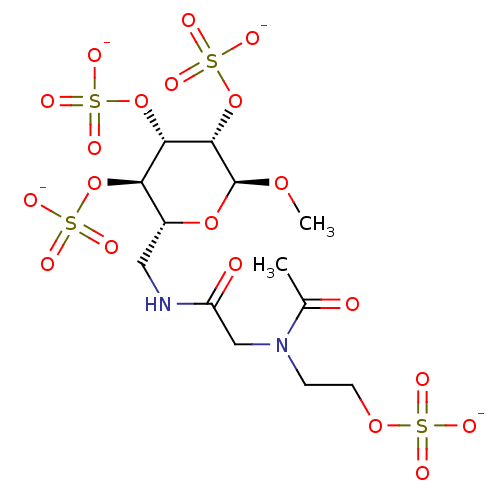
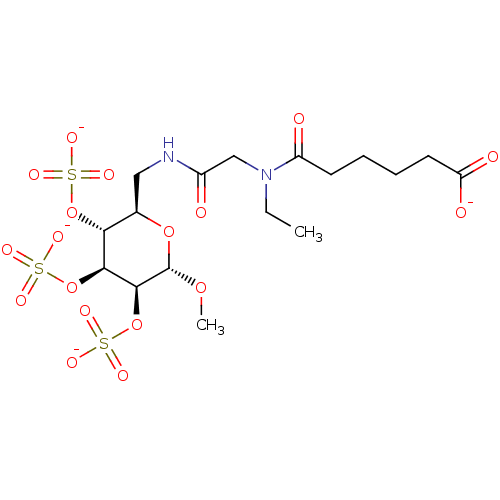
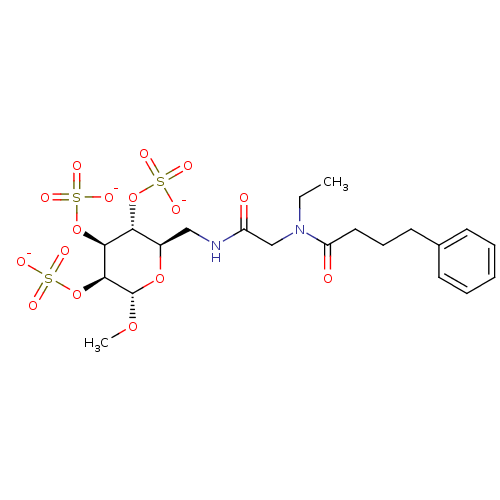
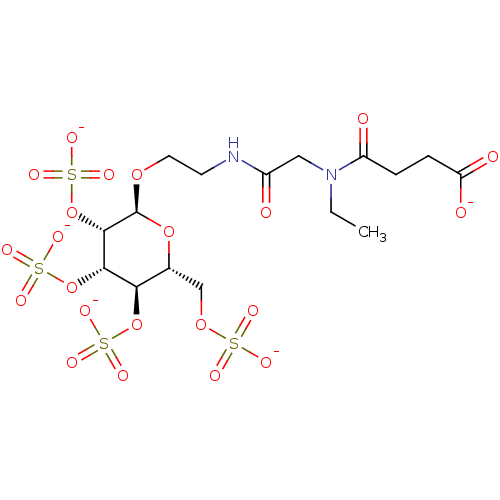
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKi: 200nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
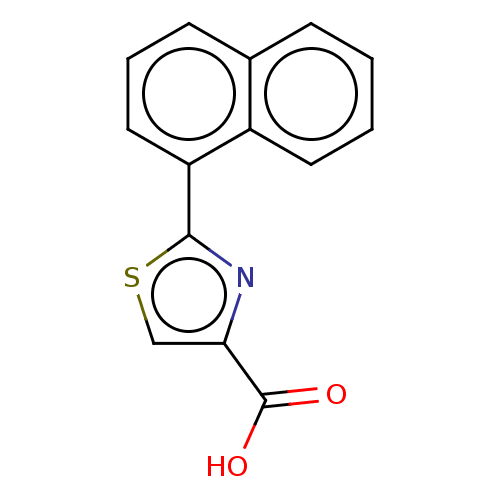
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKi: 400nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
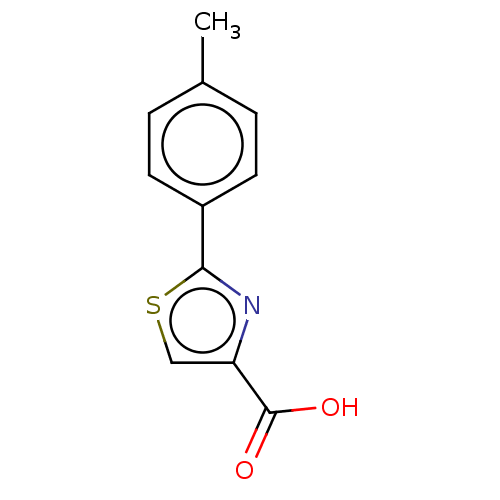
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKi: 500nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
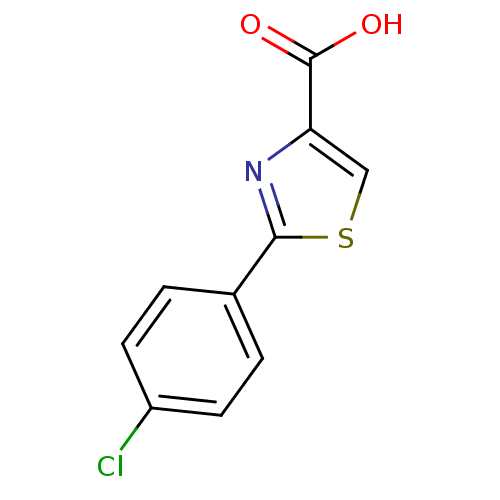
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKi: 800nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKi: 4.50E+3nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKi: 1.40E+4nMAssay Description:Competitive inhibition of red kidney bean PAP using varying levels of pNPP as substrate measured at pH 6.2 by UV-vis spectrophotometric analysisMore data for this Ligand-Target Pair
TargetTartrate-resistant acid phosphatase type 5(Sus scrofa)
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKi: 3.30E+4nMAssay Description:Competitive inhibition of pig purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV/Vis spectro...More data for this Ligand-Target Pair
TargetTartrate-resistant acid phosphatase type 5(Sus scrofa)
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKi: 4.20E+4nMAssay Description:Competitive inhibition of pig purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV/Vis spectro...More data for this Ligand-Target Pair
TargetTartrate-resistant acid phosphatase type 5(Sus scrofa)
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKi: 4.90E+4nMAssay Description:Competitive inhibition of pig purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV/Vis spectro...More data for this Ligand-Target Pair
TargetTartrate-resistant acid phosphatase type 5(Sus scrofa)
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKi: 1.20E+5nMAssay Description:Competitive inhibition of pig purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV/Vis spectro...More data for this Ligand-Target Pair
TargetFe(3+)-Zn(2+) purple acid phosphatase(Phaseolus vulgaris)
