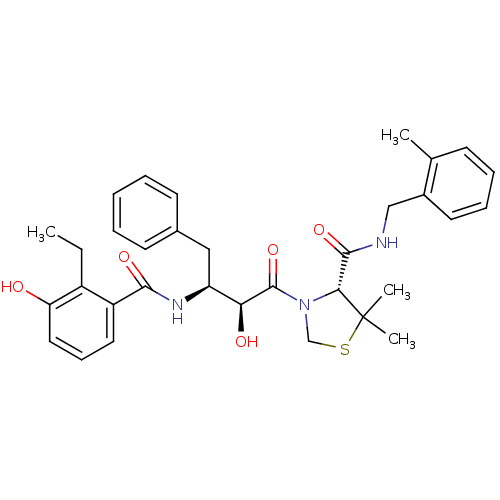
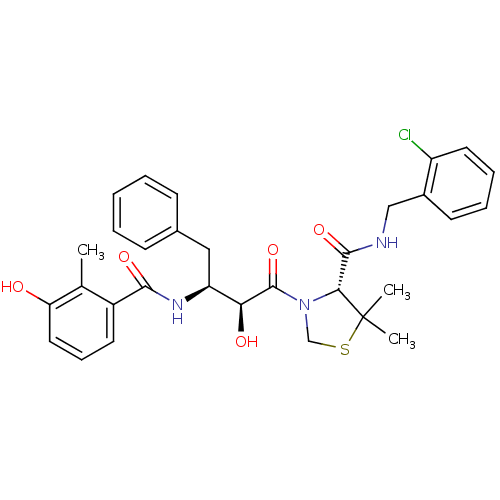
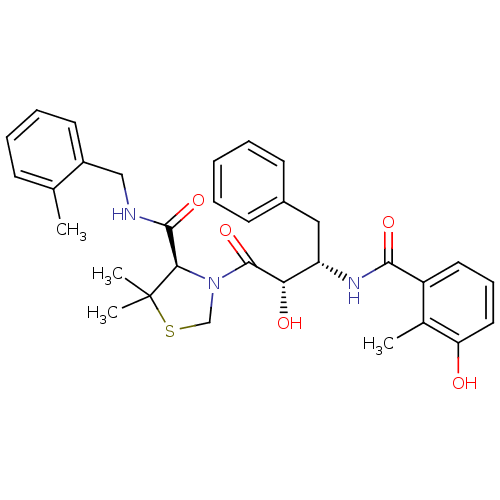
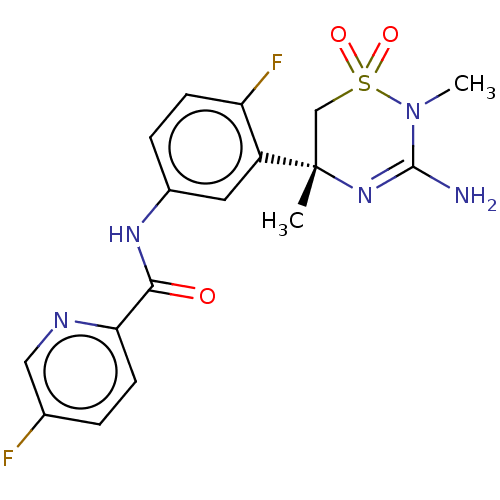
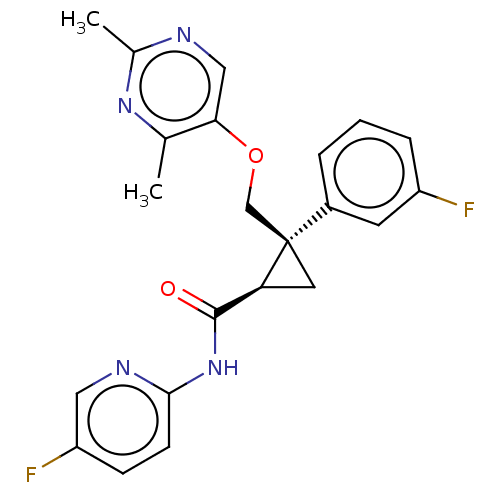
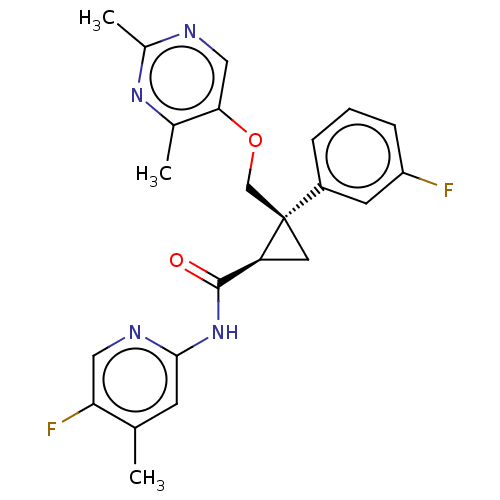
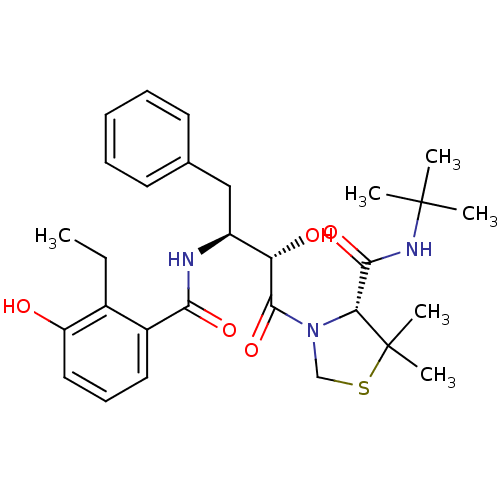
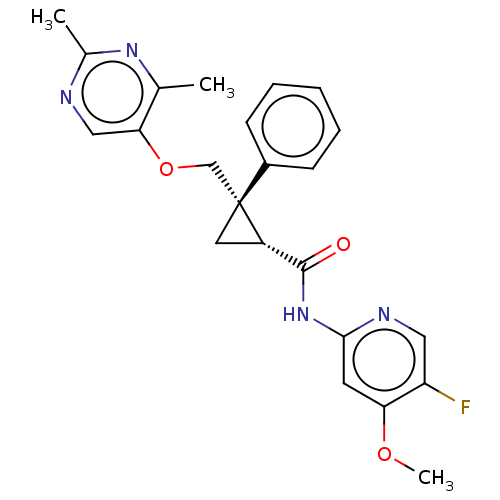
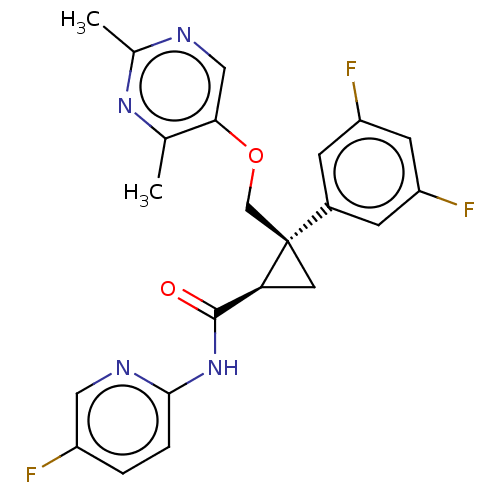
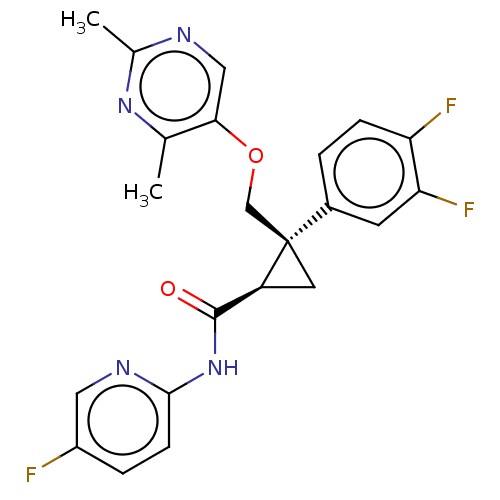
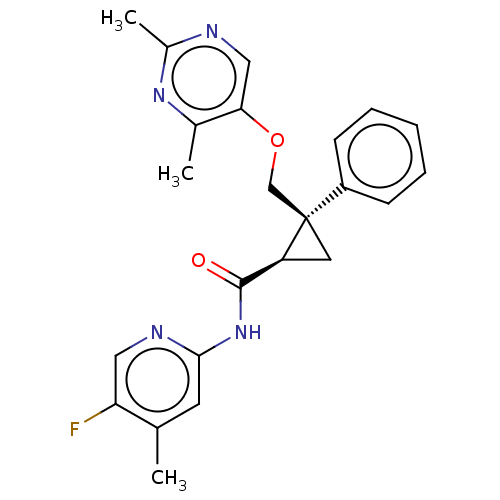
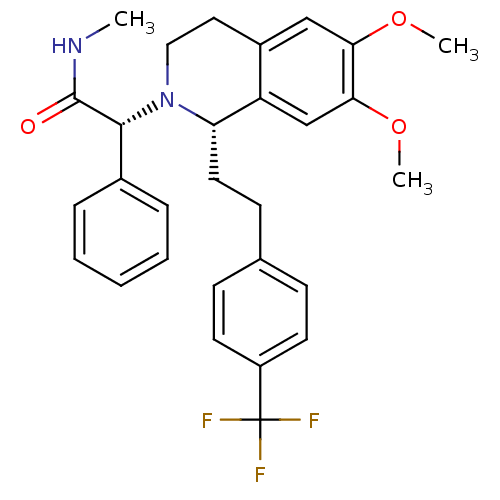
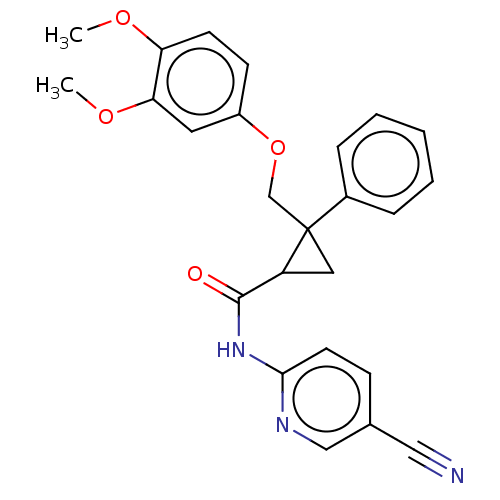
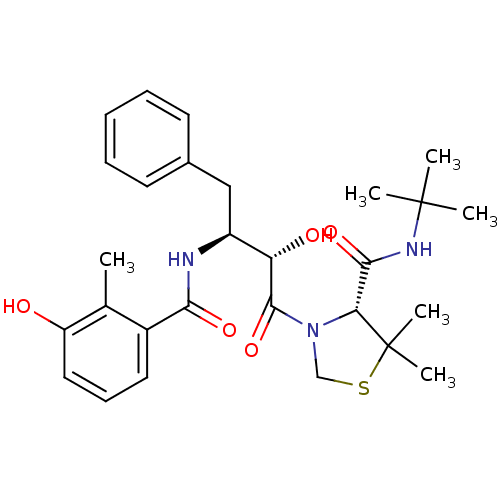
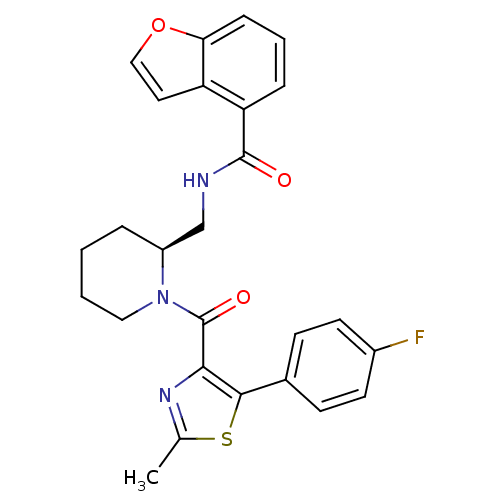
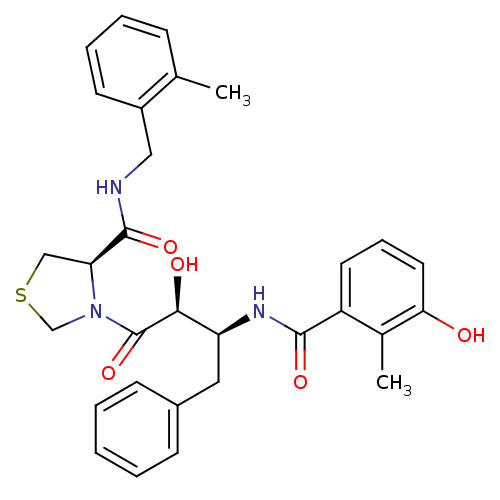
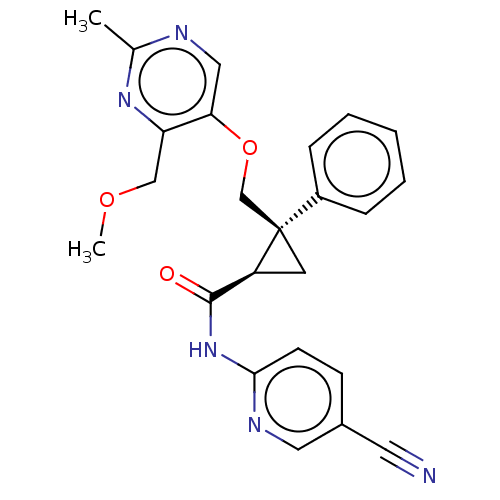
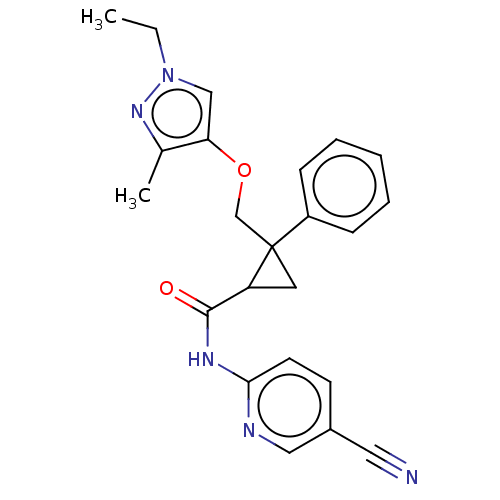
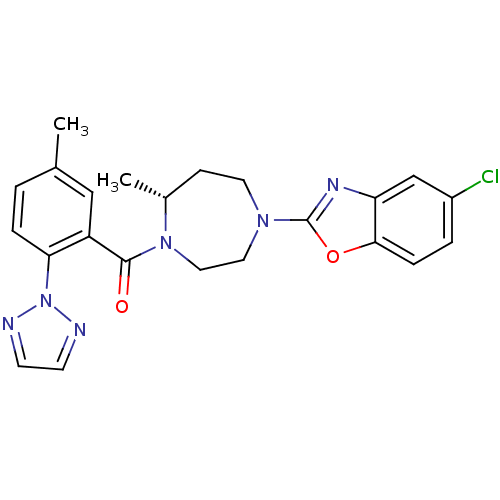
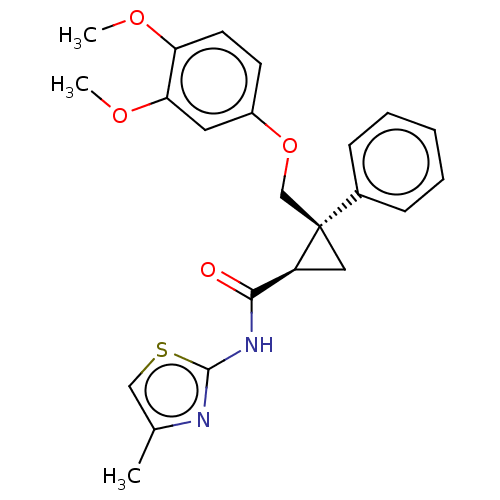
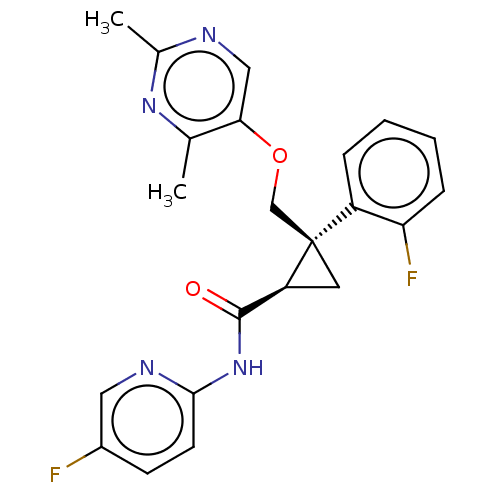
Affinity DataKi: 0.0880nM ΔG°: -59.7kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetBeta-secretase 2(Homo sapiens (Human))
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
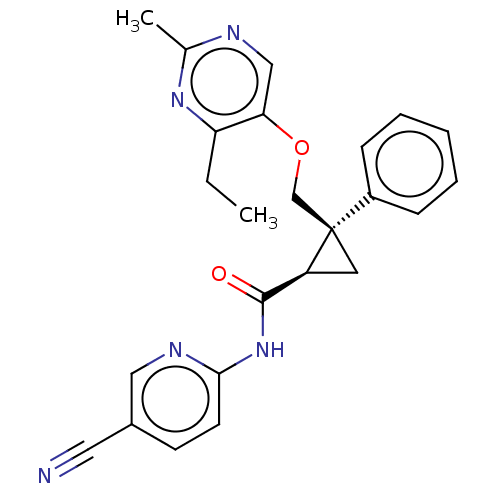
Affinity DataKi: 0.120nMAssay Description:Displacement of [3H] JNJ-962 from BACE2 (unknown origin) expressed in HEK293 cell membranes assessed as inhibition constant at pH 6.2 by scintillatio...More data for this Ligand-Target Pair
