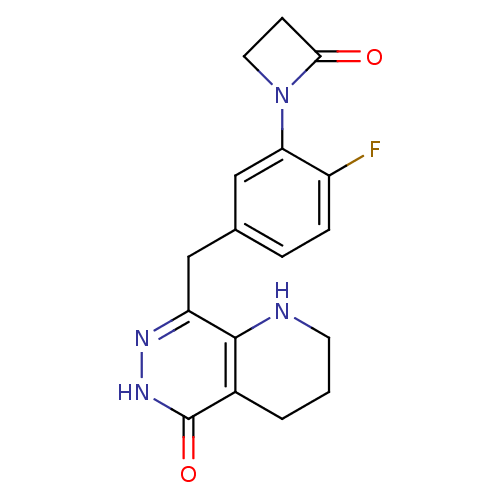
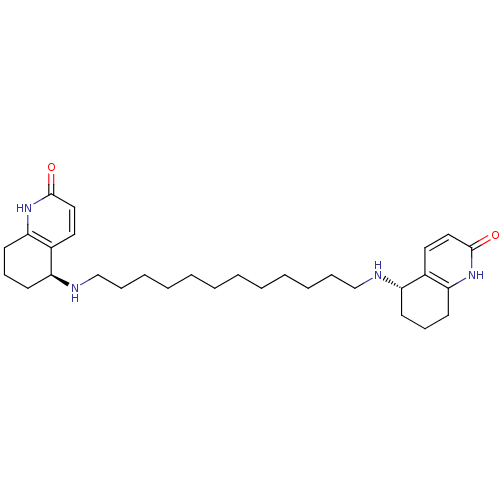
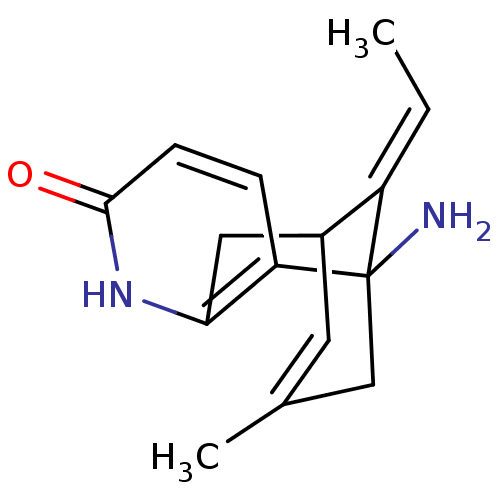
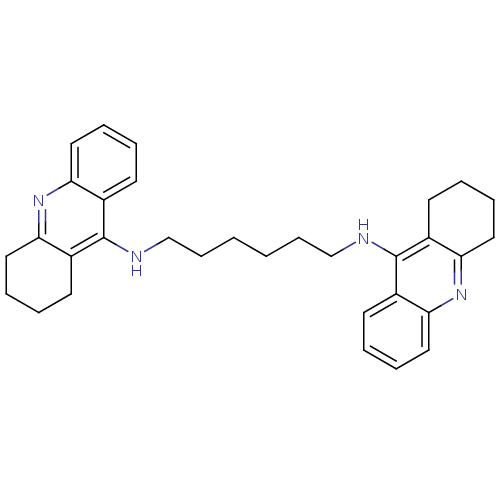
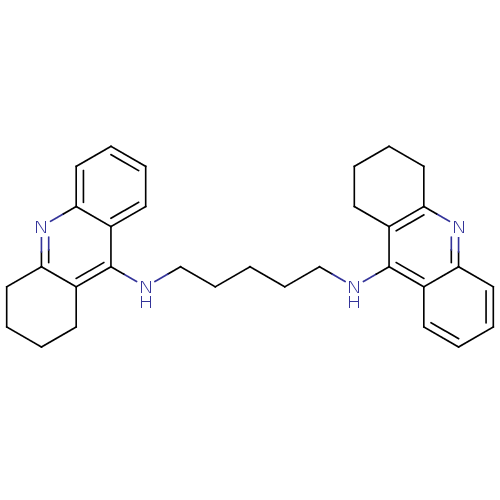
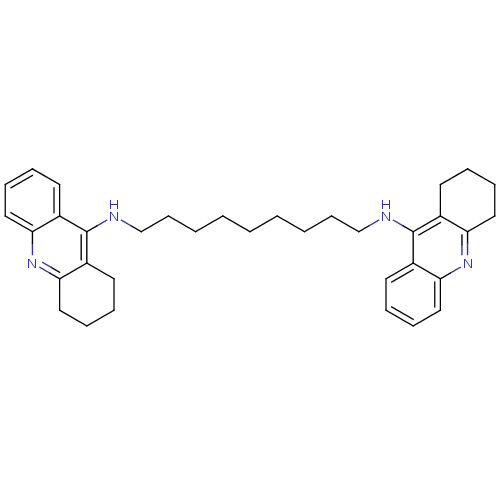
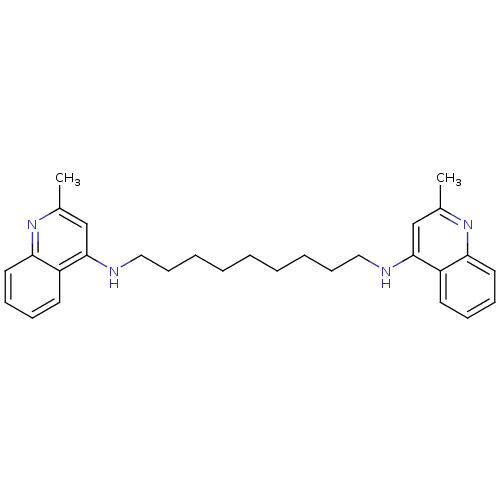
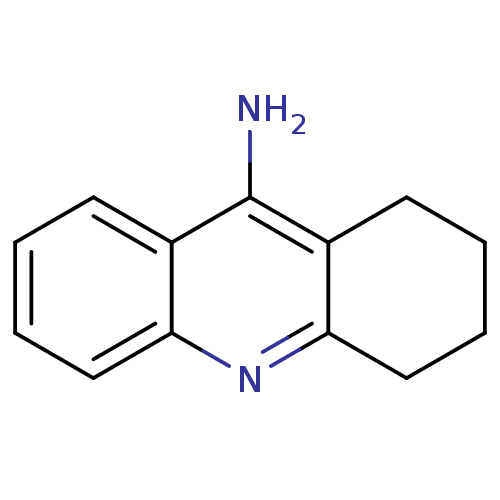
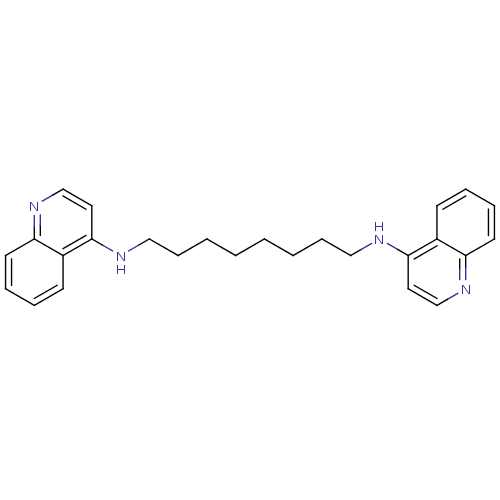
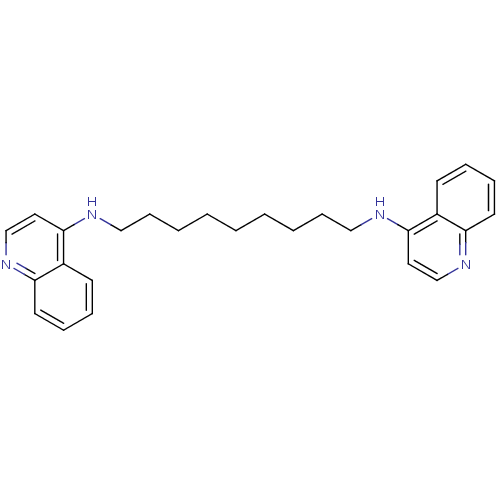
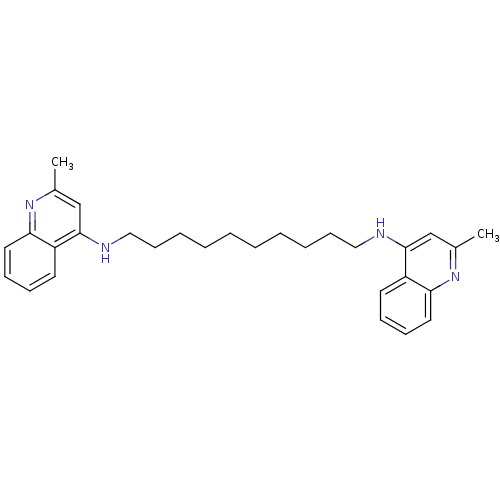
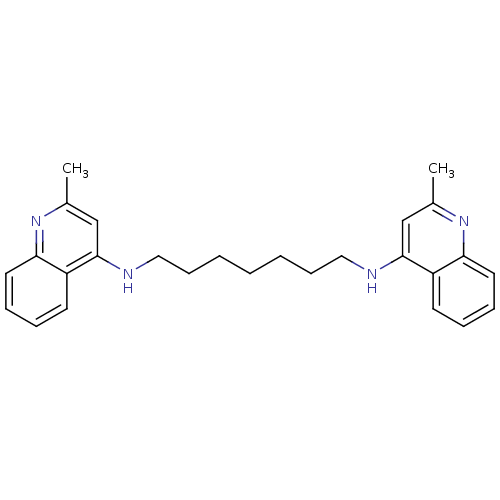
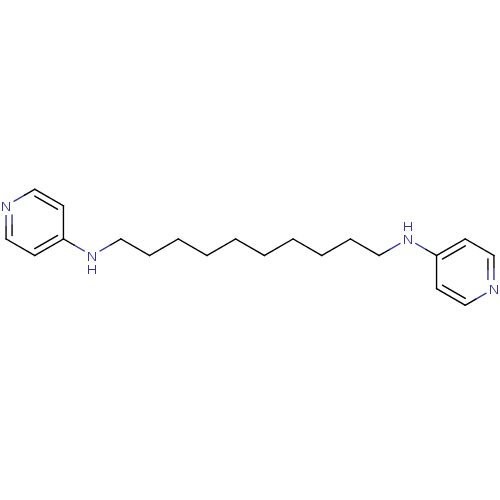
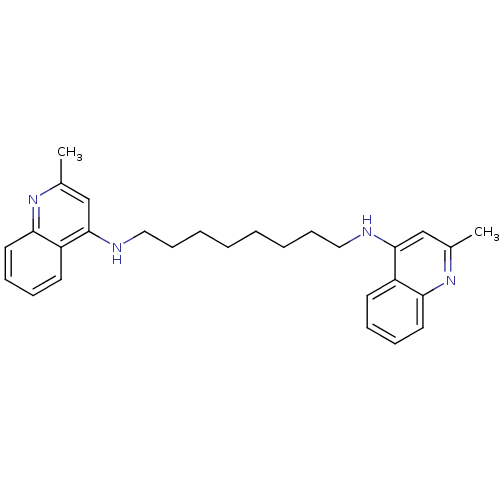
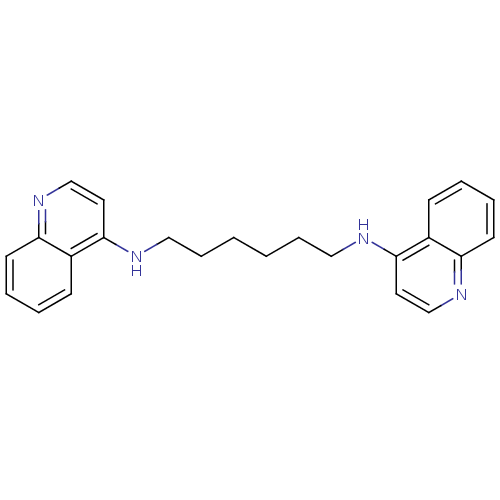
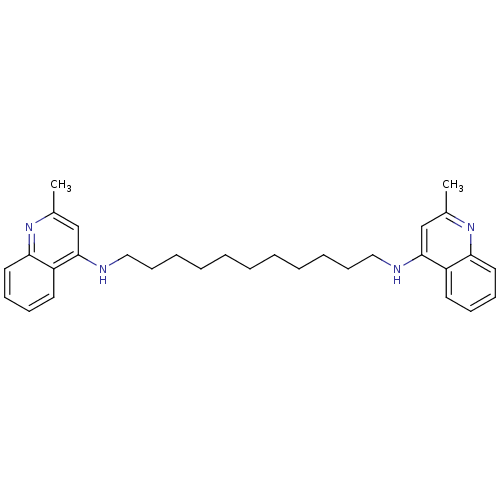
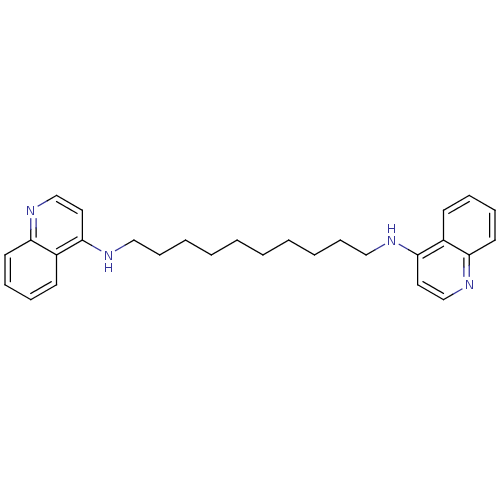
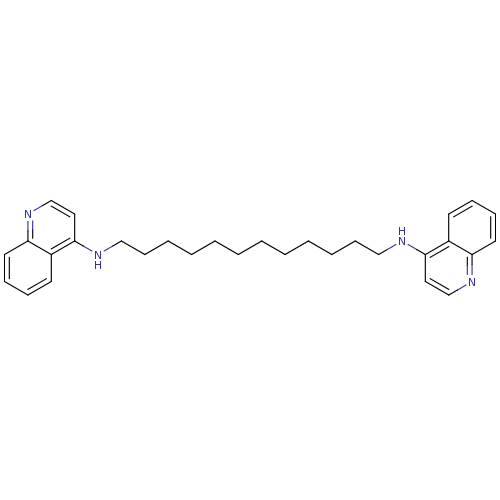
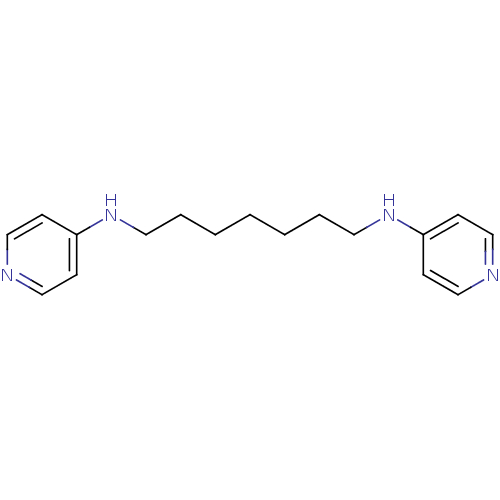
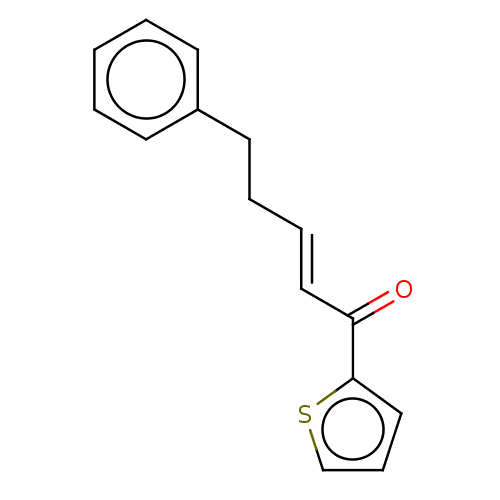
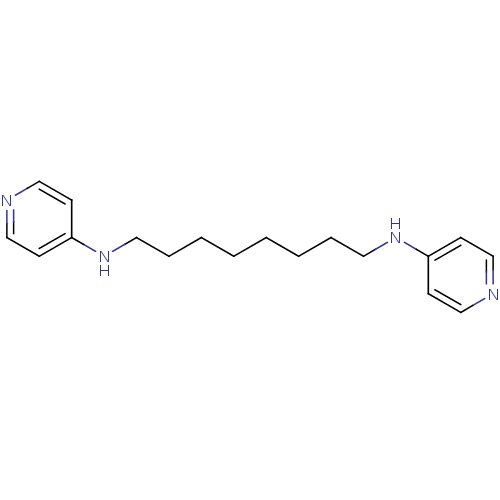
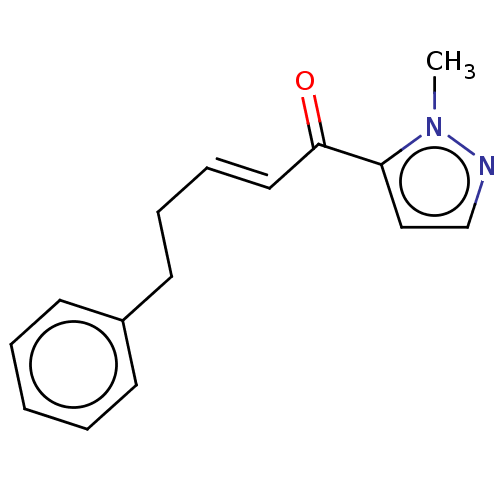
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Weizmann Institute of Science
Weizmann Institute of Science
Affinity DataKi: 0.800nM ΔG°: -51.4kJ/mole IC50: 2.40nMpH: 7.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at ...More data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Binding affinity to PARP-1 (unknown origin) assessed as inhibition constant incubated for 1 hr by topcount microplate scintillation counter analysisMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Weizmann Institute of Science
