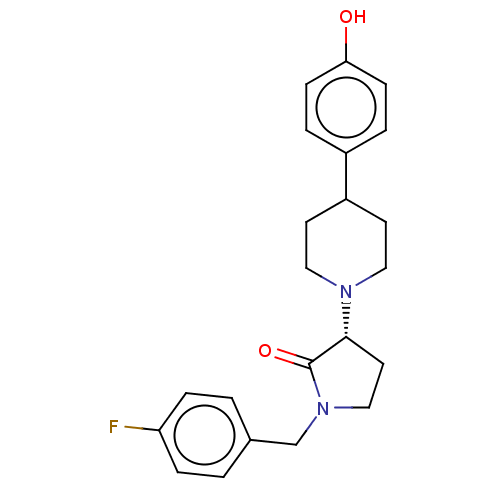
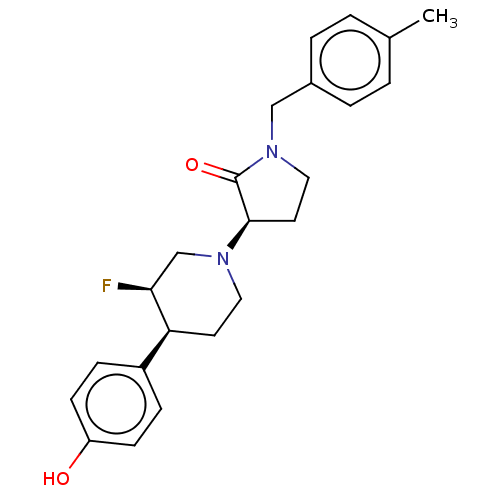
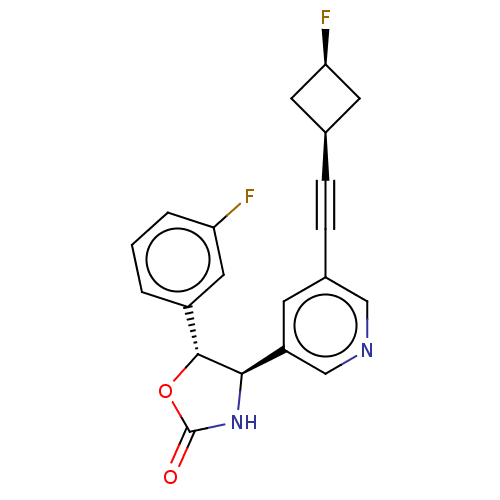
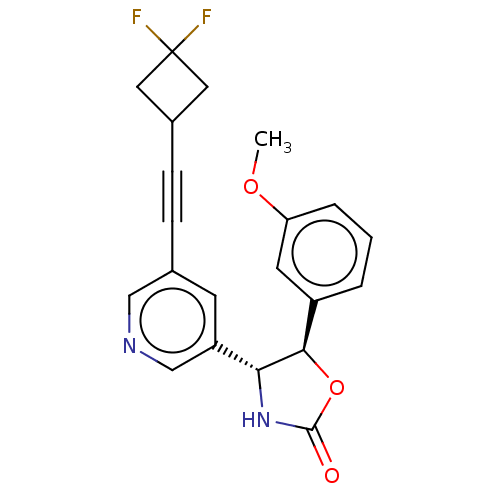
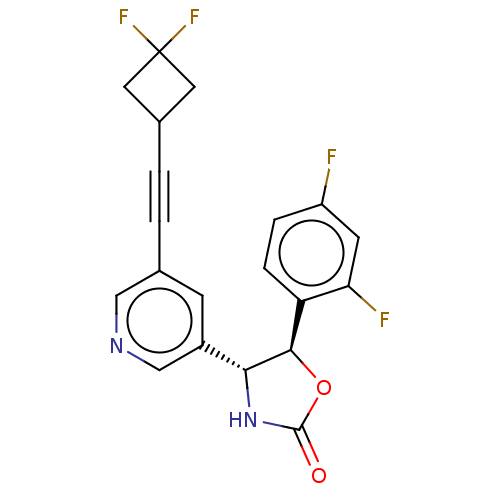
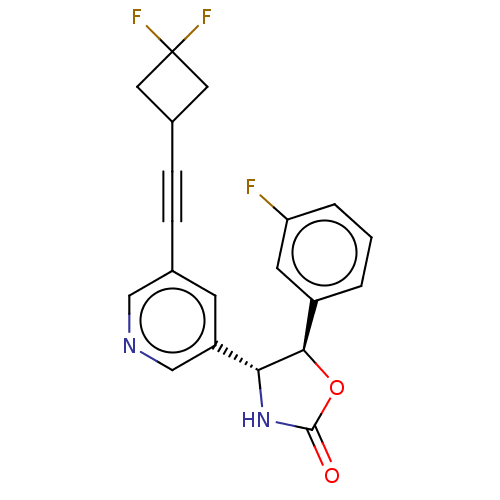
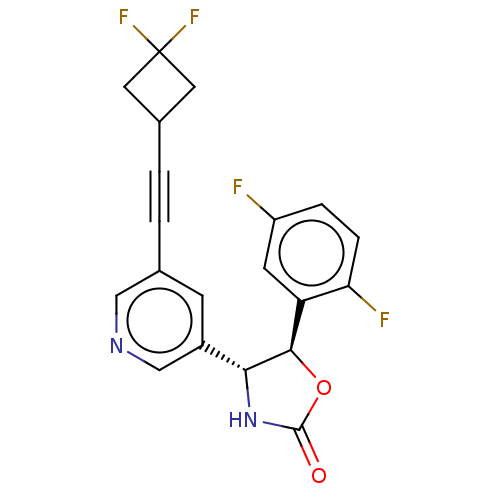
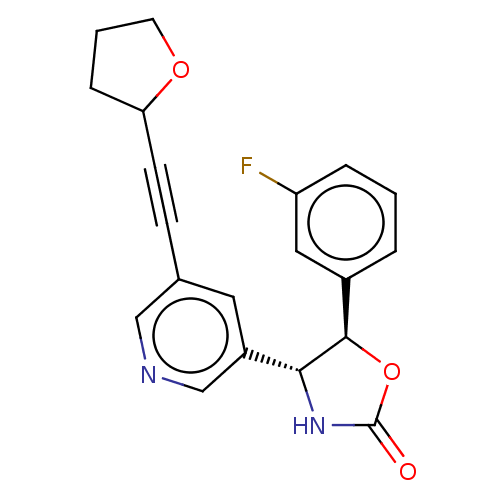
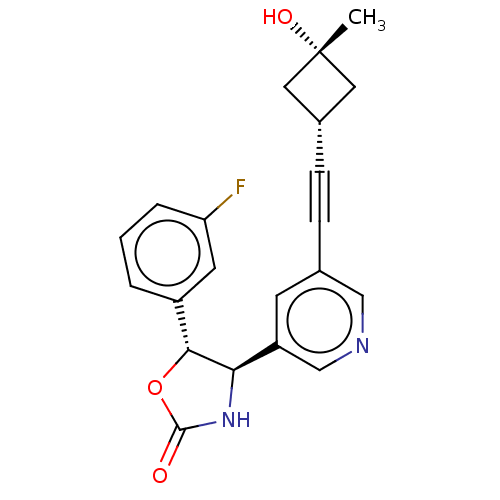
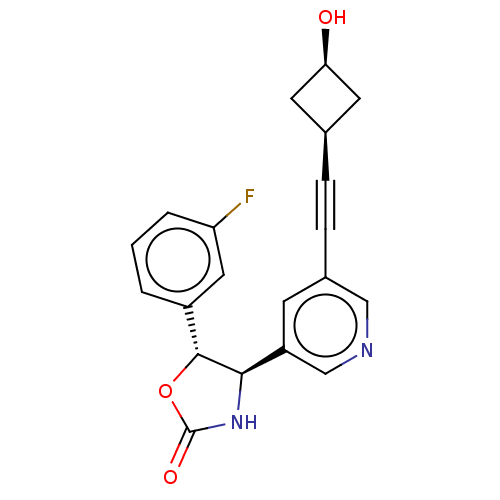
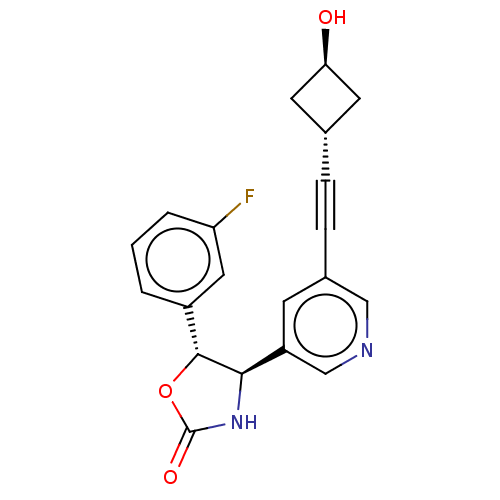
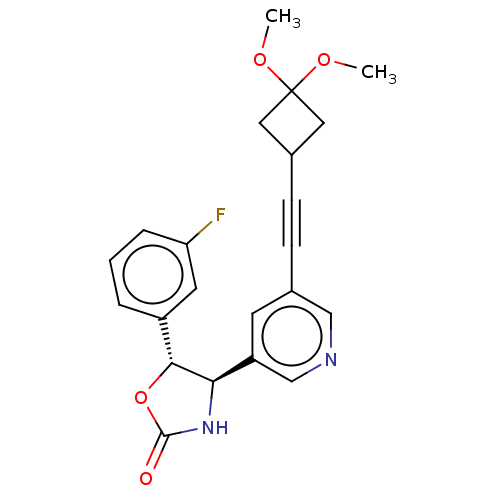
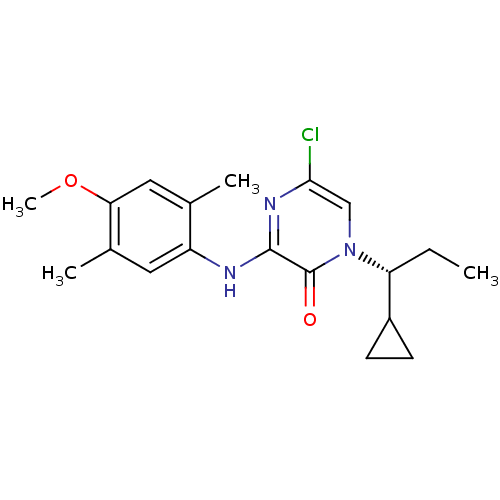
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataKi: 1.40nMAssay Description:Displacement of [3H]Ro 25-6981 from GluN2B receptor in Sprague-Dawley rat forebrain membranes incubated for 1 hr by topcount micro scintillation coun...More data for this Ligand-Target Pair
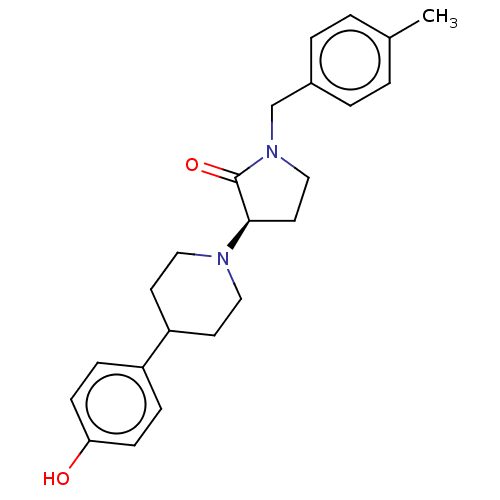
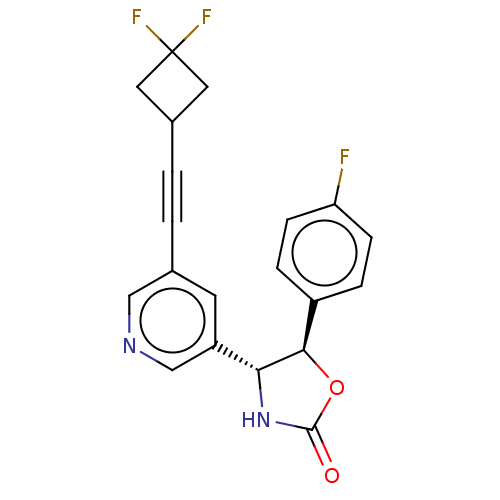
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataKi: 1.40nMAssay Description:Displacement of [3H]Ro 25-6981 from GluN2B receptor in Sprague-Dawley rat forebrain membranes incubated for 1 hr by topcount micro scintillation coun...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
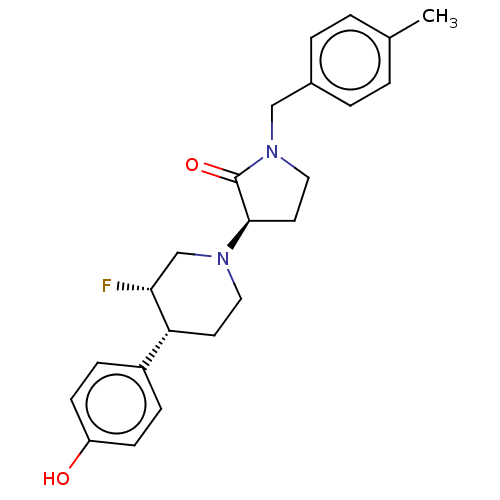
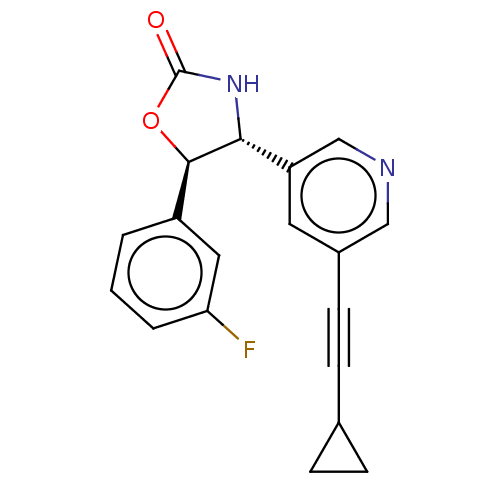
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataKi: 4nMAssay Description:Displacement of [3H]Ro 25-6981 from GluN2B receptor in Sprague-Dawley rat forebrain membranes incubated for 1 hr by topcount micro scintillation coun...More data for this Ligand-Target Pair
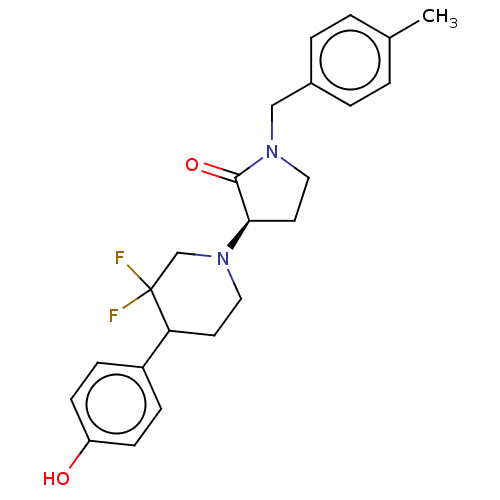