TargetBeta-secretase 1(Homo sapiens (Human))
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
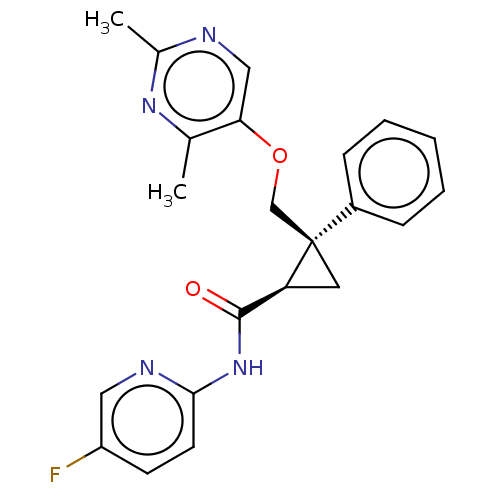
Affinity DataKi: 0.130nMAssay Description:Displacement of [3H] JNJ-962 from BACE1 (unknown origin) expressed in HEK293 cell membranes assessed as inhibition constant at pH 6.2 by scintillatio...More data for this Ligand-Target Pair
Affinity DataKi: 0.160nM ΔG°: -58.2kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetBeta-secretase 1(Homo sapiens (Human))
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
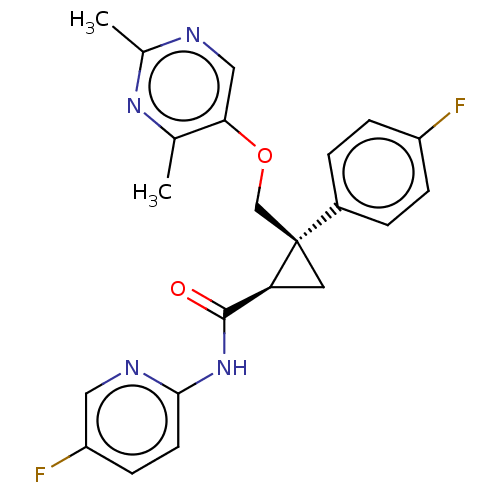
Affinity DataKi: 0.210nMAssay Description:Displacement of [3H]-JNJ962 from BACE1 (unknown origin) expressed in HEK293 cell membrane assessed as inhibition constant by scintillation counting a...More data for this Ligand-Target Pair
Affinity DataKi: 0.290nM ΔG°: -56.6kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetBeta-secretase 2(Homo sapiens (Human))
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Affinity DataKi: 0.290nMAssay Description:Binding affinity to BACE2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.330nM ΔG°: -56.3kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetBeta-secretase 1(Homo sapiens (Human))