Weizmann Institute of Science
Affinity DataKi: 4.5nM ΔG°: -47.2kJ/mole IC50: 16nMpH: 7.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at ...More data for this Ligand-Target Pair
Affinity DataKi: 19.6nM ΔG°: -43.6kJ/mole IC50: 52nMpH: 7.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at ...More data for this Ligand-Target Pair
Affinity DataKi: 47.1nM ΔG°: -41.4kJ/mole IC50: 114nMpH: 7.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at ...More data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Weizmann Institute of Science
Weizmann Institute of Science
Affinity DataKi: 175nM ΔG°: -38.2kJ/mole IC50: 414nMpH: 7.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.80nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 28nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 31nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 54.1nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 87.8nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
TargetCarboxylic ester hydrolase(Rattus norvegicus (rat))
Hong Kong University of Science and Technology
Hong Kong University of Science and Technology
Affinity DataIC50: 92nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 94.3nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
TargetCarboxylic ester hydrolase(Rattus norvegicus (rat))
Hong Kong University of Science and Technology
Hong Kong University of Science and Technology
Affinity DataIC50: 102nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
TargetCarboxylic ester hydrolase(Rattus norvegicus (rat))
Hong Kong University of Science and Technology
Hong Kong University of Science and Technology
Affinity DataIC50: 119nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 124nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 143nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 148nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 151nMpH: 7.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at ...More data for this Ligand-Target Pair
TargetCarboxylic ester hydrolase(Rattus norvegicus (rat))
Hong Kong University of Science and Technology
Hong Kong University of Science and Technology
Affinity DataIC50: 152nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 152nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
TargetCarboxylic ester hydrolase(Rattus norvegicus (rat))
Hong Kong University of Science and Technology