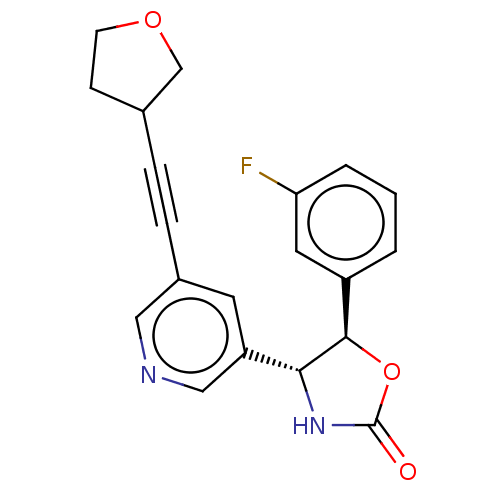
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataKi: 4.30nMAssay Description:Displacement of [3H]Ro 25-6981 from GluN2B receptor in Sprague-Dawley rat forebrain membranes incubated for 1 hr by topcount micro scintillation coun...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataKi: 4.40nMAssay Description:Displacement of [3H]Ro 25-6981 from GluN2B receptor in Sprague-Dawley rat forebrain membranes incubated for 1 hr by topcount micro scintillation coun...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Homo sapiens (Human))
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataKi: 6.30nMAssay Description:Binding affinity to GluN2B receptor in human cortexMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataKi: 7.70nMAssay Description:Displacement of [3H]Ro 25-6981 from GluN2B receptor in Sprague-Dawley rat forebrain membranes incubated for 1 hr by topcount micro scintillation coun...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataKi: 8.40nMAssay Description:Displacement of [3H]Ro 25-6981 from GluN2B receptor in Sprague-Dawley rat forebrain membranes incubated for 1 hr by topcount micro scintillation coun...More data for this Ligand-Target Pair
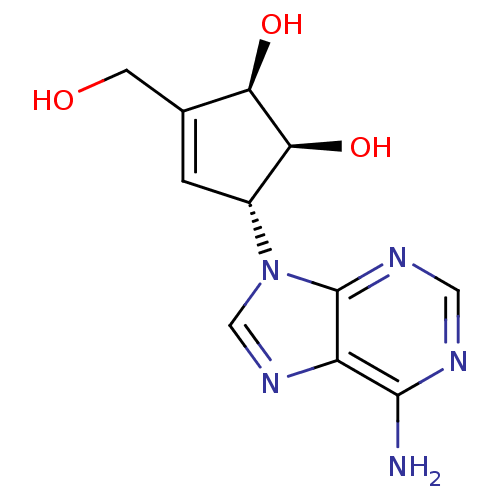
Affinity DataKi: 8.40nMAssay Description:Reversible inhibition of rat liver S- adenosyl-L-homocysteine hydrolase using SAH as substrate by spectrophotometric methodMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataKi: 11nMAssay Description:Displacement of [3H]Ro 25-6981 from GluN2B receptor in Sprague-Dawley rat forebrain membranes incubated for 1 hr by topcount micro scintillation coun...