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKi: 1.60E+5nMAssay Description:Inhibition of red kidney bean PAP using pNPP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 1.85E+5nMAssay Description:Competitive inhibition of red kidney bean purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV...More data for this Ligand-Target Pair
TargetTartrate-resistant acid phosphatase type 5(Sus scrofa)
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKi: 1.90E+5nMAssay Description:Competitive inhibition of pig purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV/Vis spectro...More data for this Ligand-Target Pair
Affinity DataKi: 3.40E+5nMAssay Description:Competitive inhibition of red kidney bean purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV...More data for this Ligand-Target Pair
Affinity DataKi: 4.00E+5nMAssay Description:Competitive inhibition of red kidney bean purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV...More data for this Ligand-Target Pair
Affinity DataKi: 5.30E+5nMAssay Description:Competitive inhibition of red kidney bean purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV...More data for this Ligand-Target Pair
TargetTartrate-resistant acid phosphatase type 5(Sus scrofa)
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKi: 6.30E+5nMAssay Description:Competitive inhibition of pig purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV/Vis spectro...More data for this Ligand-Target Pair
Affinity DataKi: 9.20E+5nMAssay Description:Competitive inhibition of red kidney bean purple acid phosphatase Fe(3)Fe(2) state assessed as enzyme-inhibitor complex using pNPP as substrate by UV...More data for this Ligand-Target Pair
TargetVascular endothelial growth factor A(Homo sapiens (Human))
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKd: 9.10E+4nMAssay Description:Binding affinity to VEGF by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor A(Homo sapiens (Human))
The University Of Queensland
Curated by ChEMBL
The University Of Queensland
Curated by ChEMBL
Affinity DataKd: 7.60E+4nMAssay Description:Binding affinity to VEGF by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 4.80E+5nMAssay Description:Binding affinity to FGF2 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 1.40E+5nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 2.24E+5nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 3.10E+5nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 