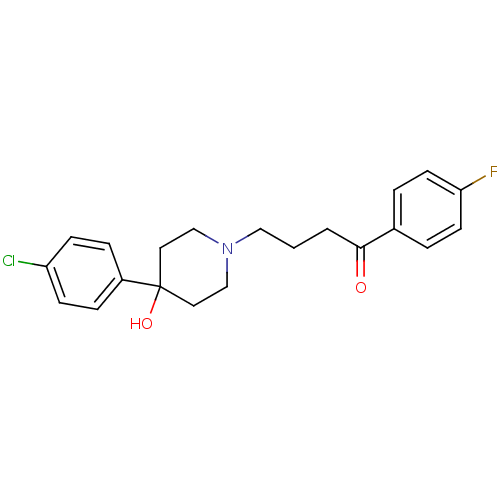
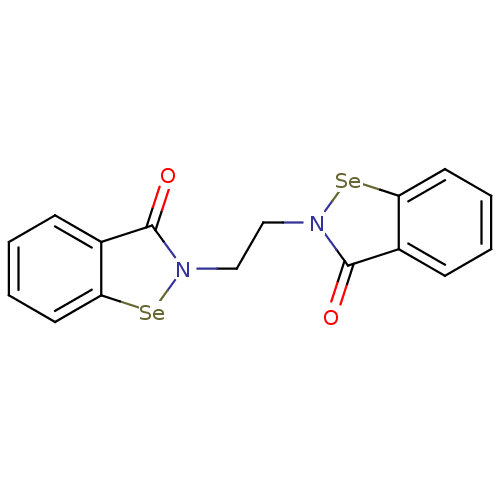
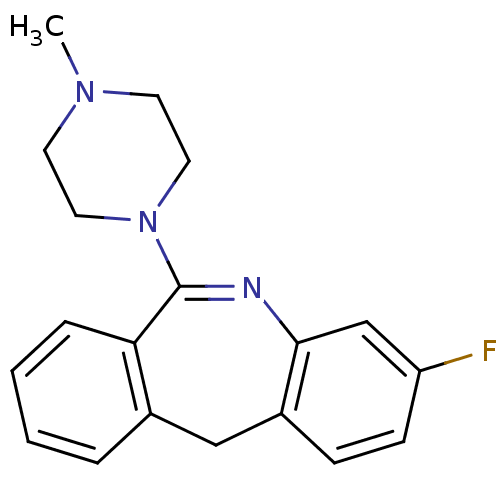
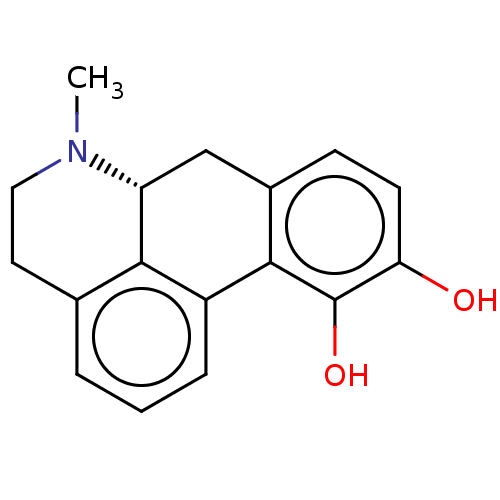
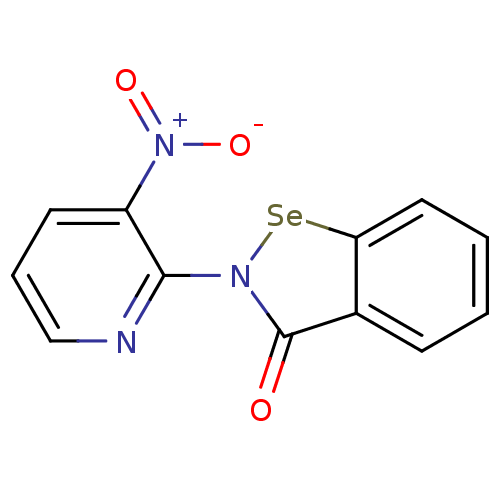
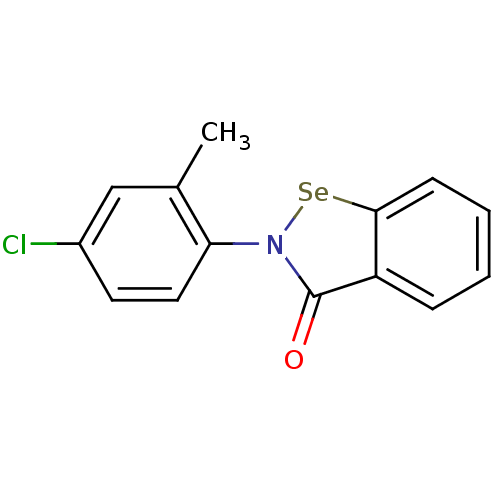
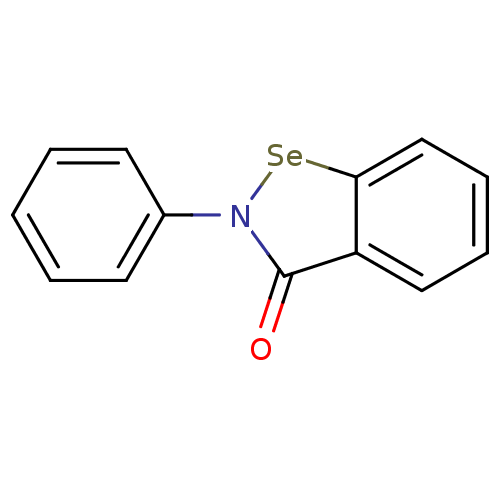
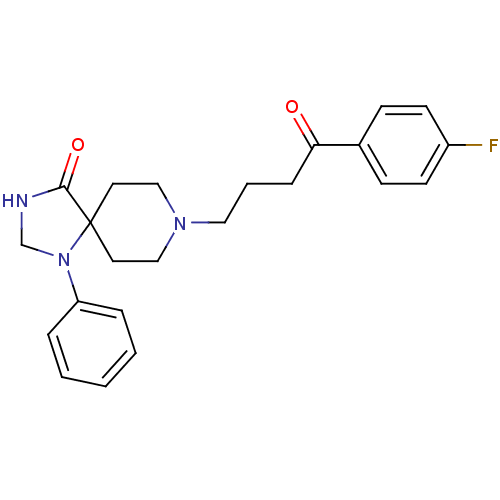
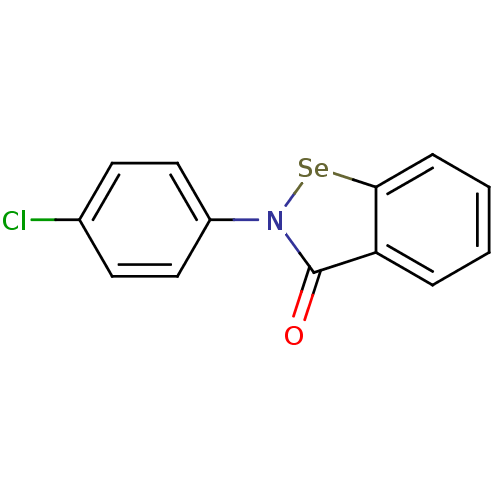
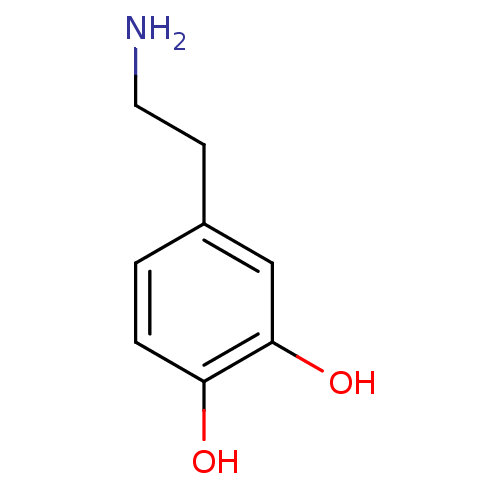
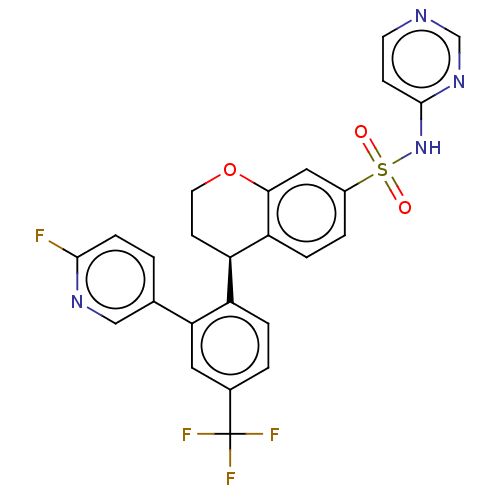
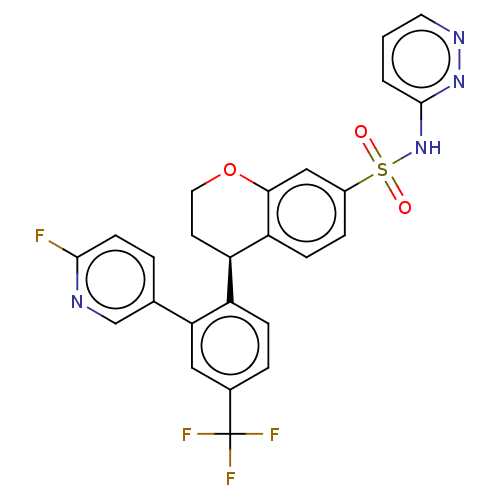
Affinity DataKi: 10nM IC50: 2.25E+3nMAssay Description:All the benzisoselenazol-3(2H)-one and bisbenzisoselenazol-3(2H)-one
derivatives were tested as potential E. coli TrxR inhibitors by standard
DTNB ...More data for this Ligand-Target Pair
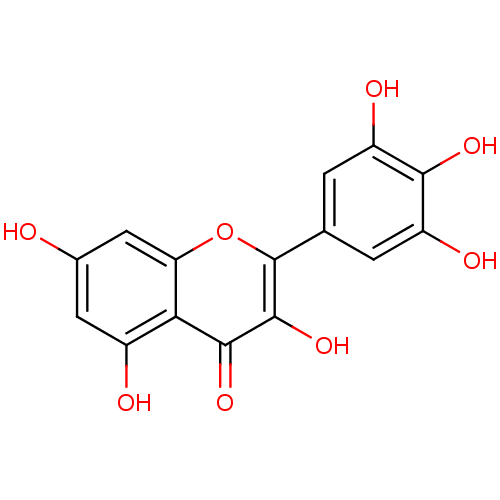
Affinity DataKi: 40nM IC50: 2.10E+3nMAssay Description:All the benzisoselenazol-3(2H)-one and bisbenzisoselenazol-3(2H)-one
derivatives were tested as potential E. coli TrxR inhibitors by standard
DTNB ...More data for this Ligand-Target Pair
TargetNonstructural protein 3(Zika virus)
University Of North Carolina At Chapel Hill