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 36nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 38nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 47nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 47nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 59nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 64nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 67nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 88nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 95nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 108nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 259nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 323nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 372nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 490nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 580nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 710nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 900nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 1.67E+3nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 1.90E+3nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 5.00E+3nMAssay Description:Displacement of [3H]MPEPy from human mGluR5 expressed in cell membranes after 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 8.25E+4nMAssay Description:Time-dependent inhibition of CYP3A4 in human liver microsomes assessed as inhibition constant by Kitz-Wilson plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 8.25E+4nMAssay Description:Time-dependent inhibition of CYP3A4 in human liver microsomes assessed as inhibition constant by Kitz-Wilson plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 8.30E+4nMAssay Description:Time dependent inhibition of CYP3A4 in human liver microsomes assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 4.70E+5nMAssay Description:Time dependent inhibition of Hafnia alvei lysine decarboxylase using L-lysine by Kitz-Wilson analysisMore data for this Ligand-Target Pair
Affinity DataKi: 6.30E+5nMAssay Description:Time dependent inhibition of Hafnia alvei lysine decarboxylase using L-lysine by Kitz-Wilson analysisMore data for this Ligand-Target Pair
TargetCorticotropin-releasing factor receptor 1(Rattus norvegicus (rat))
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.210nMAssay Description:Displacement of [125I]Tyr-o-CRF from CRF1 receptor in rat frontal cortex homogenate by binding titration assayMore data for this Ligand-Target Pair
TargetCorticotropin-releasing factor receptor 1(Rattus norvegicus (rat))
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.240nMAssay Description:Displacement of [125I]Tyr-o-CRF from CRF1 receptor in rat frontal cortex homogenate by binding titration assayMore data for this Ligand-Target Pair
TargetCorticotropin-releasing factor receptor 1(Rattus norvegicus (rat))
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.260nMAssay Description:Displacement of [125I]Tyr-ovine-CRF from CRF1 receptor in rat frontal cortex homogenate by gamma countingMore data for this Ligand-Target Pair