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Affinity DataKi: 0.340nMAssay Description:Displacement of [3H] JNJ-962 from BACE1 (unknown origin) expressed in HEK293 cell membranes assessed as inhibition constant at pH 6.2 by scintillatio...More data for this Ligand-Target Pair
TargetBeta-secretase 1(Homo sapiens (Human))
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Affinity DataKi: 0.400nMAssay Description:Displacement of [3H] JNJ-962 from BACE1 (unknown origin) expressed in HEK293 cell membranes assessed as inhibition constant at pH 6.2 by scintillatio...More data for this Ligand-Target Pair
Affinity DataKi: 0.440nMAssay Description:Binding affinity to human OX2R expressed in HEK-293 cells assessed as inhibition of orexin A-induced calcium accumulation by FLIPR assayMore data for this Ligand-Target Pair
TargetBeta-secretase 1(Homo sapiens (Human))
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Affinity DataKi: 0.690nMAssay Description:Binding affinity to BACE1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.740nM ΔG°: -54.2kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetBeta-secretase 1(Homo sapiens (Human))
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Affinity DataKi: 0.800nMAssay Description:Displacement of [3H] JNJ-962 from BACE1 (unknown origin) expressed in HEK293 cell membranes assessed as inhibition constant at pH 6.2 by scintillatio...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nM ΔG°: -52.6kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Binding affinity to human OX1R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.24nM ΔG°: -51.4kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Binding affinity to human OX1R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.70nMAssay Description:Displacement of [125I]-Orexin A from human OX2R expressed in CHO cells after 30 mins by topcount analysisMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of [125I]-Orexin A from human OX2R expressed in CHO cells after 30 mins by topcount analysisMore data for this Ligand-Target Pair
Affinity DataKi: 5.14nM ΔG°: -49.2kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 5.70nMAssay Description:Binding affinity to human OX1R expressed in HEK-293 cells assessed as inhibition of orexin A-induced calcium accumulation by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Binding