Curated by ChEMBL
University Of North Carolina At Chapel Hill
Curated by ChEMBL
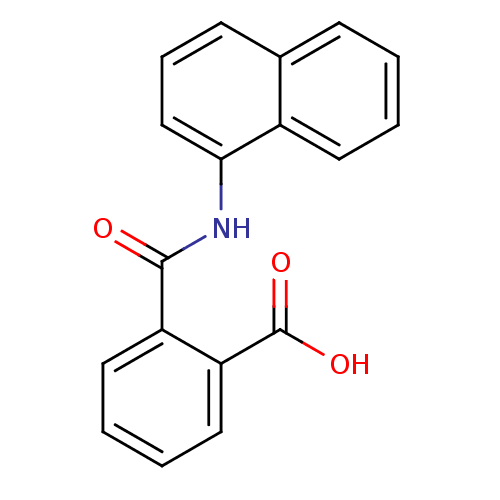
Affinity DataKi: 40nMAssay Description:Inhibition of N-terminal His6-tagged Zika virus NS2B (49 to 95 residues) - NS3 (1 to 170 residues) protease domain expressed in Escherichia coli BL21...More data for this Ligand-Target Pair
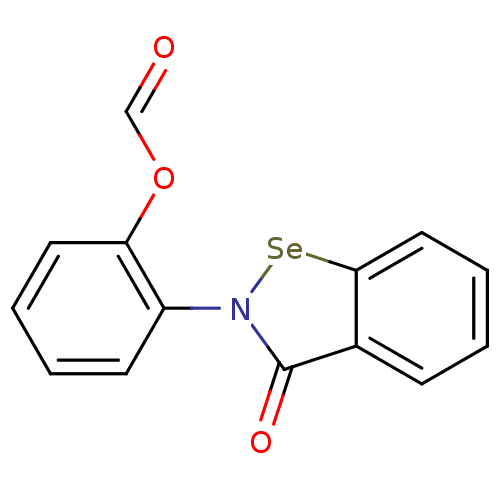
Affinity DataKi: 50nM IC50: 2.00E+3nMAssay Description:All the benzisoselenazol-3(2H)-one and bisbenzisoselenazol-3(2H)-one
derivatives were tested as potential E. coli TrxR inhibitors by standard
DTNB ...More data for this Ligand-Target Pair
Affinity DataKi: 250nM IC50: 3.00E+3nMAssay Description:All the benzisoselenazol-3(2H)-one and bisbenzisoselenazol-3(2H)-one
derivatives were tested as potential E. coli TrxR inhibitors by standard
DTNB ...More data for this Ligand-Target Pair
Affinity DataKi: 250nM IC50: 6.00E+3nMAssay Description:All the benzisoselenazol-3(2H)-one and bisbenzisoselenazol-3(2H)-one
derivatives were tested as potential E. coli TrxR inhibitors by standard