2.65E+5nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 2.43E+5nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 3.70E+4nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 9.50E+4nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 1.74E+5nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 1.60E+5nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 1.11E+5nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 3.66E+5nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 5.00E+5nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 2.50E+4nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 1.18E+4nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 4.30E+4nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 2.00E+4nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 3.43E+5nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 1.30E+5nMAssay Description:Binding affinity to FGF1 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 4.80E+5nMAssay Description:Binding affinity to FGF2 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 6.80E+5nMAssay Description:Binding affinity to FGF2 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 4.13E+5nMAssay Description:Binding affinity to FGF2 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 4.10E+5nMAssay Description:Binding affinity to FGF2 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 5.40E+5nMAssay Description:Binding affinity to FGF2 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 2.40E+5nMAssay Description:Binding affinity to FGF2 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 2.91E+5nMAssay Description:Binding affinity to FGF2 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 4.80E+5nMAssay Description:Binding affinity to FGF2 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 4.80E+5nMAssay Description:Binding affinity to FGF2 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 4.00E+5nMAssay Description:Binding affinity to FGF2 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair
Affinity DataKd: 2.40E+5nMAssay Description:Binding affinity to FGF2 by surface plasmon resonance-based solution affinity assayMore data for this Ligand-Target Pair

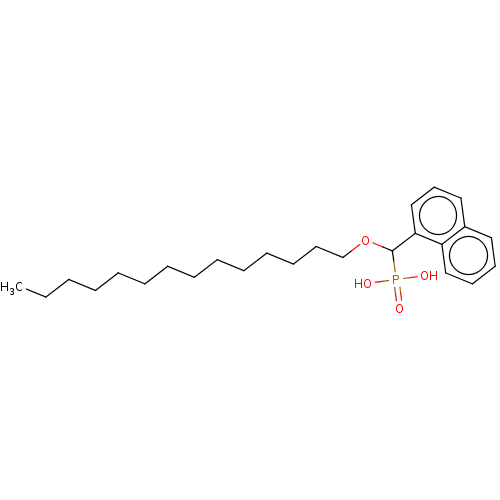
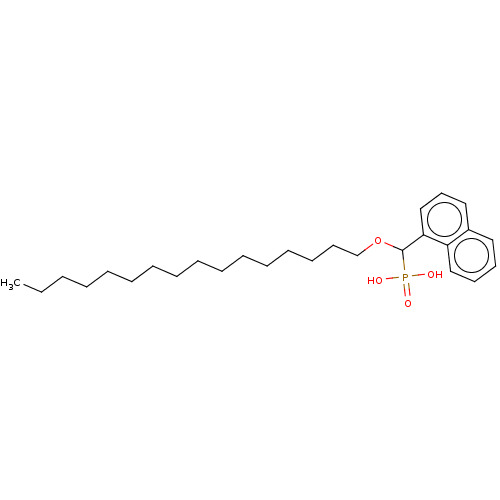
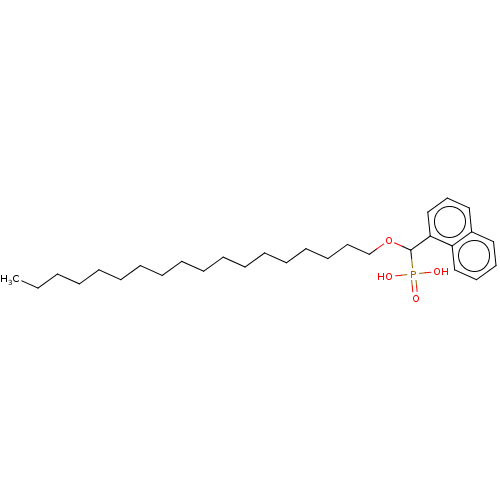
 3D Structure (crystal)
3D Structure (crystal)