TargetCorticotropin-releasing factor receptor 1(Rattus norvegicus (rat))
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.270nMAssay Description:Displacement of [125I]Tyr-ovine-CRF from CRF1 receptor in rat frontal cortex homogenate by gamma countingMore data for this Ligand-Target Pair
TargetCorticotropin-releasing factor receptor 1(Rattus norvegicus (rat))
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.280nMAssay Description:Displacement of [125I]Tyr-o-CRF from CRF1 receptor in rat frontal cortex homogenate by binding titration assayMore data for this Ligand-Target Pair
TargetCorticotropin-releasing factor receptor 1(Rattus norvegicus (rat))
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.290nMAssay Description:Displacement of [125I]Tyr-ovine-CRF from CRF1 receptor in rat frontal cortex homogenate by gamma countingMore data for this Ligand-Target Pair
TargetCorticotropin-releasing factor receptor 1(Rattus norvegicus (rat))
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:Displacement of [125I]Tyr-o-CRF from CRF1 receptor in rat frontal cortex homogenate by binding titration assayMore data for this Ligand-Target Pair
TargetCorticotropin-releasing factor receptor 1(Rattus norvegicus (rat))
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataIC50: 0.420nMAssay Description:Displacement of [125I]Tyr-ovine-CRF from CRF1 receptor in rat frontal cortex homogenate by gamma countingMore data for this Ligand-Target Pair