Hong Kong University of Science and Technology
Affinity DataIC50: 155nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 157nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 161nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
TargetCarboxylic ester hydrolase(Rattus norvegicus (rat))
Hong Kong University of Science and Technology
Hong Kong University of Science and Technology
Affinity DataIC50: 167nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 215nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 223nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 244nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
TargetCarboxylic ester hydrolase(Rattus norvegicus (rat))
Hong Kong University of Science and Technology
Hong Kong University of Science and Technology
Affinity DataIC50: 252nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 254nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 312nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
TargetCarboxylic ester hydrolase(Rattus norvegicus (rat))
Hong Kong University of Science and Technology
Hong Kong University of Science and Technology
Affinity DataIC50: 329nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
TargetCarboxylic ester hydrolase(Rattus norvegicus (rat))
Hong Kong University of Science and Technology
Hong Kong University of Science and Technology
Affinity DataIC50: 340nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 364nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 403nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 454nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 483nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 630nMAssay Description:Antagonist activity at GPR52 (unknown origin) expressed in HEK293 cells assessed as inhibition of WO459-induced cAMP accumulation preincubated for 15...More data for this Ligand-Target Pair
Affinity DataIC50: 650nMAssay Description:Antagonist activity at GPR52 (unknown origin) expressed in HEK293 cells assessed as inhibition of WO459-induced cAMP accumulation preincubated for 15...More data for this Ligand-Target Pair
Affinity DataIC50: 711nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 786nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 809nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 880nMAssay Description:Antagonist activity at GPR52 (unknown origin) expressed in HEK293 cells assessed as inhibition of WO459-induced cAMP accumulation preincubated for 15...More data for this Ligand-Target Pair