affinity to human OX1R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Binding affinity to human OX1R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Binding affinity to human OX1R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 7.20nMAssay Description:Displacement of [125I]-Orexin A from human OX2R expressed in CHO cells after 30 mins by topcount analysisMore data for this Ligand-Target Pair
Affinity DataKi: 7.20nMAssay Description:Displacement of [125I]-Orexin A from human OX1R expressed in CHO cells after 30 mins by topcount analysisMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 8.5nMAssay Description:Displacement of [125I]-Orexin A from human OX2R expressed in CHO cells after 30 mins by topcount analysisMore data for this Ligand-Target Pair
Affinity DataKi: 8.91nM ΔG°: -47.8kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
TargetBeta-secretase 2(Homo sapiens (Human))
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Affinity DataKi: 9.60nMAssay Description:Displacement of [3H] JNJ-962 from BACE2 (unknown origin) expressed in HEK293 cell membranes assessed as inhibition constant at pH 6.2 by scintillatio...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair
TargetBeta-secretase 2(Homo sapiens (Human))
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Shionogi Pharmaceutical Research Center
Curated by ChEMBL
Affinity DataKi: 11nMAssay Description:Displacement of [3H] JNJ-962 from BACE2 (unknown origin) expressed in HEK293 cell membranes assessed as inhibition constant at pH 6.2 by scintillatio...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [125I]-Orexin A from human OX2R expressed in CHO cells after 30 mins by topcount analysisMore data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Displacement of [125I]-Orexin A from human OX2R expressed in CHO cells after 30 mins by topcount analysisMore data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Displacement of [125I]-Orexin A from human OX2R expressed in CHO cells after 30 mins by topcount analysisMore data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Binding affinity to human OX1R by radioligand displacement binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Binding affinity to human OX2R by radioligand displacement binding assayMore data for this Ligand-Target Pair

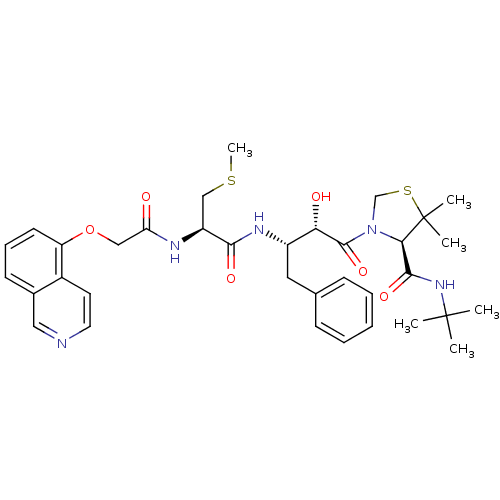

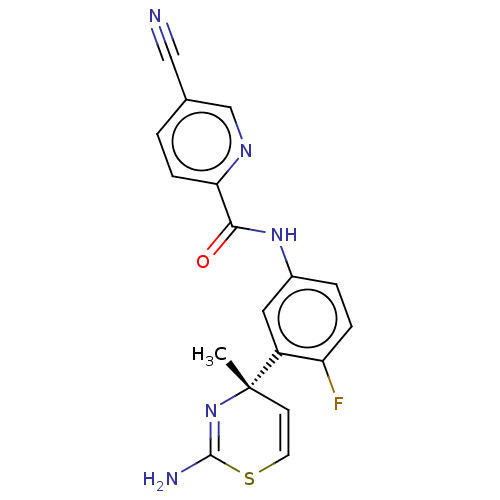
 3D Structure (crystal)
3D Structure (crystal)