DTNB ...More data for this Ligand-Target Pair
Affinity DataKi: 300nM IC50: 6.00E+3nMAssay Description:All the benzisoselenazol-3(2H)-one and bisbenzisoselenazol-3(2H)-one
derivatives were tested as potential E. coli TrxR inhibitors by standard
DTNB ...More data for this Ligand-Target Pair
Affinity DataKi: 550nM IC50: 7.00E+3nMAssay Description:All the benzisoselenazol-3(2H)-one and bisbenzisoselenazol-3(2H)-one
derivatives were tested as potential E. coli TrxR inhibitors by standard
DTNB ...More data for this Ligand-Target Pair
TargetNonstructural protein 3(Zika virus)
University Of North Carolina At Chapel Hill
Curated by ChEMBL
University Of North Carolina At Chapel Hill
Curated by ChEMBL
Affinity DataKi: 800nMAssay Description:Inhibition of Zika virus Asian/8375 NS2B (48 to 100 residues)-NS3 (14 to 185 residues) expressed in Escherichia coli BL21 (DE3) Star cells preincubat...More data for this Ligand-Target Pair
Affinity DataKi: 1.00E+3nM IC50: 1.50E+4nMAssay Description:All the benzisoselenazol-3(2H)-one and bisbenzisoselenazol-3(2H)-one