Affinity DataIC50: 908nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 910nMAssay Description:Antagonist activity at GPR52 (unknown origin) expressed in HEK293 cells assessed as inhibition of WO459-induced cAMP accumulation preincubated for 15...More data for this Ligand-Target Pair
Affinity DataIC50: 1.01E+3nMAssay Description:Antagonist activity at GPR52 (unknown origin) expressed in HEK293 cells assessed as inhibition of WO459-induced cAMP accumulation preincubated for 15...More data for this Ligand-Target Pair
TargetCarboxylic ester hydrolase(Rattus norvegicus (rat))
Hong Kong University of Science and Technology
Hong Kong University of Science and Technology
Affinity DataIC50: 1.06E+3nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 1.08E+3nMAssay Description:Antagonist activity at GPR52 (unknown origin) expressed in HEK293 cells assessed as inhibition of WO459-induced cAMP accumulation preincubated for 15...More data for this Ligand-Target Pair
Affinity DataIC50: 1.23E+3nMAssay Description:Antagonist activity at GPR52 (unknown origin) expressed in HEK293 cells assessed as inhibition of WO459-induced cAMP accumulation preincubated for 15...More data for this Ligand-Target Pair

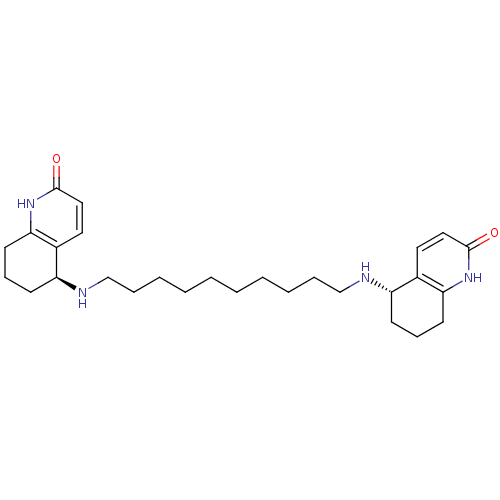
 3D Structure (crystal)
3D Structure (crystal)