derivatives were tested as potential E. coli TrxR inhibitors by standard
DTNB ...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nM IC50: 7.50E+3nMAssay Description:All the benzisoselenazol-3(2H)-one and bisbenzisoselenazol-3(2H)-one
derivatives were tested as potential E. coli TrxR inhibitors by standard
DTNB ...More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+3nM IC50: 3.00E+3nMAssay Description:All the benzisoselenazol-3(2H)-one and bisbenzisoselenazol-3(2H)-one
derivatives were tested as potential E. coli TrxR inhibitors by standard
DTNB ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0400nMAssay Description:HEK-293 cells overexpressing the channel of interest were seeded in a 96-well plate at a density of 30000 cells/well and incubated at 37� C./5% CO2 f...More data for this Ligand-Target Pair
Affinity DataIC50: 0.110nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:HEK-293 cells overexpressing the channel of interest were seeded in a 96-well plate at a density of 30000 cells/well and incubated at 37� C./5% CO2 f...More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:HEK-293 cells overexpressing the channel of interest were seeded in a 96-well plate at a density of 30000 cells/well and incubated at 37� C./5% CO2 f...More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:HEK-293 cells overexpressing the channel of interest were seeded in a 96-well plate at a density of 30000 cells/well and incubated at 37� C./5% CO2 f...More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:HEK-293 cells overexpressing the channel of interest were seeded in a 96-well plate at a density of 30000 cells/well and incubated at 37� C./5% CO2 f...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMAssay Description:HEK-293 cells overexpressing the channel of interest were seeded in a 96-well plate at a density of 30000 cells/well and incubated at 37� C./5% CO2 f...More data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair

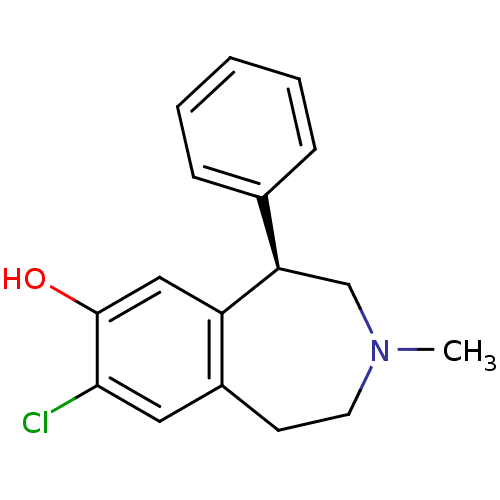

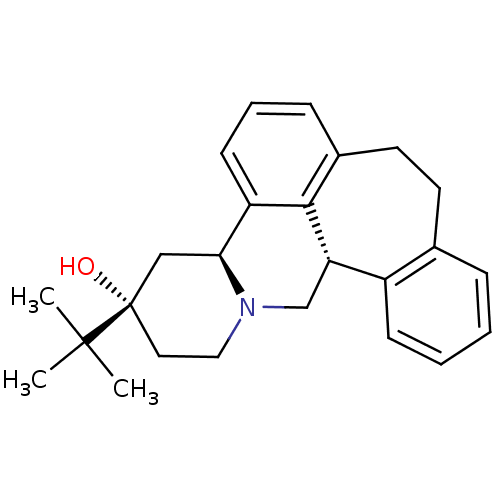
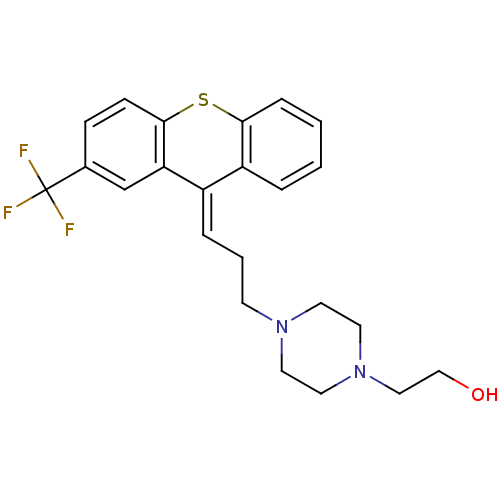
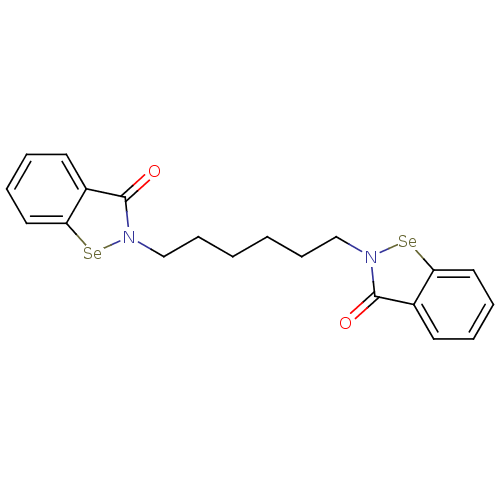

 3D Structure (crystal)
3D Structure (crystal)